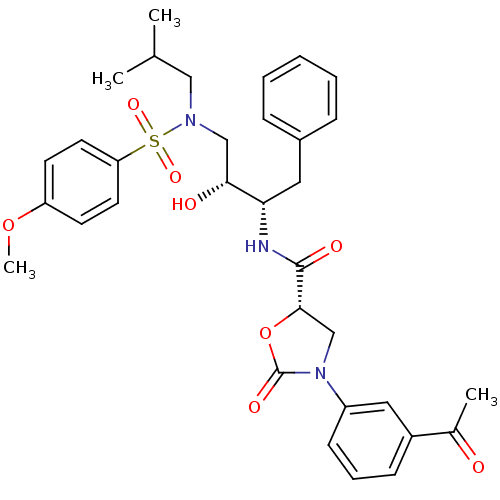
BDBM12877 (5S)-3-(3-Acetylphenyl)-N-[(1S,2R)-1-benzyl-2-hydroxy-3-[isobutyl[(4-methoxyphenyl)sulfonyl]amino]propyl]-2-oxooxazolidine-5-carboxamide::(5S)-3-(3-acetylphenyl)-N-[(2S,3R)-3-hydroxy-4-[(4-methoxybenzene)(2-methylpropyl)sulfonamido]-1-phenylbutan-2-yl]-2-oxo-1,3-oxazolidine-5-carboxamide::N-Aryl-oxazolidinone-5-carboxamide Analogue 21e
SMILES COc1ccc(cc1)S(=O)(=O)N(CC(C)C)C[C@@H](O)[C@H](Cc1ccccc1)NC(=O)[C@@H]1CN(C(=O)O1)c1cccc(c1)C(C)=O
InChI Key InChIKey=SLMZIXVIEWNEAM-UHFFFAOYSA-N
Data 8 KI
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 8 hits for monomerid = 12877
Found 8 hits for monomerid = 12877
Affinity DataKi: 0.000794nMAssay Description:Inhibition of HIV1 protease Q7K mutant by FRET methodMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q496K](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 0.000800nM ΔG°: -16.8kcal/molepH: 4.7 T: 2°CAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetProtease(Human immunodeficiency virus type 1)
University of Massachusetts Medical School
Curated by ChEMBL
University of Massachusetts Medical School
Curated by ChEMBL
Affinity DataKi: 0.000800nMAssay Description:Inhibition of wild type HIV1 protease by FRETMore data for this Ligand-Target Pair
Affinity DataKi: 0.000800nMAssay Description:Inhibition of HIV1 protease Q7K mutant by FRET methodMore data for this Ligand-Target Pair
TargetProtease(Human immunodeficiency virus type 1)
University of Massachusetts Medical School
Curated by ChEMBL
University of Massachusetts Medical School
Curated by ChEMBL
Affinity DataKi: 0.00100nMAssay Description:Inhibition of wild type HIV1 protease by FRET assayMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,D521N,V555I,N579D](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 0.0390nM ΔG°: -14.4kcal/molepH: 4.7 T: 2°CAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,L501I,G539V,I545V,V555I,V573A](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 0.160nM ΔG°: -13.6kcal/molepH: 4.7 T: 2°CAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [491-589,Q498K,L501I,V555I,A562V,G564S,I575V,L581M](Human immunodeficiency virus type 1)
University of Massachusetts Medical School
University of Massachusetts Medical School
Affinity DataKi: 3.36nM ΔG°: -11.7kcal/molepH: 4.7 T: 2°CAssay Description:HIV-1 protease inhibitor activities were determined by the fluorescence resonance energy transfer (FRET) method. The energy transfer donor (EDANS) an...More data for this Ligand-Target Pair