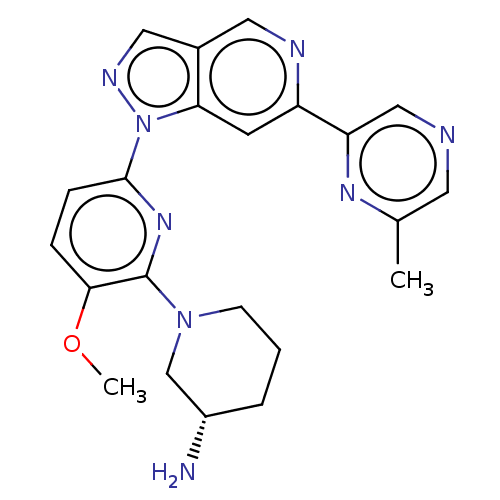
BDBM248955 US9434725, 186
SMILES COc1ccc(nc1N1CCC[C@H](N)C1)-n1ncc2cnc(cc12)-c1cncc(C)n1
InChI Key InChIKey=FPBWUTAWSOLPQI-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 7 hits for monomerid = 248955
Found 7 hits for monomerid = 248955
Affinity DataKi: 0.00800nMAssay Description:Inhibition of PIM3 (unknown origin) using 5FAM-ARKRRRHPSGPPTA as substrate after 90 mins in presence of ATP by caliper microfluidic mobility shift as...More data for this Ligand-Target Pair
Affinity DataKi: 0.0130nM ΔG°: -14.8kcal/molepH: 7.2 T: 2°CAssay Description:PIM-1, -2, and -3 enzymes were generated as fusion proteins expressed in bacteria and purified by IMAC column chromatography (Sun, X., Chiu, J. F., a...More data for this Ligand-Target Pair
Affinity DataKi: 0.0180nMAssay Description:Inhibition of PIM1 (unknown origin) using 5FAM-ARKRRRHPSGPPTA as substrate after 90 mins in presence of ATP by caliper microfluidic mobility shift as...More data for this Ligand-Target Pair
Affinity DataKi: 0.0212nM ΔG°: -14.6kcal/molepH: 7.2 T: 2°CAssay Description:PIM-1, -2, and -3 enzymes were generated as fusion proteins expressed in bacteria and purified by IMAC column chromatography (Sun, X., Chiu, J. F., a...More data for this Ligand-Target Pair
Affinity DataKi: 0.110nMAssay Description:Inhibition of PIM2 (unknown origin) using 5FAM-ARKRRRHPSGPPTA as substrate after 90 mins in presence of ATP by caliper microfluidic mobility shift as...More data for this Ligand-Target Pair
Affinity DataKi: 0.134nM ΔG°: -13.5kcal/molepH: 7.2 T: 2°CAssay Description:PIM-1, -2, and -3 enzymes were generated as fusion proteins expressed in bacteria and purified by IMAC column chromatography (Sun, X., Chiu, J. F., a...More data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:BaF3, a murine interleukin-3 dependent pro-B cell line, parental cells, BaF3 PIM1 cells, BaF3 PIM2 cells, and MM1.S (multiple myeloma) cells were see...More data for this Ligand-Target Pair