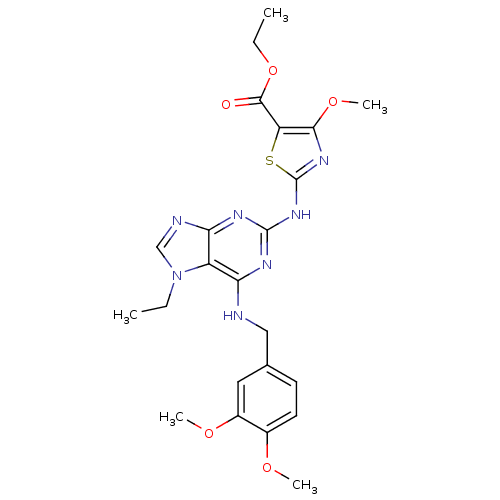
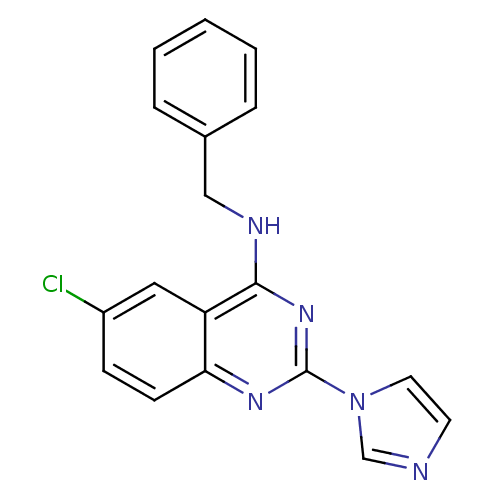
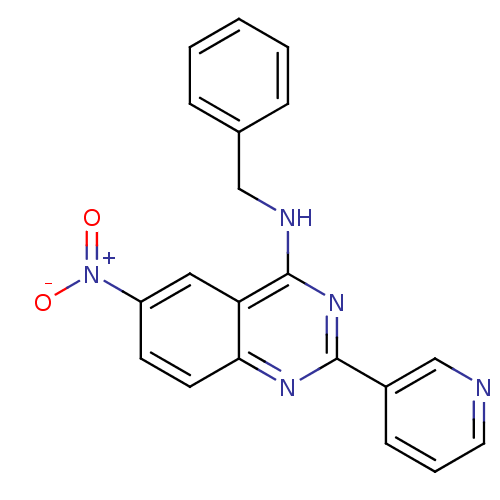
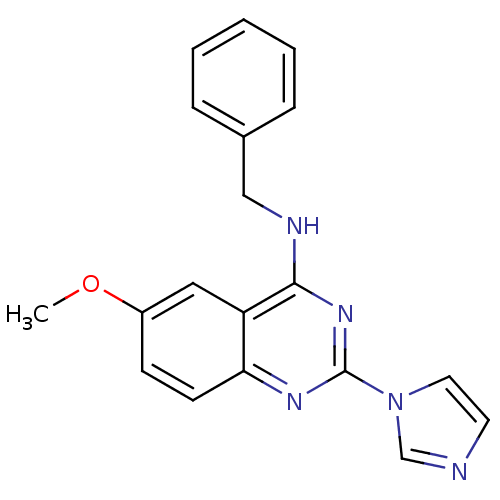
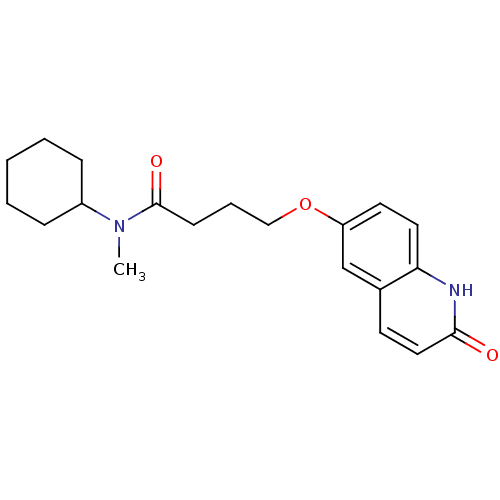
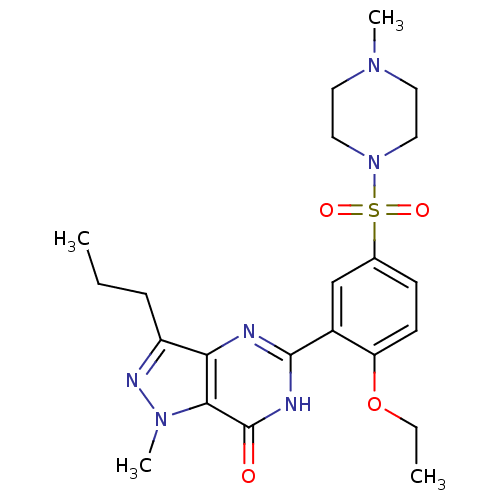
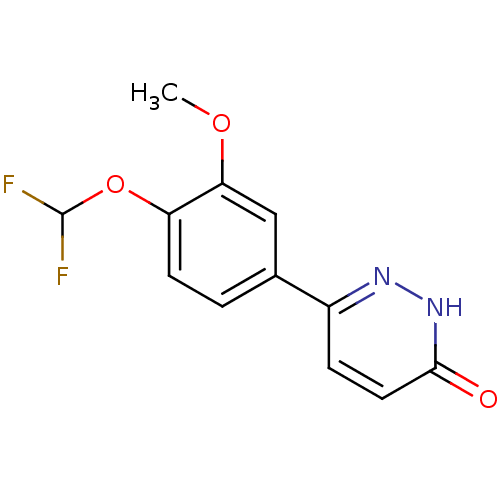
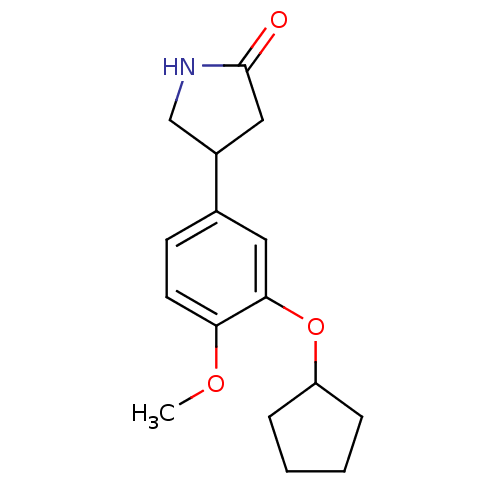
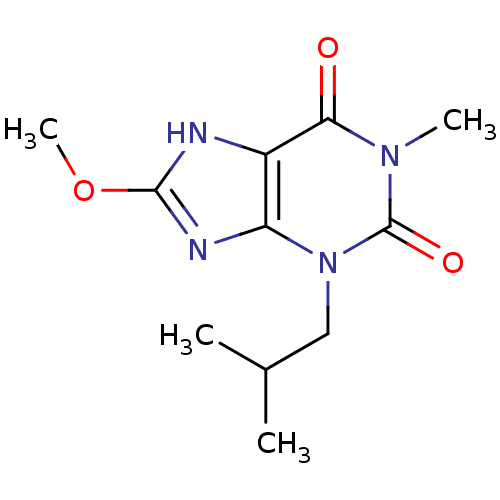
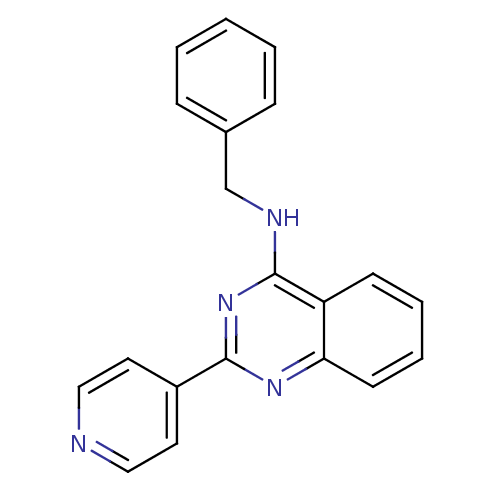
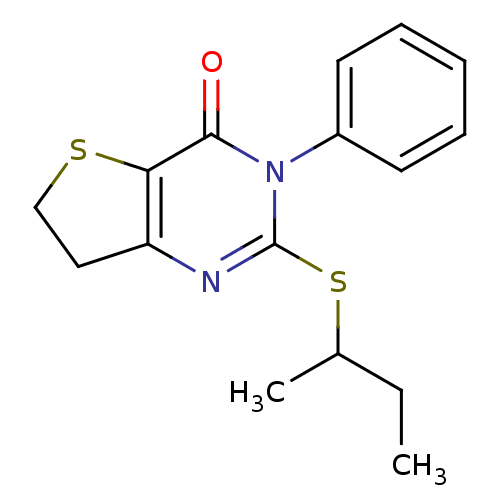
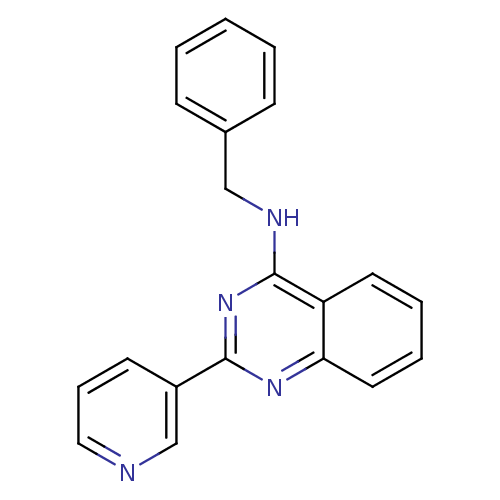
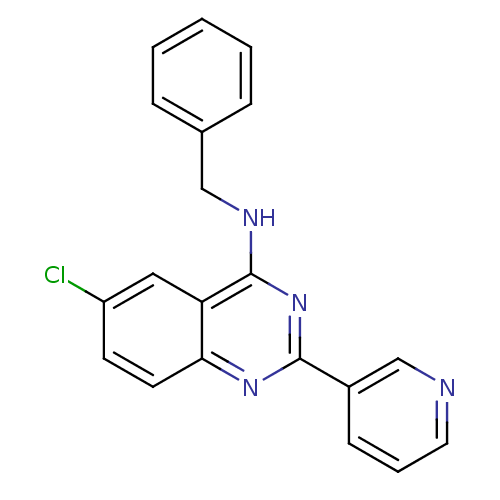
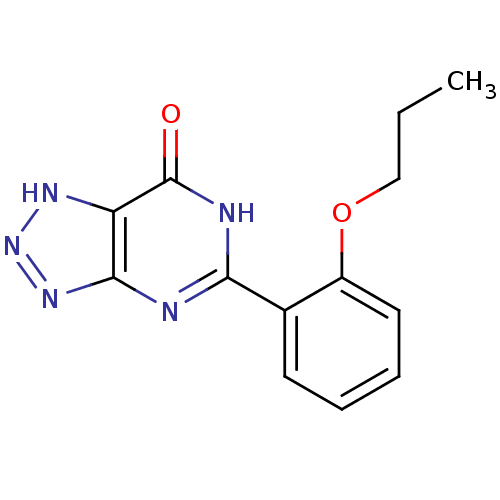
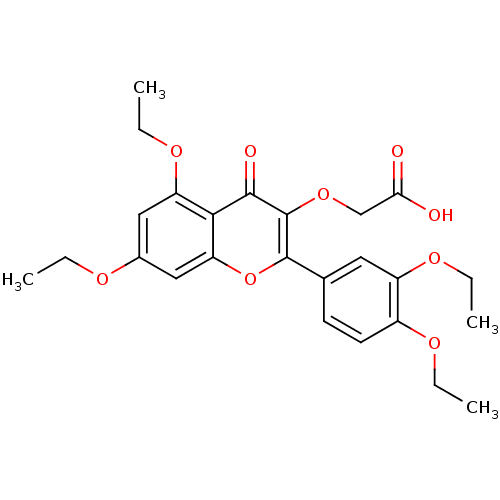
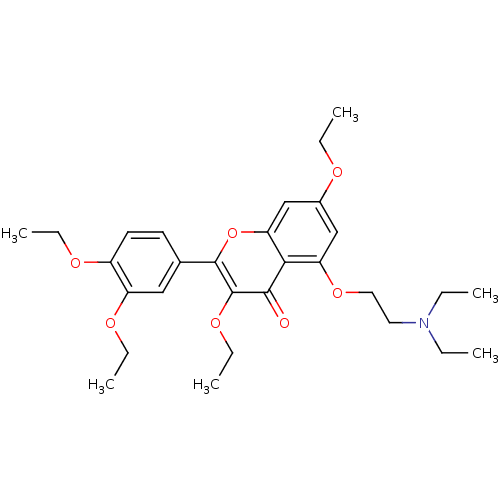
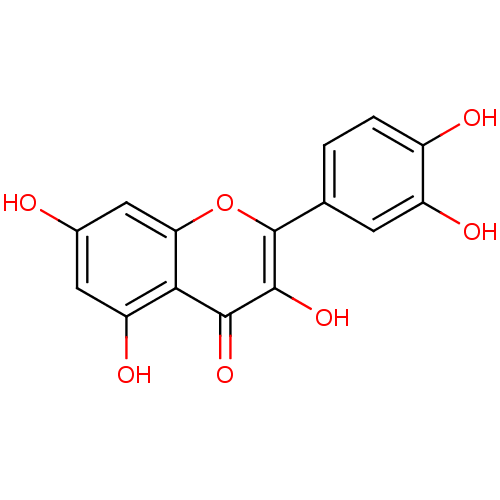
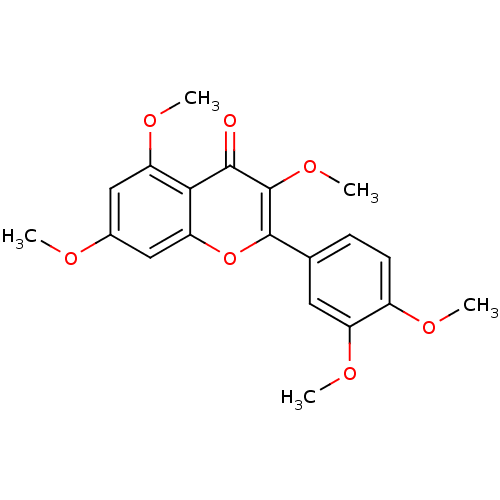
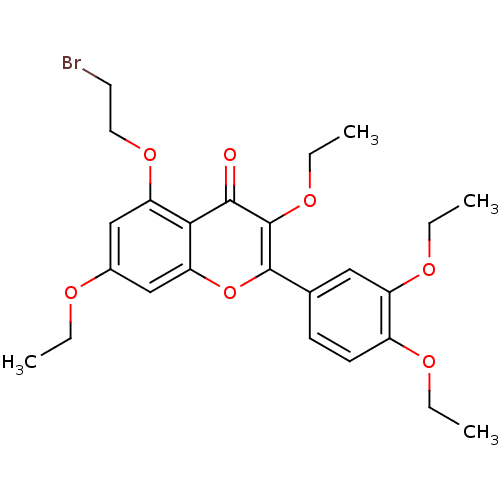
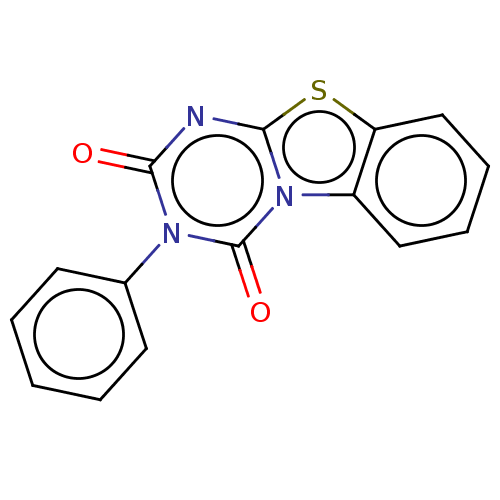
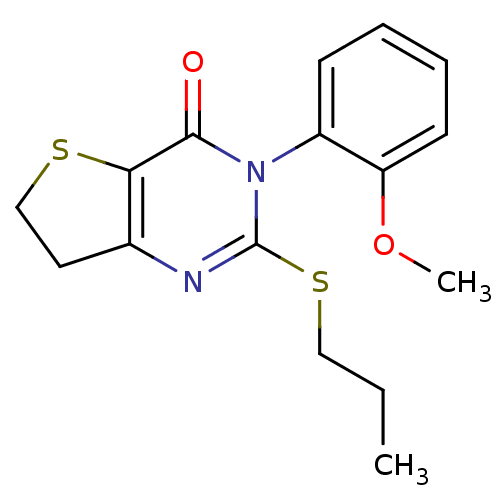
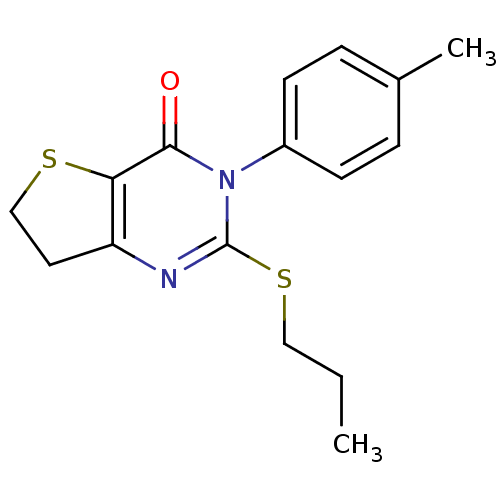
