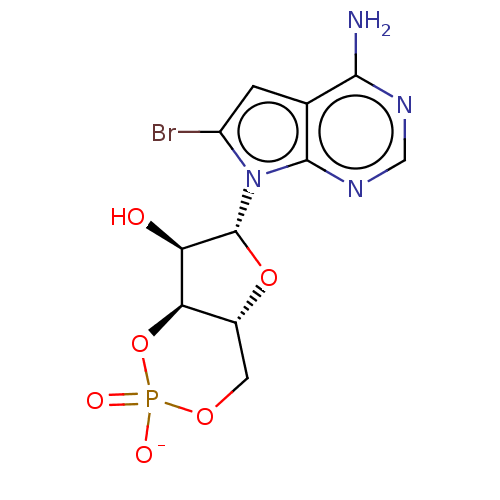
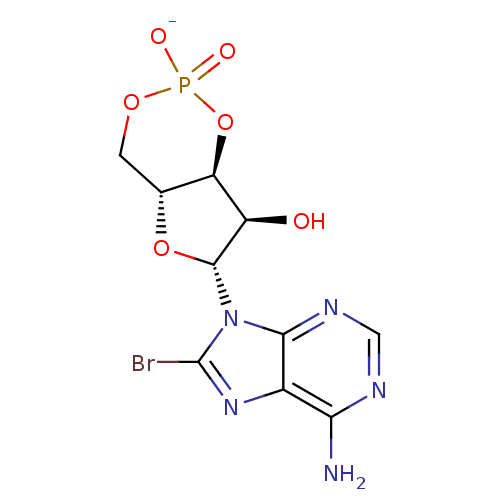
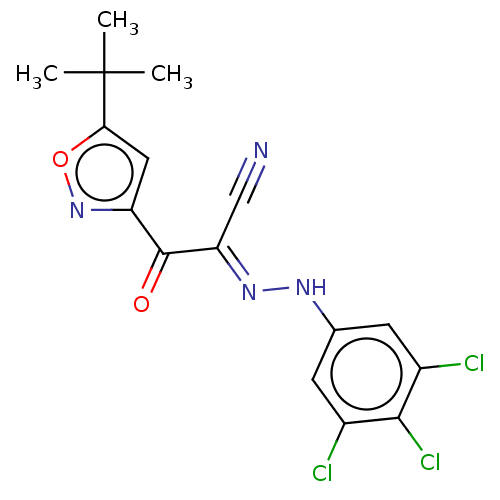
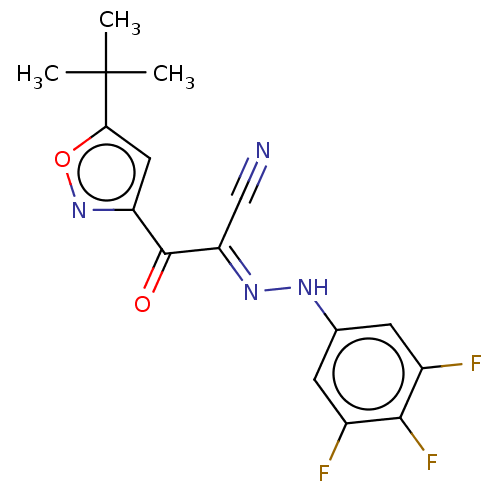
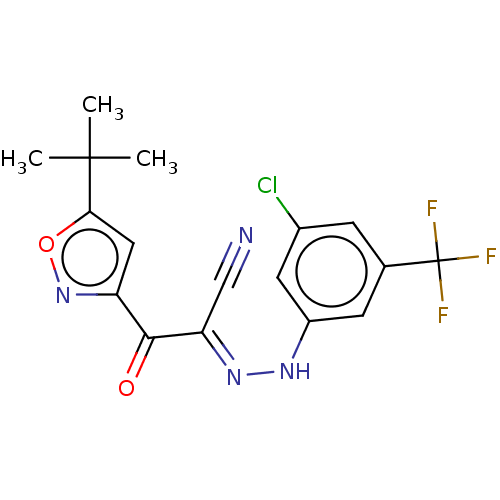
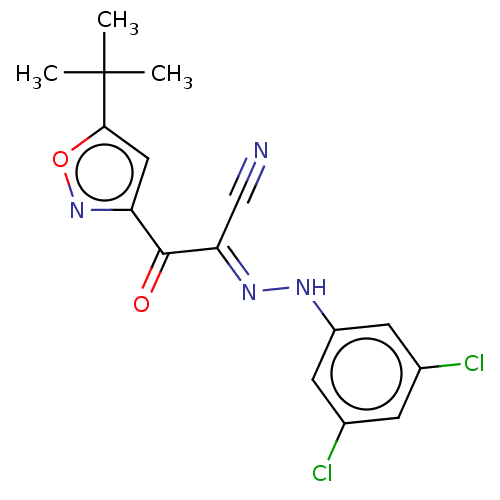
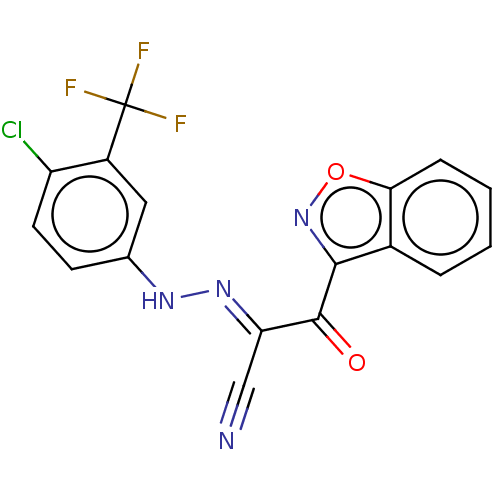
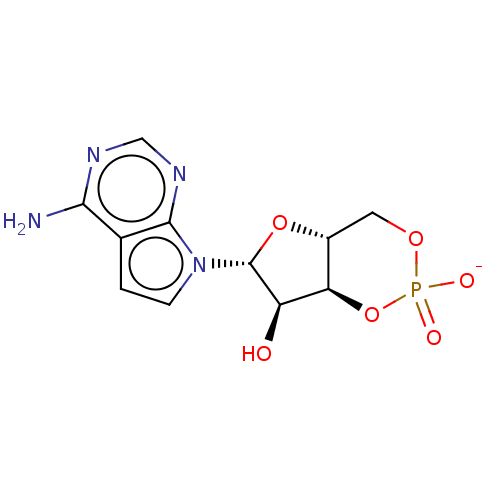
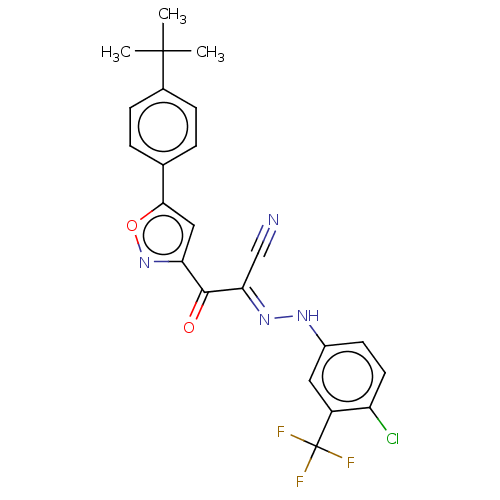
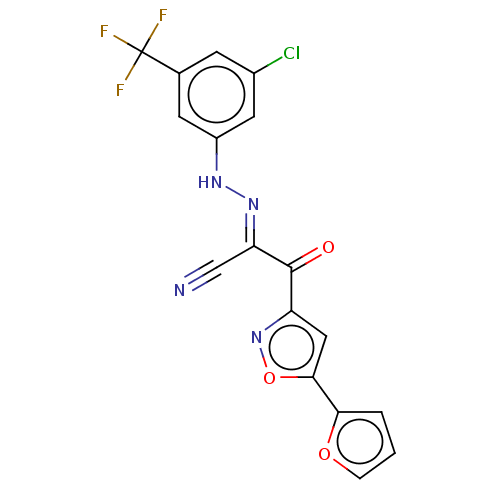
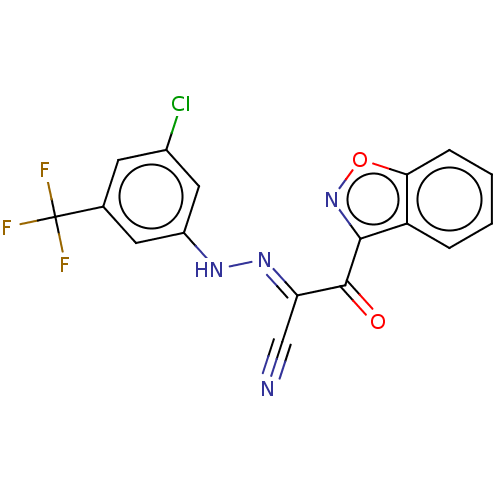
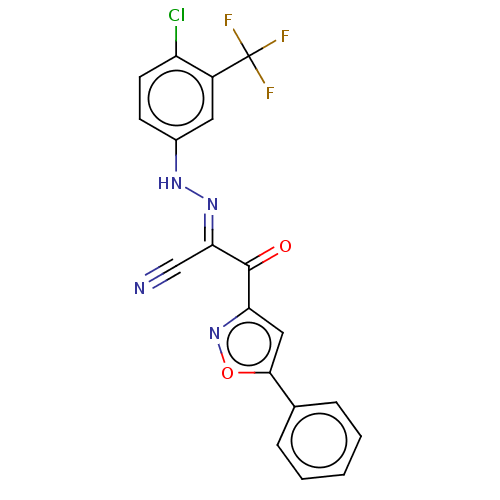
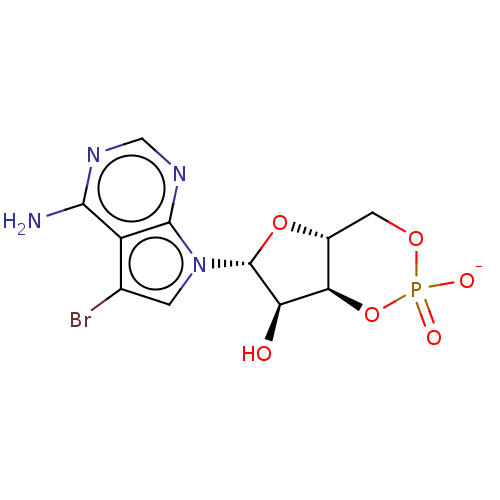
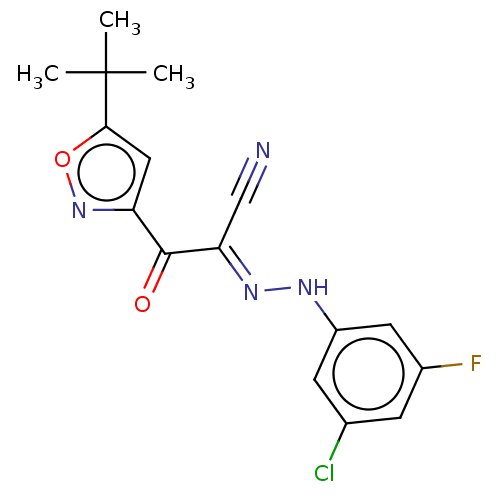
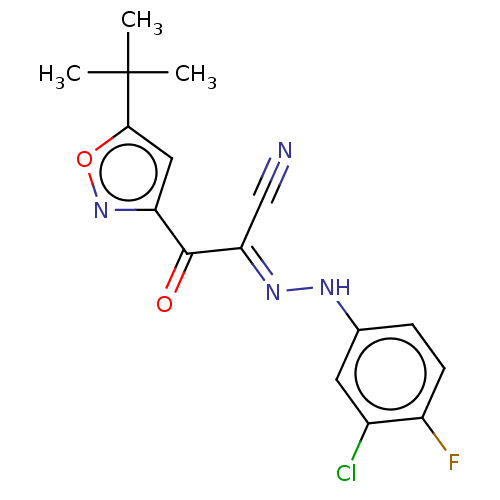
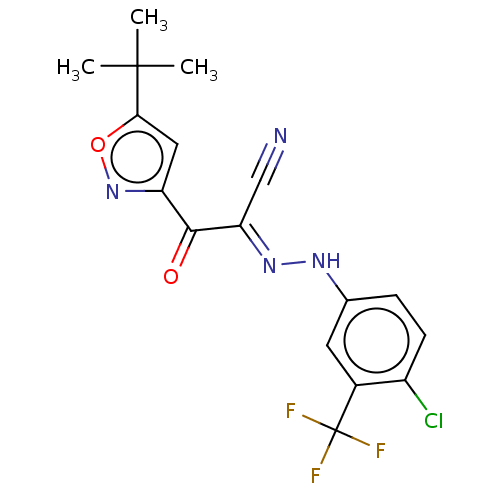
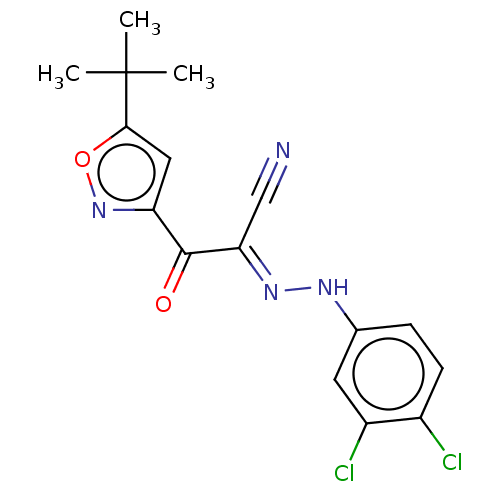
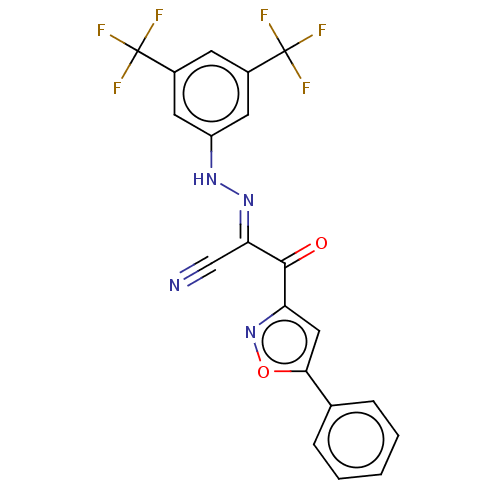
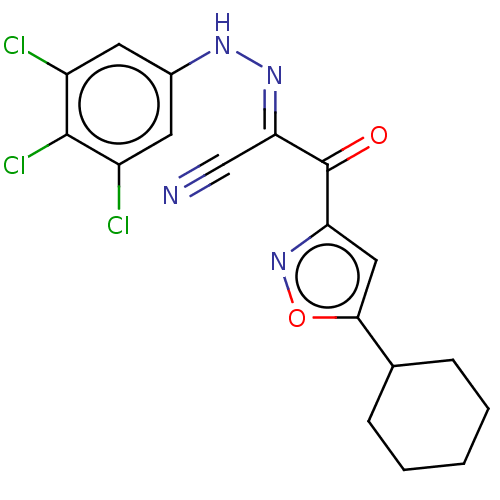
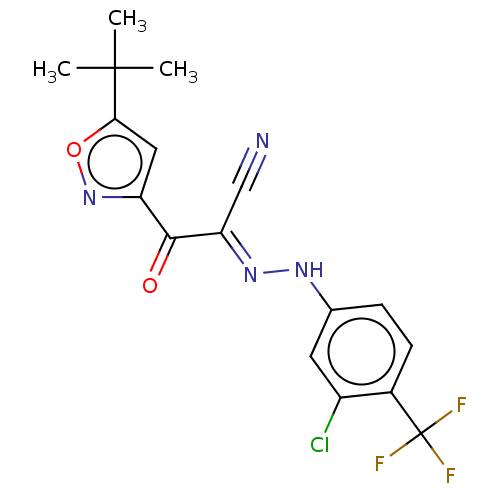
