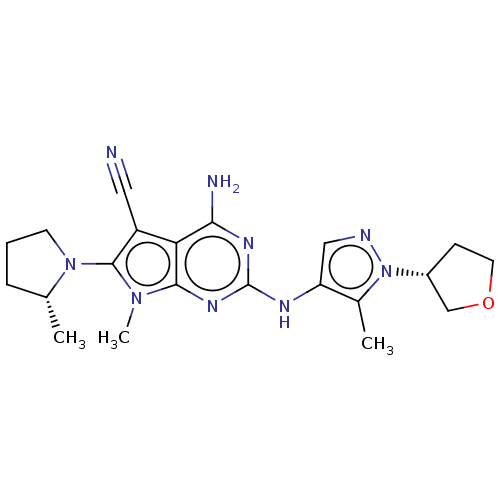
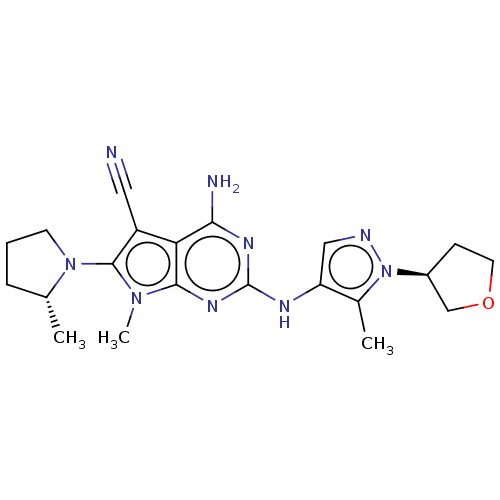
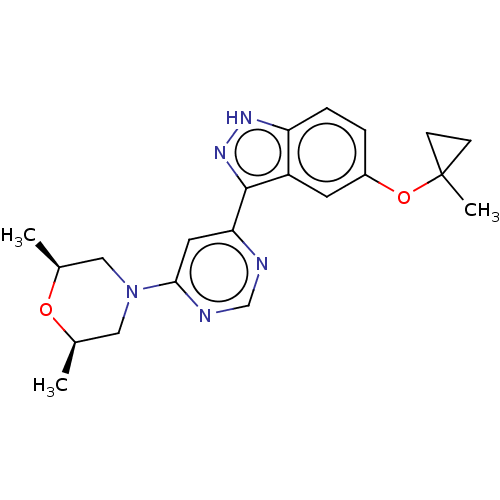
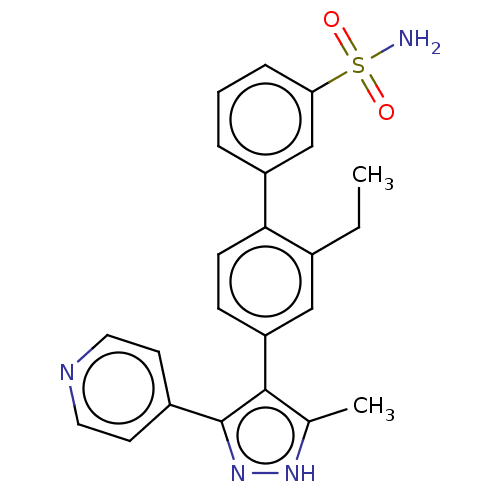
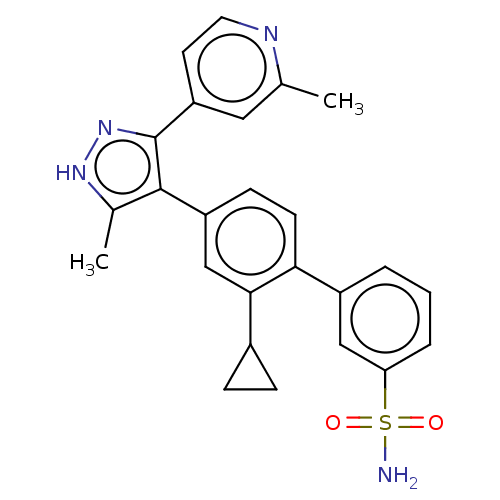
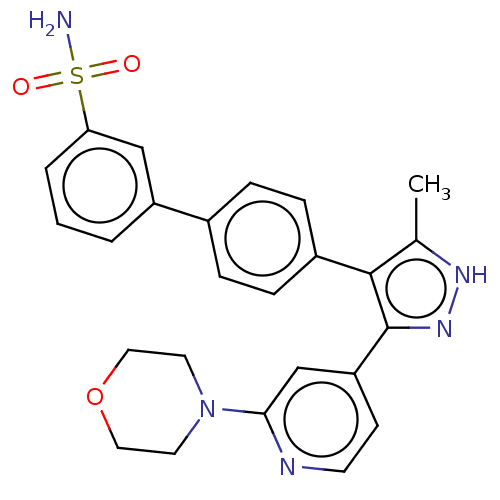
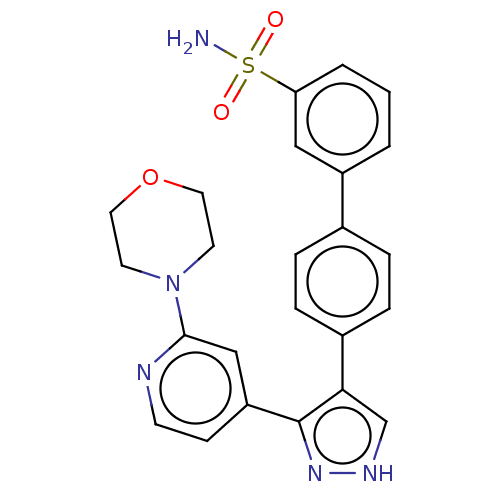
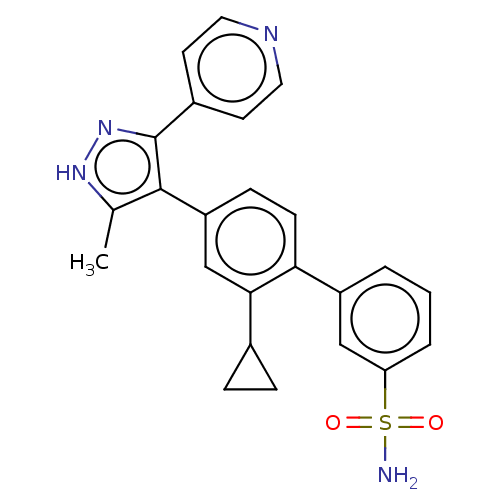
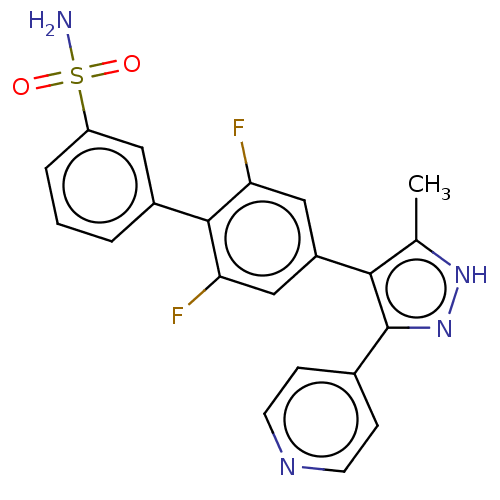
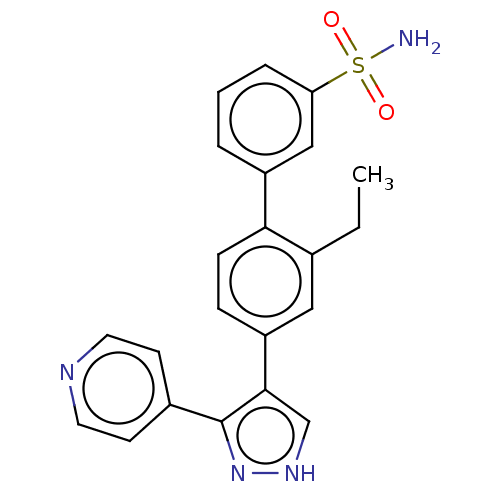
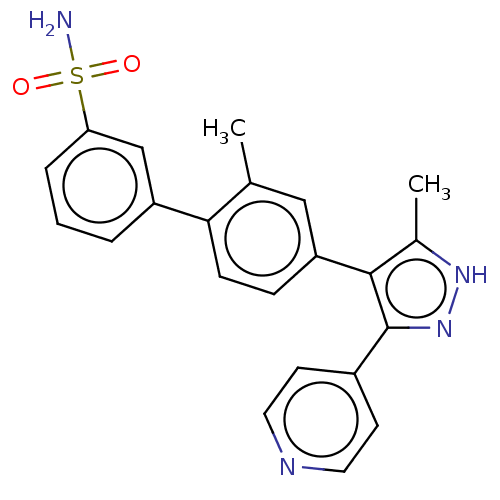
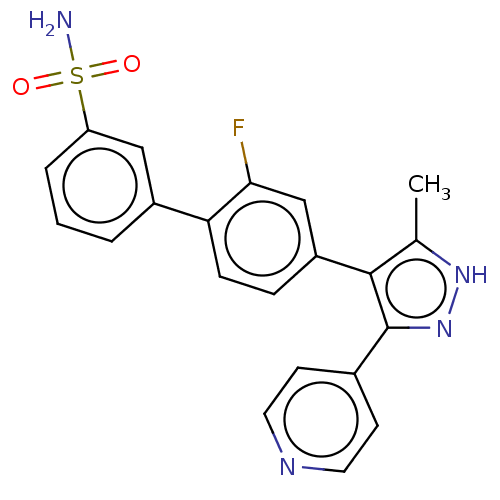
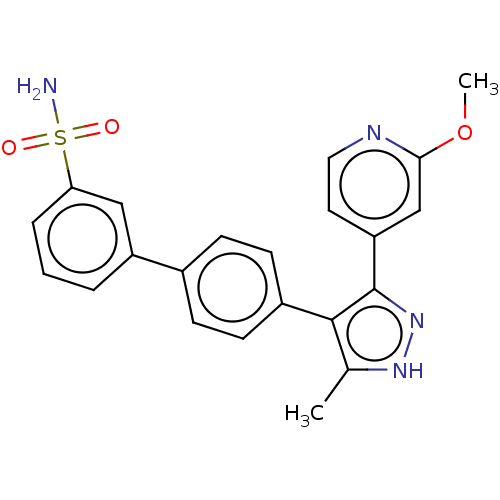
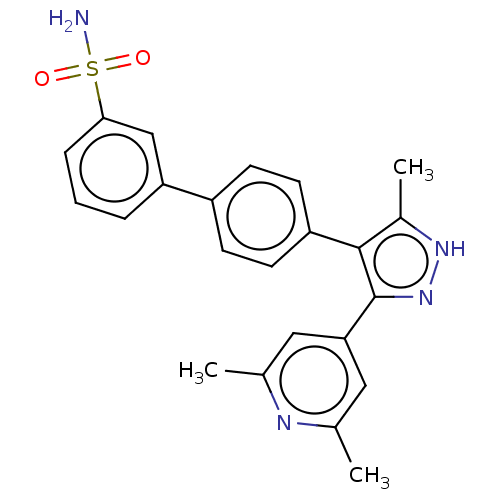
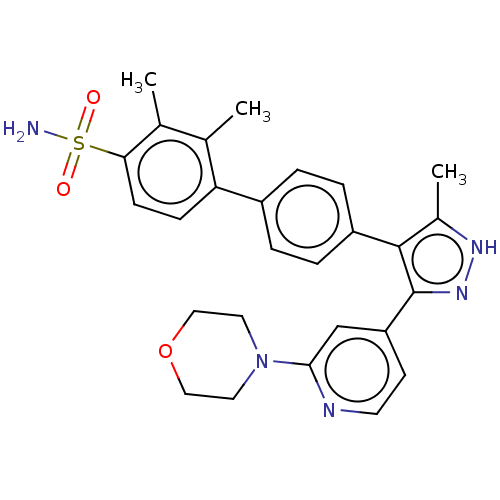
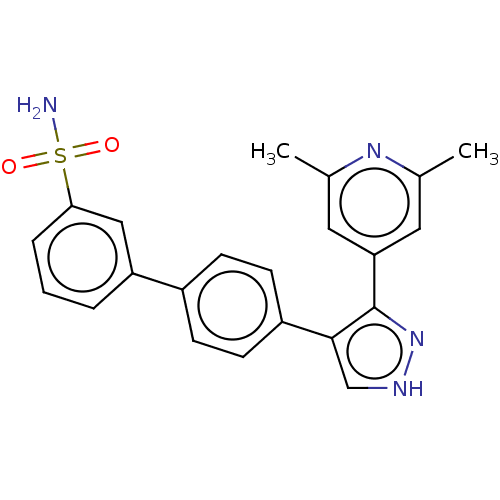
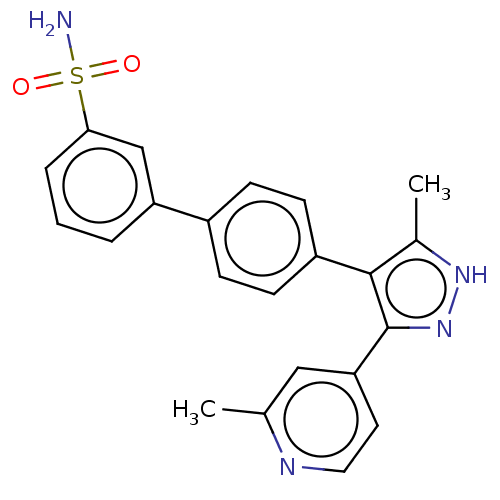
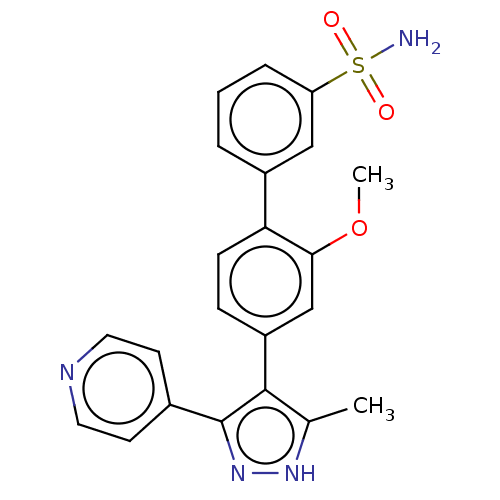