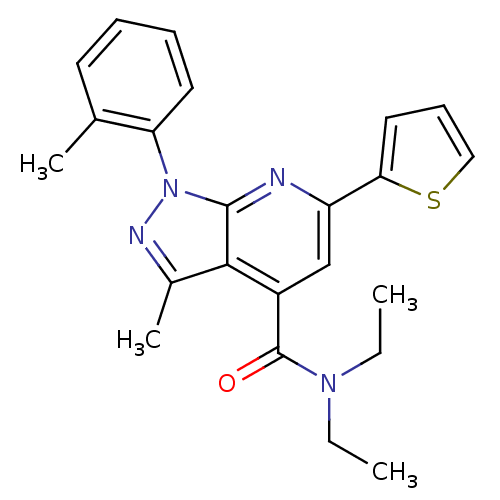
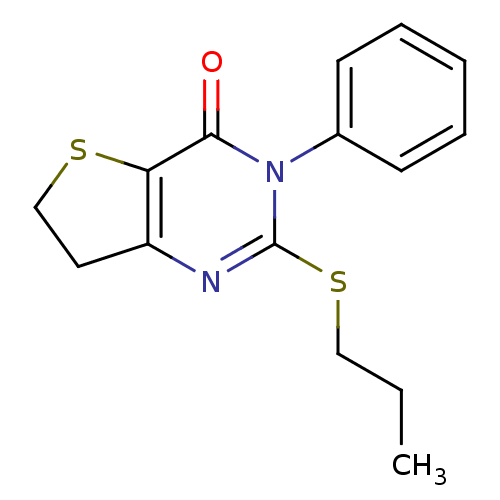
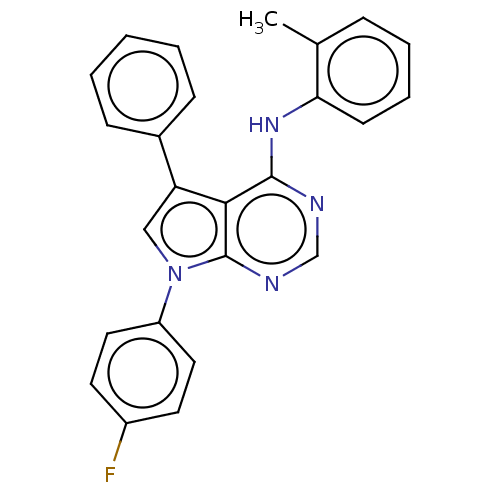
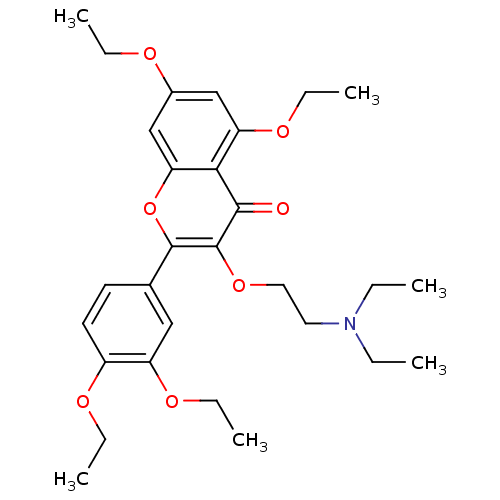
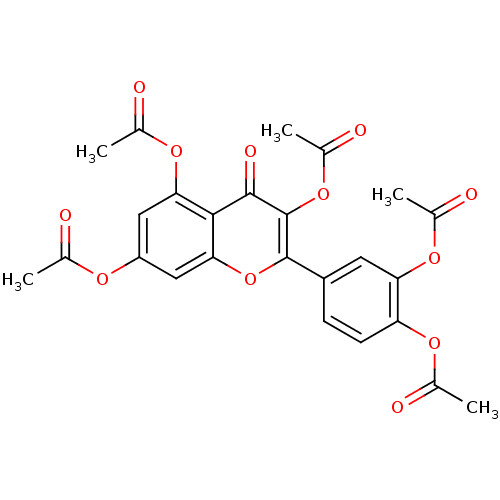
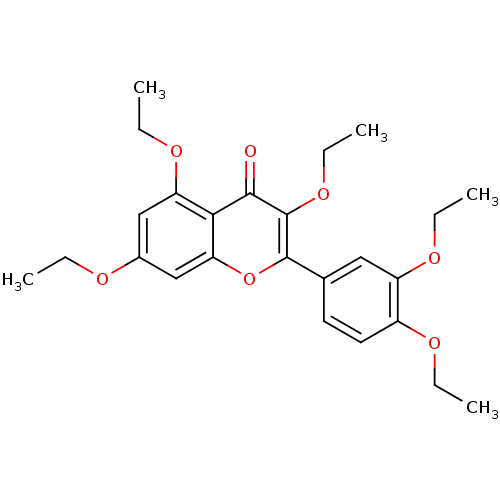
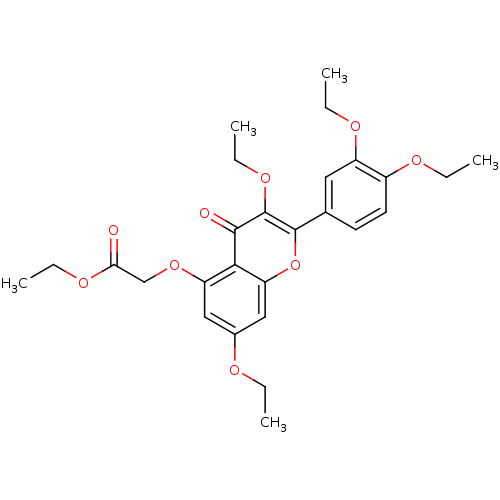
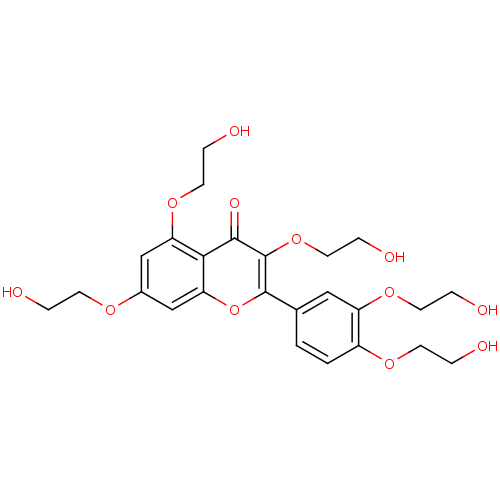
Affinity DataKd: 2.80nMAssay Description:Binding affinity to PDE2A in rat striatal membranes after 30 mins by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 4.20nMAssay Description:Displacement of radiolabeled 4-(azetidin-1-yl)-3-[5-[4-(trifluoromethyl)phenyl]-1H-pyrazol-4-yl]-1-(tritritiomethyl)pyrazolo[3,4-d]pyrimidine from PD...More data for this Ligand-Target Pair
Affinity DataEC50: 670nMAssay Description:Inhibition of PDE2a in rat primary cortical neurons assessed as increase in Bay 41-8543-stimulated intracellular cGMP level preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibitory concentration against phosphodiesterase 2 from rat kidneyMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of rat brain Phosphodiesterase 2More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of rat brain Phosphodiesterase 2More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of rat brain Phosphodiesterase 2More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibitory concentration against phosphodiesterase 2 from rat kidneyMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibitory concentration against phosphodiesterase 2 from rat kidneyMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibitory concentration against phosphodiesterase 2 from rat kidneyMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibitory concentration against phosphodiesterase 2 from rat kidneyMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibitory concentration against phosphodiesterase 2 from rat kidneyMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nMAssay Description:Inhibition of rat brain Phosphodiesterase 2More data for this Ligand-Target Pair
Affinity DataIC50: 1.84E+4nMAssay Description:In vitro enzyme assays were conducted via the Ba(OH)2 precipiation method using recombinant human PDE1C, PDE3B, PDE5A1, PDE6C, PDE8A, PDE9A2, PDE10A1...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+4nMAssay Description:Inhibition of rat brain Phosphodiesterase 2More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+4nMAssay Description:Inhibition of rat brain Phosphodiesterase 2More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+4nMAssay Description:Inhibition of rat brain Phosphodiesterase 2More data for this Ligand-Target Pair
Affinity DataIC50: 2.45E+4nMAssay Description:Inhibition of rat brain particulate cGMP phosphodiesteraseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of