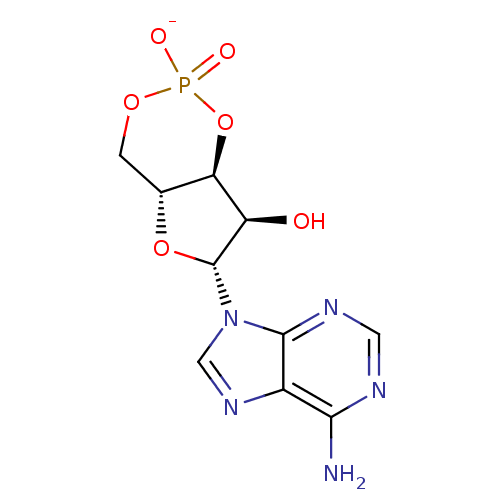
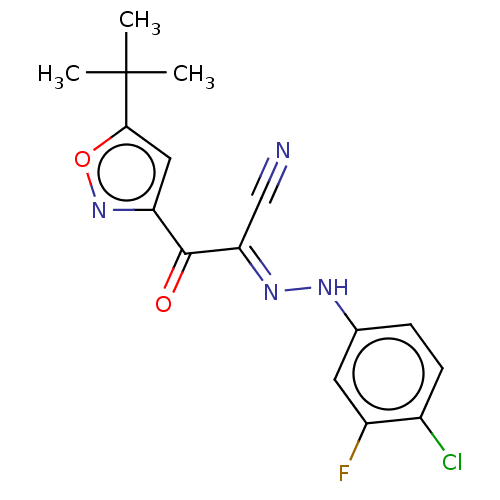
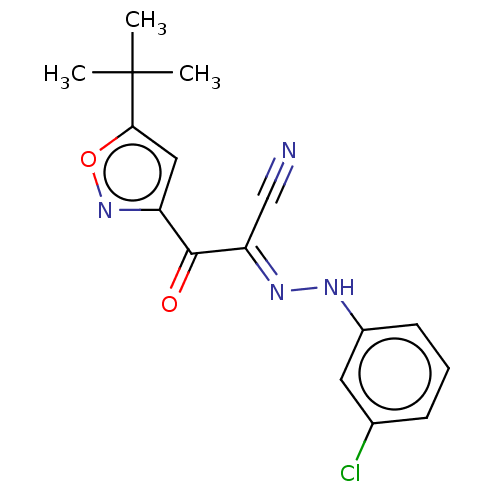
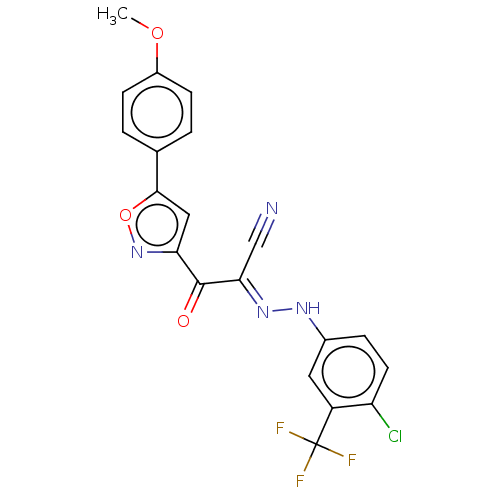
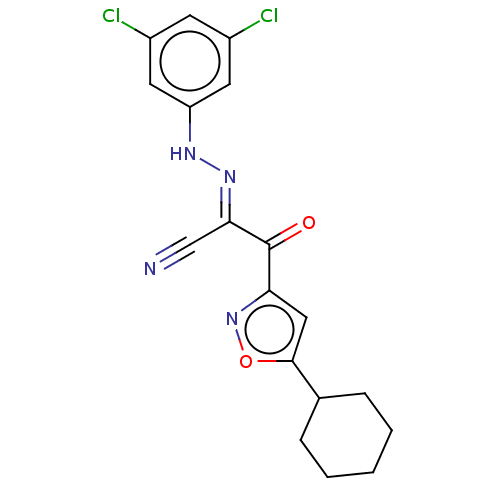
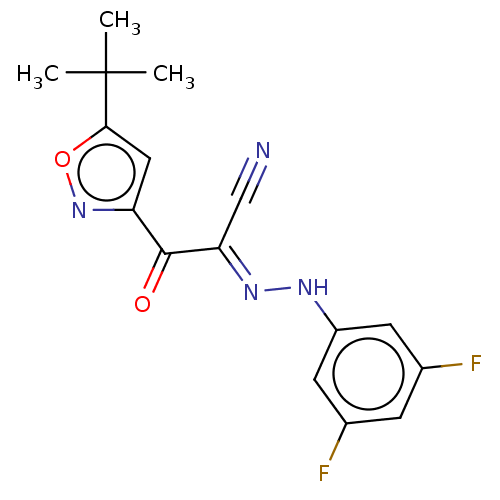
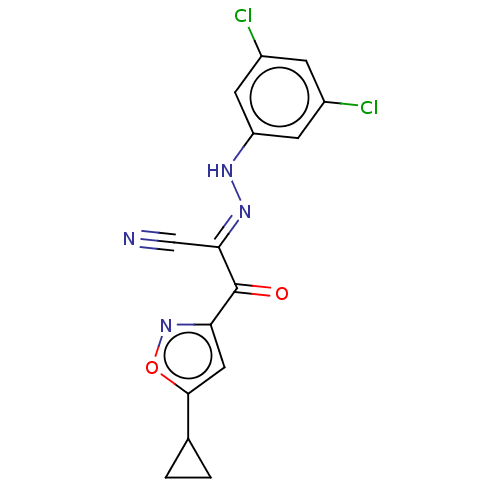
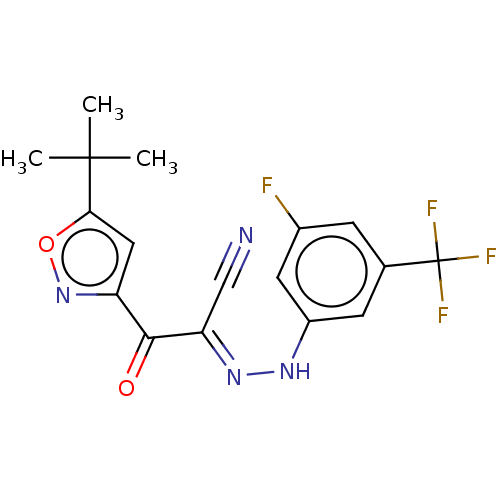
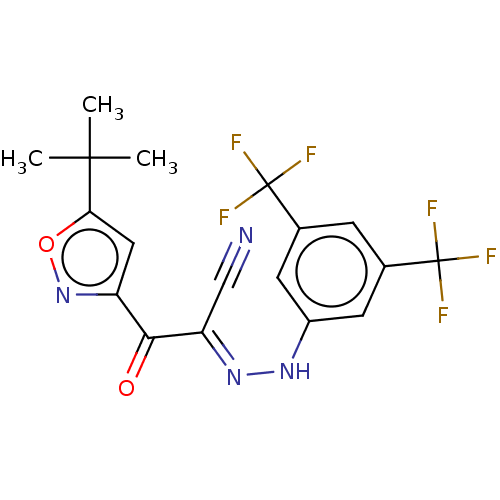
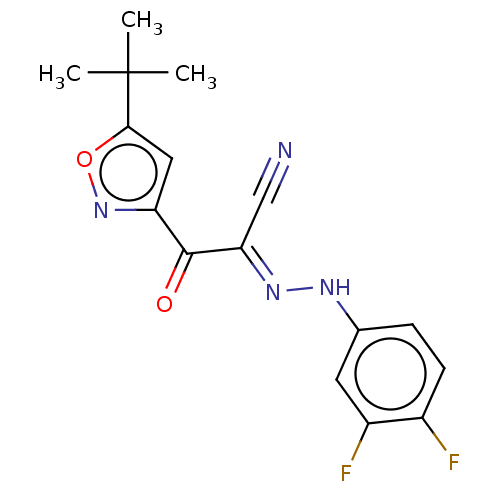
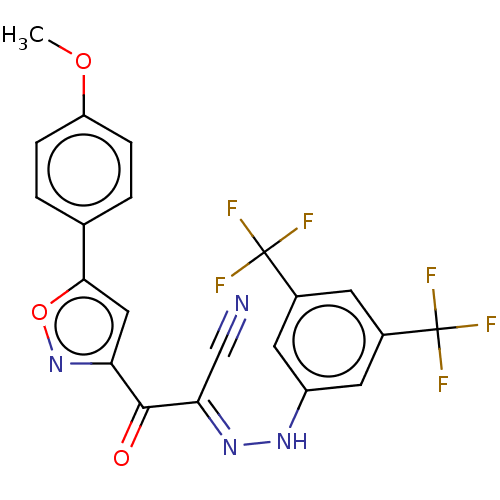
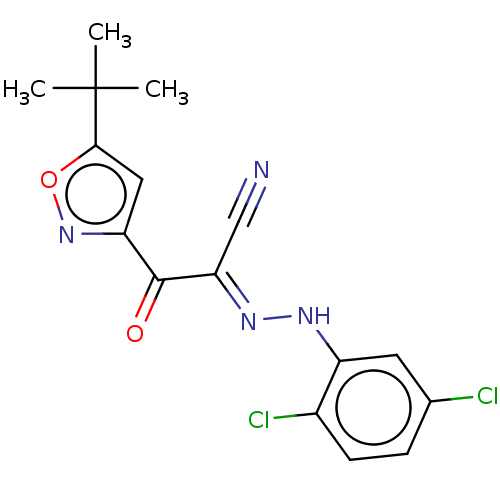
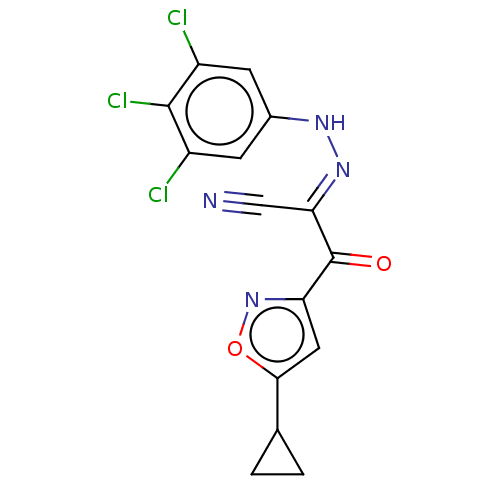
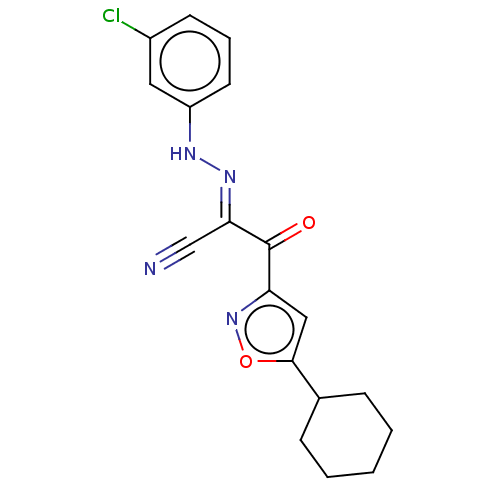
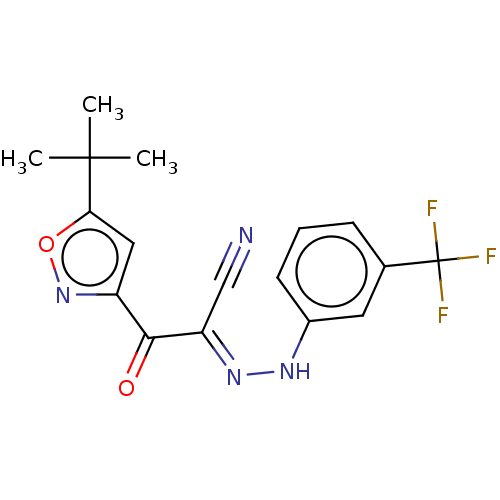
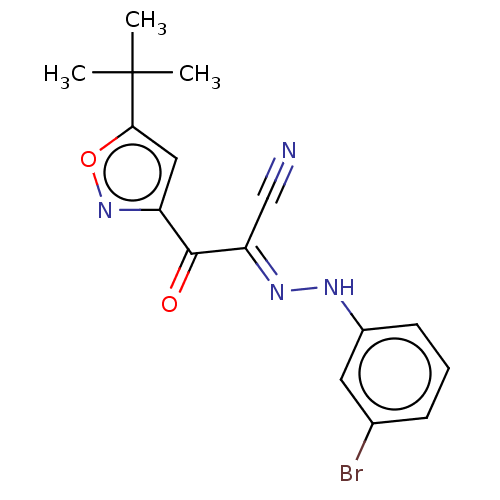
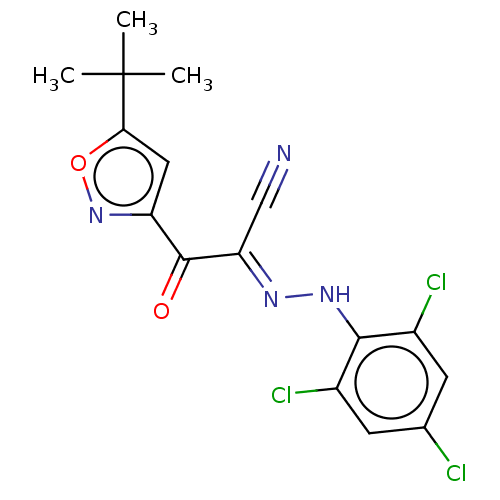
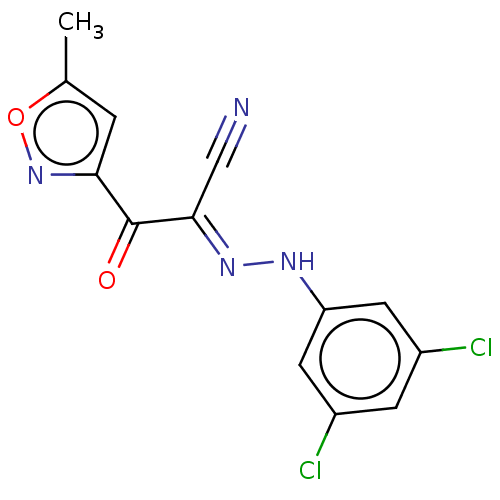
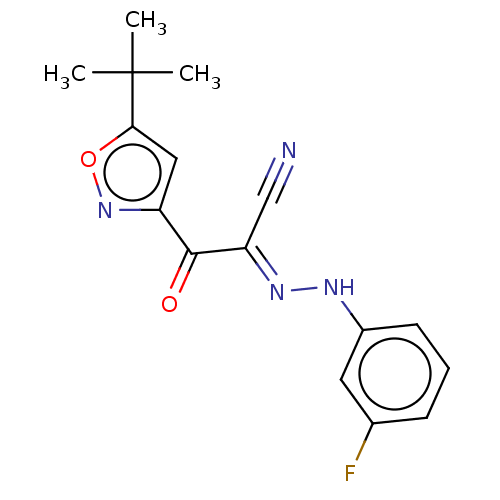
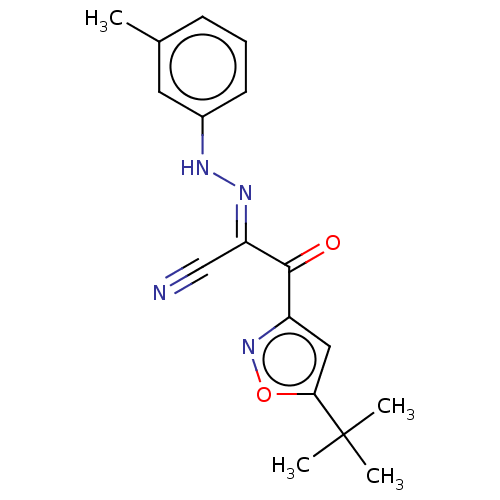
