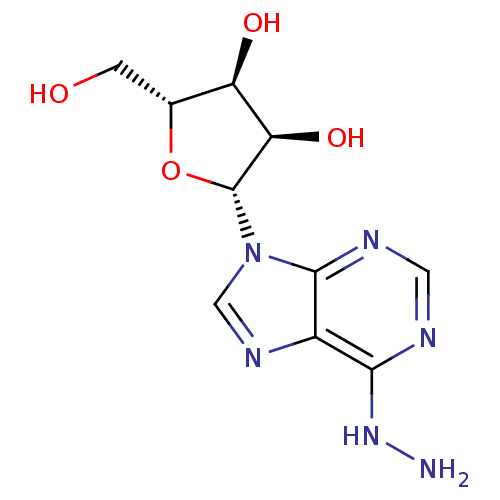
BDBM50214817 CHEMBL236757::N6-aminoadenosine
SMILES NNc1ncnc2n(cnc12)[C@@H]1O[C@H](CO)[C@@H](O)[C@H]1O
InChI Key InChIKey=DGKZTAGCCXJUAT-KQYNXXCUSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 11 hits for monomerid = 50214817
Found 11 hits for monomerid = 50214817
TargetHeat shock protein HSP 90-alpha(Homo sapiens (Human))
National Institute Of Diabetes And Digestive And Kidney Diseases
Curated by ChEMBL
National Institute Of Diabetes And Digestive And Kidney Diseases
Curated by ChEMBL
Affinity DataKi: >3.00E+3nMAssay Description:Displacement of FITC-geldanamycin from HSP90alpha (unknown origin) after 24 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetAdenosine receptor A2a(Rattus norvegicus (rat))
National Institute Of Diabetes
Curated by ChEMBL
National Institute Of Diabetes
Curated by ChEMBL
Affinity DataKi: 7.34E+3nMAssay Description:Binding affinity to adenosine A2A receptor in rat striatal membranes by measuring displacement of specific [3H]-CGS- 21680 as radioligandMore data for this Ligand-Target Pair
TargetAdenosine receptor A2a(Rattus norvegicus (rat))
National Institute Of Diabetes
Curated by ChEMBL
National Institute Of Diabetes
Curated by ChEMBL
Affinity DataKi: 7.34E+3nMAssay Description:Displacement of [3H]-CGS- 21680 from Adenosine A2 receptor of rat striatal membranesMore data for this Ligand-Target Pair
TargetEndoplasmin(Homo sapiens (Human))
National Institute Of Diabetes And Digestive And Kidney Diseases
Curated by ChEMBL
National Institute Of Diabetes And Digestive And Kidney Diseases
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of FITC-geldanamycin from GRP94 (unknown origin) after 24 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.69E+4nMAssay Description:Binding affinity determined by displacement of specific binding of [125I]N-(4-amino-3-iodophenethyl)-adenosine in membranes of CHO cells stably trans...More data for this Ligand-Target Pair
Affinity DataKi: 2.97E+4nMAssay Description:Binding affinity at Adenosine A1 receptor in rat brain membranes by [3H](R)-PIA displacement.More data for this Ligand-Target Pair
Affinity DataKi: 2.97E+4nMAssay Description:Binding affinity to adenosine A1 receptor in rat brain membranes by measuring displacement of specific [3H]PIA as radioligand.More data for this Ligand-Target Pair
TargetHeat shock protein HSP 90-alpha(Homo sapiens (Human))
National Institute Of Diabetes And Digestive And Kidney Diseases
Curated by ChEMBL
National Institute Of Diabetes And Digestive And Kidney Diseases
Curated by ChEMBL
Affinity DataKd: >5.00E+4nMAssay Description:Displacement of FITC-geldanamycin from HSP90alpha (unknown origin) after 24 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetEndoplasmin(Homo sapiens (Human))
National Institute Of Diabetes And Digestive And Kidney Diseases
Curated by ChEMBL
National Institute Of Diabetes And Digestive And Kidney Diseases
Curated by ChEMBL
Affinity DataKd: >5.00E+4nMAssay Description:Displacement of FITC-geldanamycin from GRP94 (unknown origin) after 24 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetRibonucleoside-diphosphate reductase large subunit/subunit M2 B(Homo sapiens (Human))
University Of Camerino
Curated by ChEMBL
University Of Camerino
Curated by ChEMBL
Affinity DataIC50: 2.44E+5nMAssay Description:Inhibition of human recombinant ribonucleotide reductase p53R2/M1 expressed in Escherichia coli BL21 (DE3) after 30 mins by [3H]CDP reduction methodMore data for this Ligand-Target Pair
TargetRibonucleoside-diphosphate reductase large subunit/subunit M2(Homo sapiens (Human))
University Of Camerino
Curated by ChEMBL
University Of Camerino
Curated by ChEMBL
Affinity DataIC50: 2.32E+5nMAssay Description:Inhibition of human recombinant ribonucleotide reductase M2/M1 expressed in Escherichia coli BL21 (DE3) after 30 mins by [3H]CDP reduction methodMore data for this Ligand-Target Pair