rat brain Phosphodiesterase 2More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+4nMAssay Description:In vitro enzyme assays were conducted via the Ba(OH)2 precipiation method using recombinant human PDE1C, PDE3B, PDE5A1, PDE6C, PDE8A, PDE9A2, PDE10A1...More data for this Ligand-Target Pair
Affinity DataIC50: 4.80E+4nMAssay Description:Inhibition of rat brain particulate cGMP phosphodiesteraseMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of rat brain Phosphodiesterase 2More data for this Ligand-Target Pair
Affinity DataIC50: 6.31E+4nMAssay Description:Inhibition of rat brain particulate cGMP phosphodiesteraseMore data for this Ligand-Target Pair
Affinity DataIC50: 7.10E+4nMAssay Description:Inhibition of rat brain particulate cGMP phosphodiesteraseMore data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+4nMAssay Description:Inhibition of rat brain particulate cGMP phosphodiesteraseMore data for this Ligand-Target Pair
Affinity DataIC50: 8.13E+4nMAssay Description:Inhibition of rat brain particulate cGMP phosphodiesteraseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.60E+4nMAssay Description:Inhibition of rat brain particulate cGMP phosphodiesteraseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.60E+4nMAssay Description:Inhibition of rat brain particulate cGMP phosphodiesteraseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:In vitro enzyme assays were conducted via the Ba(OH)2 precipiation method using recombinant human PDE1C, PDE3B, PDE5A1, PDE6C, PDE8A, PDE9A2, PDE10A1...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:In vitro enzyme assays were conducted via the Ba(OH)2 precipiation method using recombinant human PDE1C, PDE3B, PDE5A1, PDE6C, PDE8A, PDE9A2, PDE10A1...