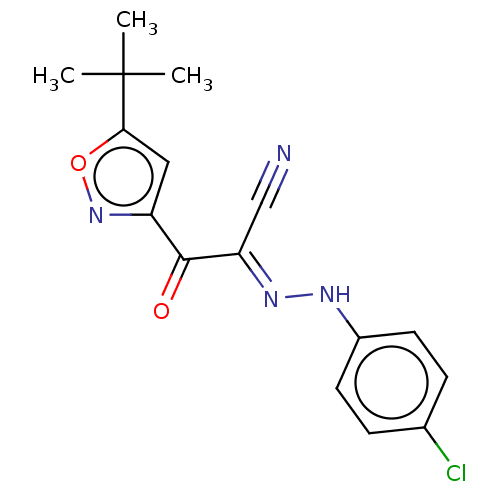
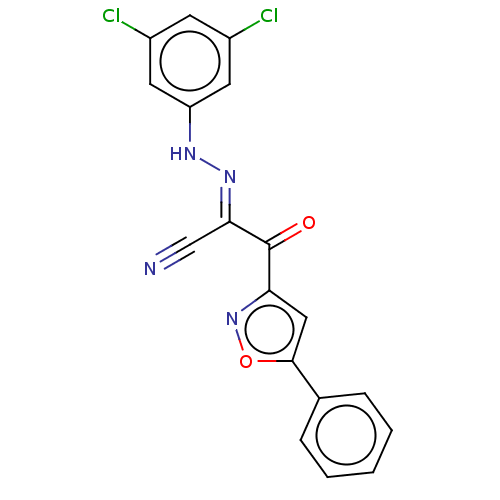
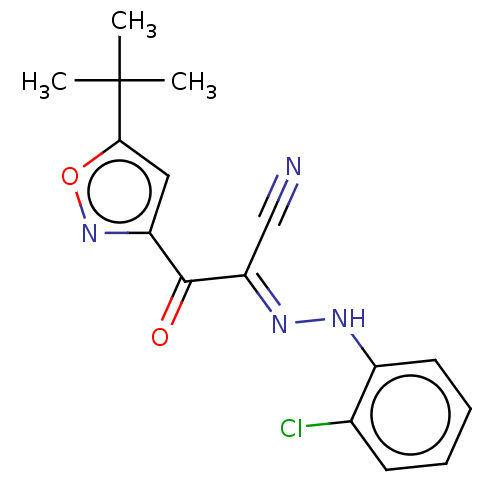
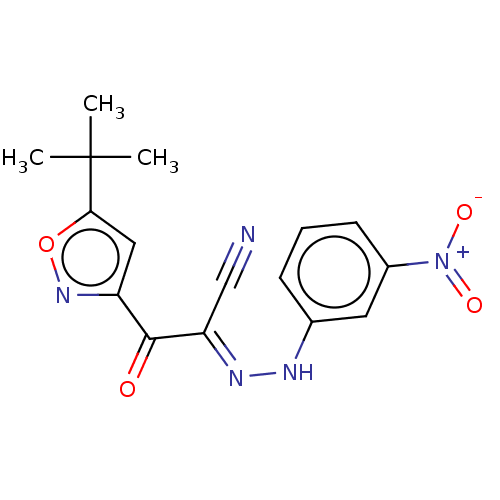
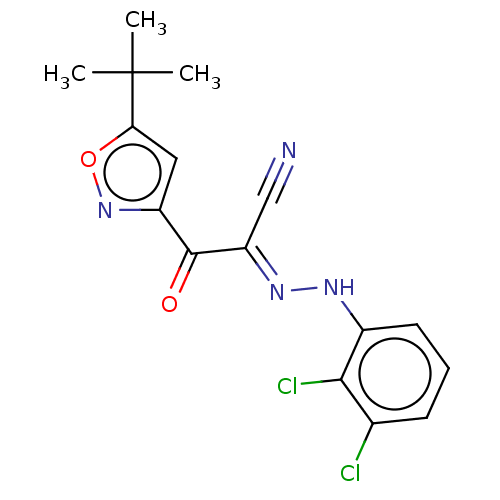
Affinity DataEC50: 300nMAssay Description:Inhibition of 8Fluo-cAMP binding to DEP domain deficient mouse EPAC2 (280 to 988 residues) measured after 5 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataEC50: 780nMAssay Description:Inhibition of 8Fluo-cAMP binding to DEP domain deficient mouse EPAC2 (280 to 988 residues) measured after 5 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 900nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 900nMAssay Description:Displacement of 8-NBD-cAMP from mouse EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMAssay Description:Antagonist activity at mouse EPAC2 assessed as inhibition of EPAC2-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide exchange activity by 8-NB...