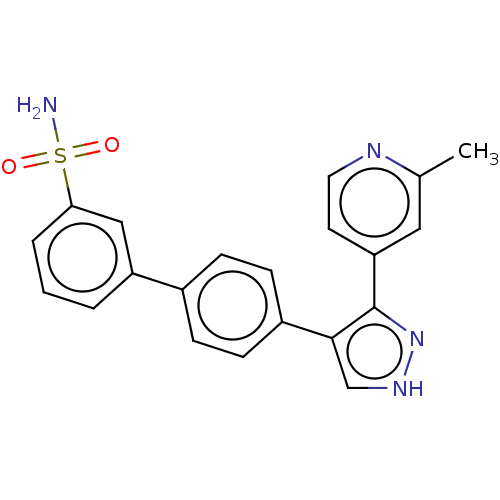
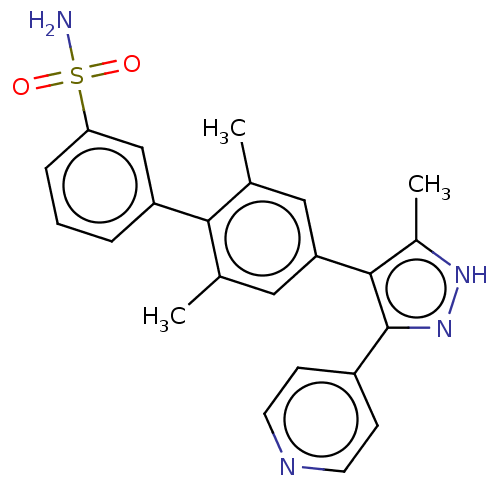
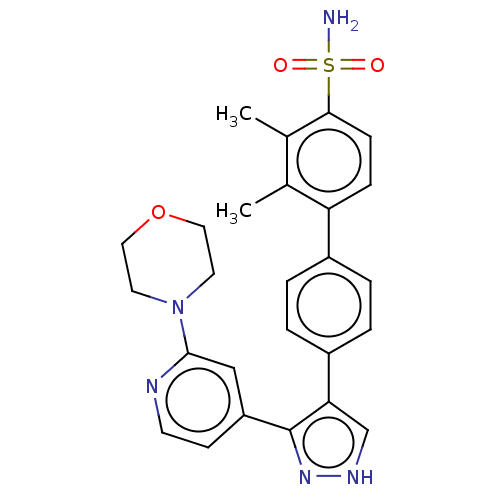
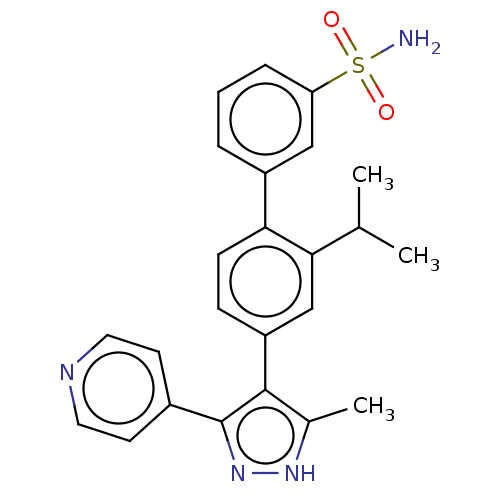
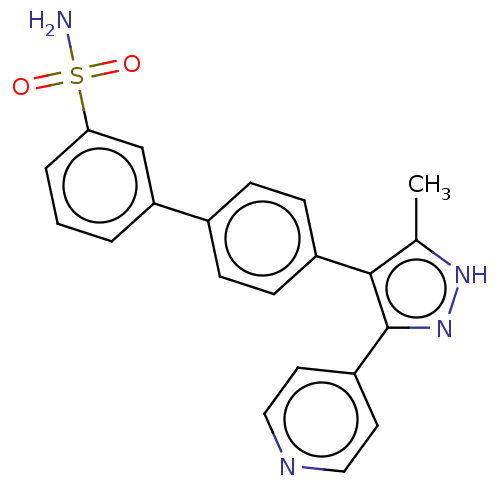
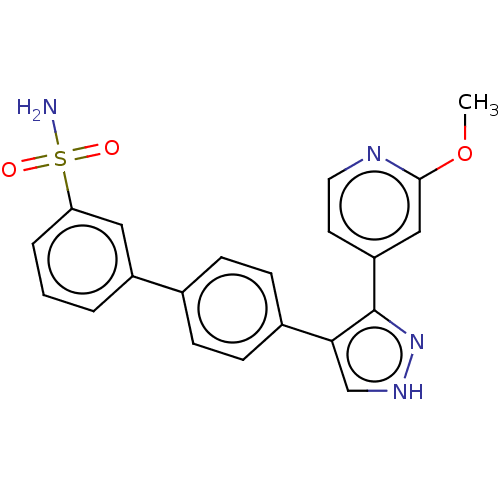
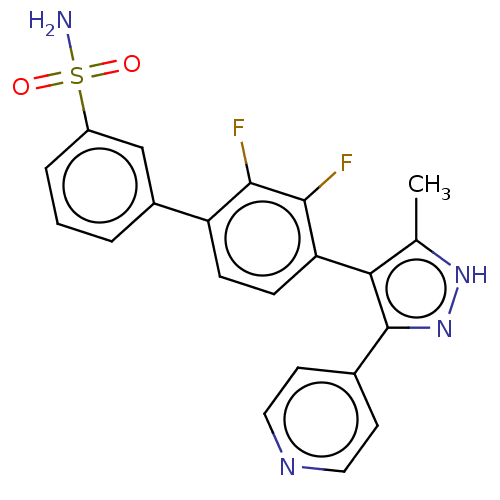
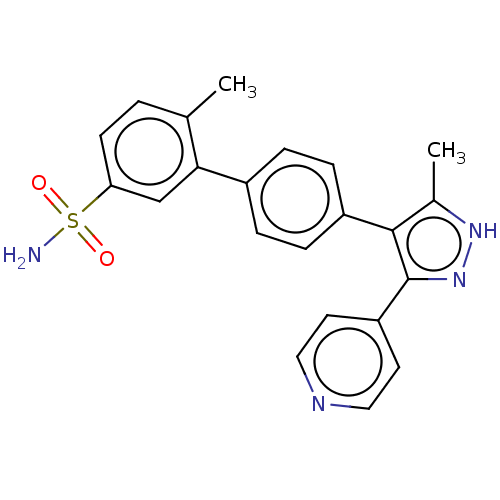
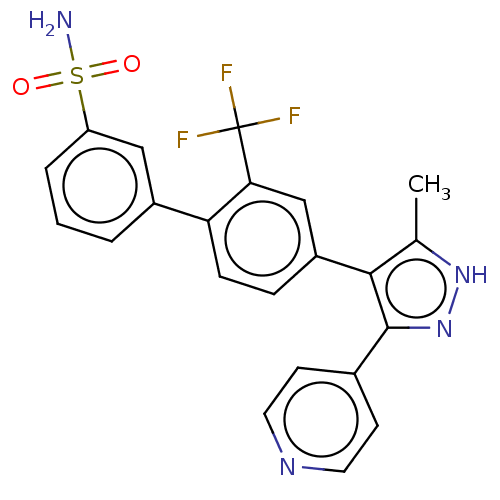
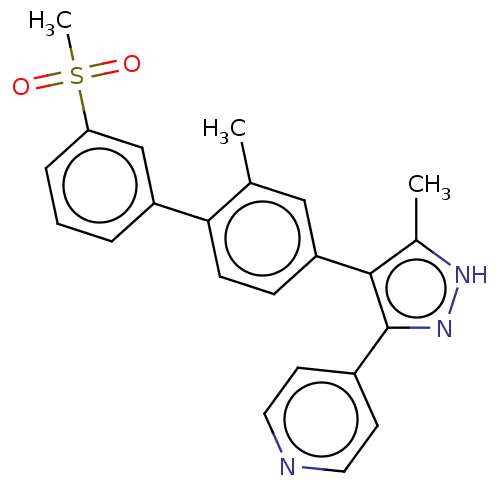
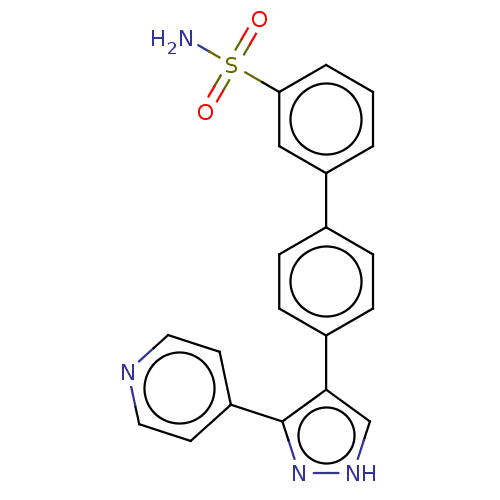
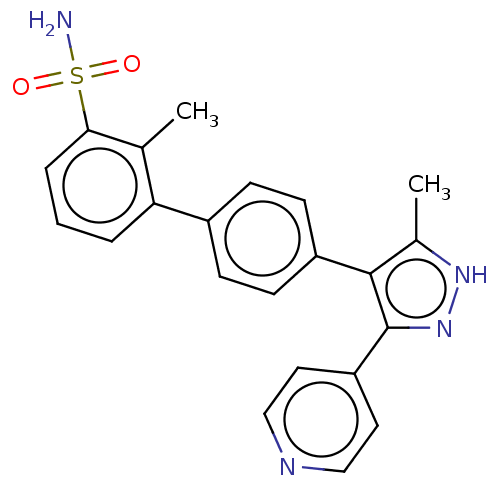
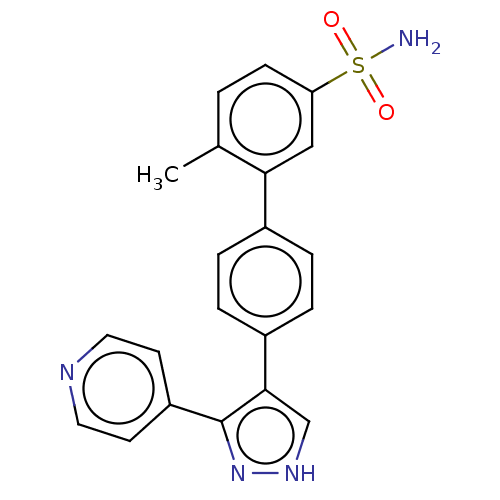
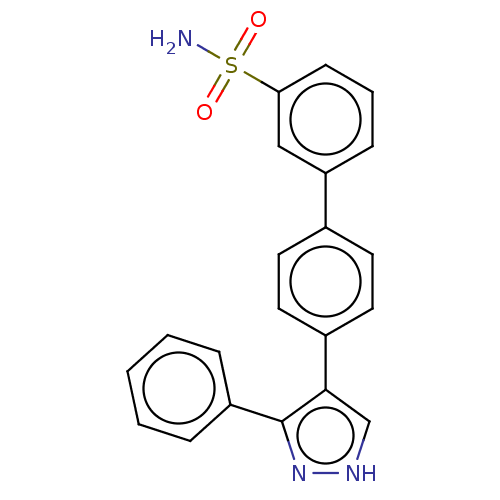
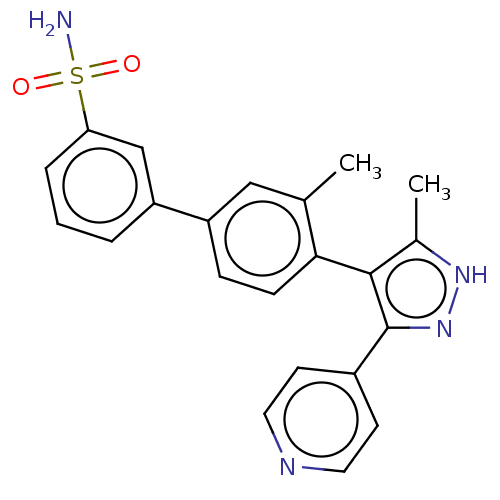
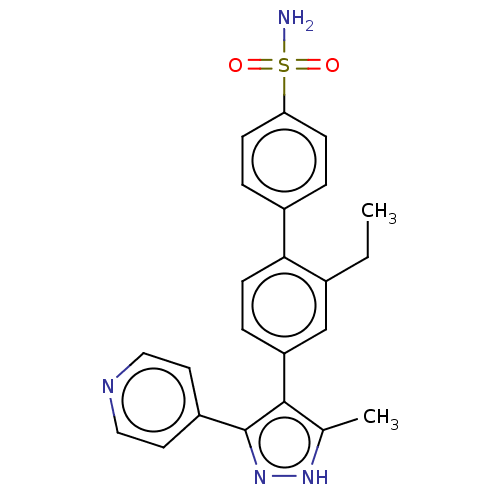
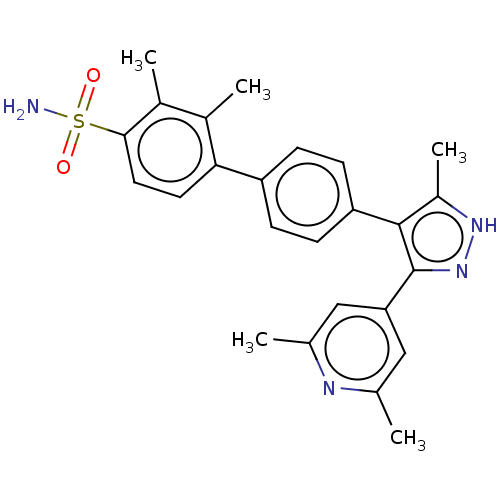