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:In vitro enzyme assays were conducted via the Ba(OH)2 precipiation method using recombinant human PDE1C, PDE3B, PDE5A1, PDE6C, PDE8A, PDE9A2, PDE10A1...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of rat brain Phosphodiesterase 2More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:In vitro enzyme assays were conducted via the Ba(OH)2 precipiation method using recombinant human PDE1C, PDE3B, PDE5A1, PDE6C, PDE8A, PDE9A2, PDE10A1...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:In vitro enzyme assays were conducted via the Ba(OH)2 precipiation method using recombinant human PDE1C, PDE3B, PDE5A1, PDE6C, PDE8A, PDE9A2, PDE10A1...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:In vitro enzyme assays were conducted via the Ba(OH)2 precipiation method using recombinant human PDE1C, PDE3B, PDE5A1, PDE6C, PDE8A, PDE9A2, PDE10A1...More data for this Ligand-Target Pair
Affinity DataIC50: 1.26E+5nMAssay Description:Inhibition of rat brain particulate cGMP phosphodiesteraseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.75E+5nMAssay Description:Inhibition of rat brain particulate cGMP phosphodiesteraseMore data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+5nMAssay Description:Inhibition of rat brain particulate cGMP phosphodiesteraseMore data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+5nMAssay Description:Inhibition of rat brain particulate cGMP phosphodiesteraseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of rat brain particulate cGMP phosphodiesteraseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+6nMAssay Description:Inhibition of rat brain particulate cGMP phosphodiesteraseMore data for this Ligand-Target Pair
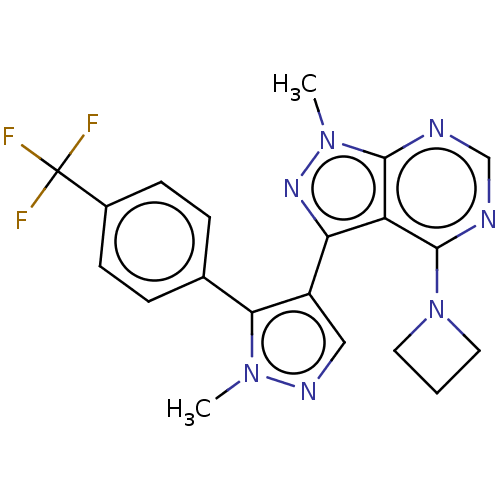
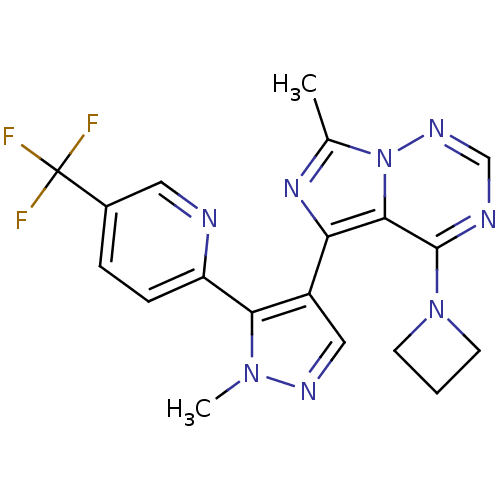
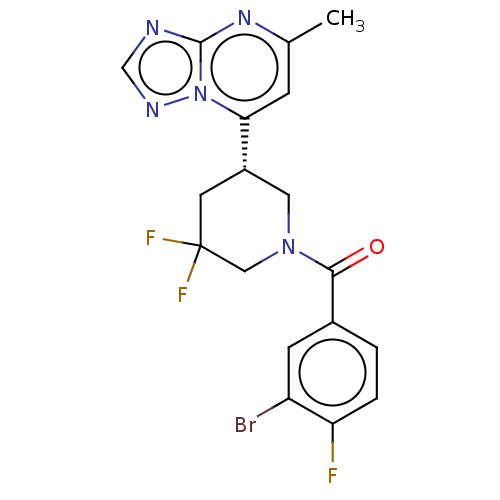
 3D Structure (crystal)
3D Structure (crystal)