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataEC50: 2.06E+3nMAssay Description:Inhibition of 8Fluo-cAMP binding to DEP domain deficient mouse EPAC2 (280 to 988 residues) measured after 5 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Antagonist activity at mouse EPAC2 assessed as inhibition of EPAC2-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide exchange activity by 8-NB...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Antagonist activity at mouse EPAC2 assessed as inhibition of EPAC2-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide exchange activity by 8-NB...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Antagonist activity at mouse EPAC2 assessed as inhibition of EPAC2-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide exchange activity by 8-NB...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Antagonist activity at mouse EPAC2 assessed as inhibition of EPAC2-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide exchange activity by 8-NB...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+3nMAssay Description:Antagonist activity at mouse EPAC2 assessed as inhibition of EPAC2-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide exchange activity by 8-NB...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+3nMAssay Description:Antagonist activity at mouse EPAC2 assessed as inhibition of EPAC2-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide exchange activity by 8-NB...More data for this Ligand-Target Pair
Affinity DataEC50: 2.36E+3nMAssay Description:Inhibition of 8Fluo-cAMP binding to DEP domain deficient mouse EPAC2 (280 to 988 residues) measured after 5 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMAssay Description:Antagonist activity at mouse EPAC2 assessed as inhibition of EPAC2-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide exchange activity by 8-NB...More data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataEC50: 3.87E+3nMAssay Description:Inhibition of 8Fluo-cAMP binding to DEP domain deficient mouse EPAC2 (280 to 988 residues) measured after 5 mins by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.40E+3nMAssay Description:Antagonist activity at mouse EPAC2 assessed as inhibition of EPAC2-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide exchange activity by 8-NB...More data for this Ligand-Target Pair
Affinity DataIC50: 4.50E+3nMAssay Description:Antagonist activity at mouse EPAC2 assessed as inhibition of EPAC2-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide exchange activity by 8-NB...More data for this Ligand-Target Pair
Affinity DataIC50: 4.80E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.50E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.40E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.60E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+3nMAssay Description:Antagonist activity at mouse EPAC2 assessed as inhibition of EPAC2-mediated Rap1B (1 to 167 residues)-BODIPY GDP nucleotide exchange activity by 8-NB...More data for this Ligand-Target Pair
Affinity DataIC50: 7.30E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 8.80E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 8.90E+3nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.13E+4nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.19E+4nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+4nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.23E+4nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.24E+4nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.51E+4nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.65E+4nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+4nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.83E+4nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.13E+4nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.56E+4nMAssay Description:Displacement of 8-NBD-cAMP from mouse recombinant EPAC2 by fluorescence assayMore data for this Ligand-Target Pair