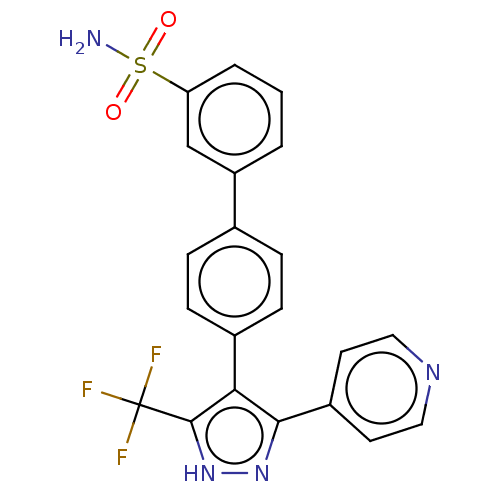
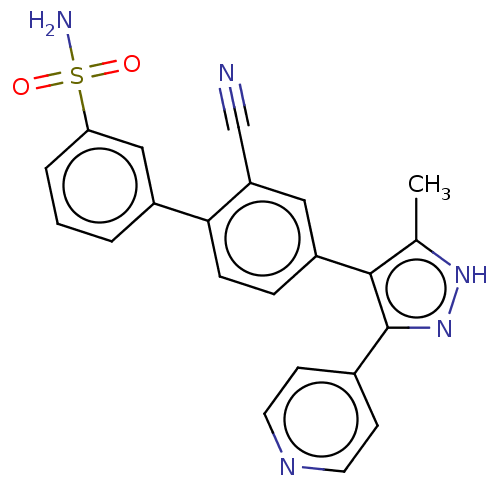
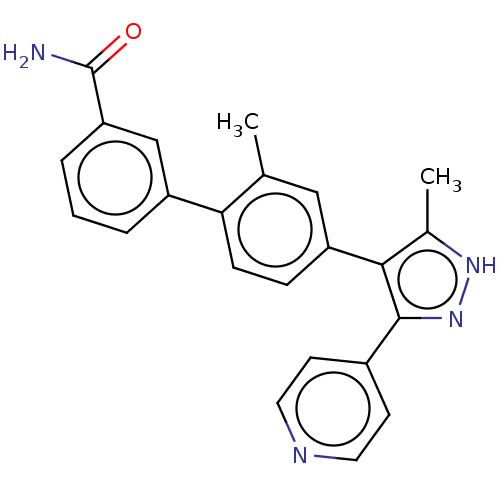
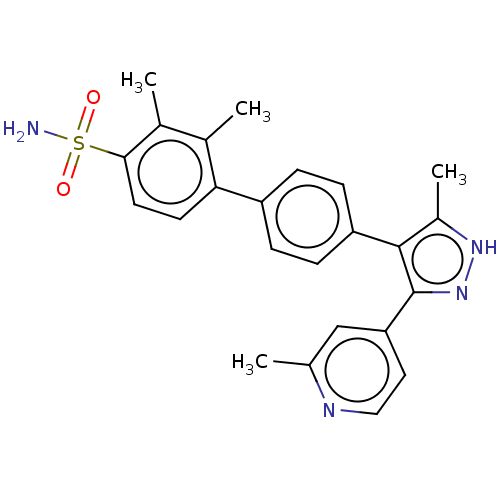
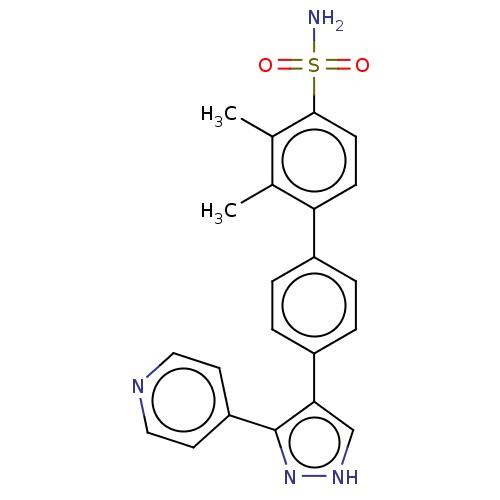
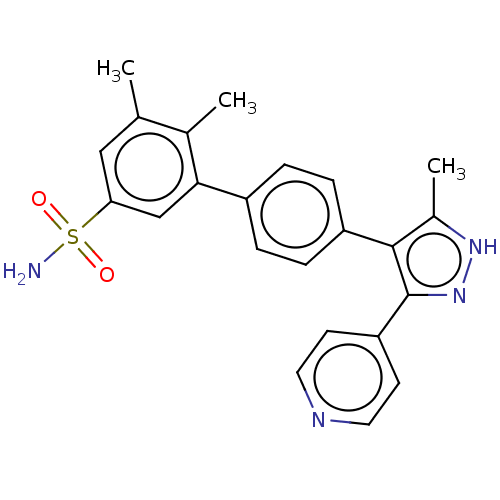
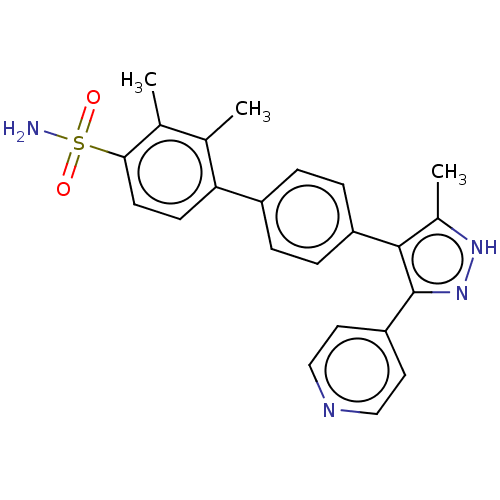
Affinity DataIC50: 11nMAssay Description:Inhibition of LRRK2-WT in C57BL/6 mouse brain at 10 to 150 mg/kg and measured after 1 hrs by western blottingMore data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:PROTAC activity at CRBN/LRRK2 in mouse RAW264.7 cells assessed as reduction of protein degradation measured after 24 hrs by Western blot analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Inhibition of LRRK2-WT-pSER935 in C57BL/6 mouse brain at 10 to 150 mg/kg and measured after 1 hrs by western blottingMore data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Inhibition of LRRK2-WT in C57BL/6 mouse brain at 10 to 150 mg/kg and measured after 1 hrs by western blottingMore data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Inhibition of LRRK2-WT-pSER935 in C57BL/6 mouse brain at 10 to 150 mg/kg and measured after 1 hrs by western blottingMore data for this Ligand-Target Pair
Affinity DataEC50: 30nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMAssay Description:Inhibition of mouse LRRK2More data for this Ligand-Target Pair
Affinity DataEC50: 260nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 280nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 290nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 340nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 390nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 390nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 600nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 600nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 690nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 750nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 760nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 800nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 920nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.08E+3nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.28E+3nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.38E+3nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.44E+3nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.50E+3nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.71E+3nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 2.10E+3nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 2.31E+3nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 3.12E+3nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 3.20E+3nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 4.78E+3nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 4.90E+3nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 5.10E+3nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 5.40E+3nMAssay Description:Inhibition of mouse full-length wild type LRRK2 transfected in HEK293 cells using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-F...More data for this Ligand-Target Pair
Affinity DataEC50: 6.40E+3nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.22E+4nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.22E+4nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.34E+4nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.43E+4nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.50E+4nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.67E+4nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 1.96E+4nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 2.17E+4nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 5.70E+4nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: 7.60E+4nMAssay Description:Inhibition of mouse full-length wild type LRRK2 transfected in HEK293 cells using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-F...More data for this Ligand-Target Pair
Affinity DataEC50: 8.00E+4nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of mouse full-length wild type LRRK2 transfected in HEK293 cells using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-F...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of mouse full-length wild type LRRK2 transfected in HEK293 cells using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-F...More data for this Ligand-Target Pair