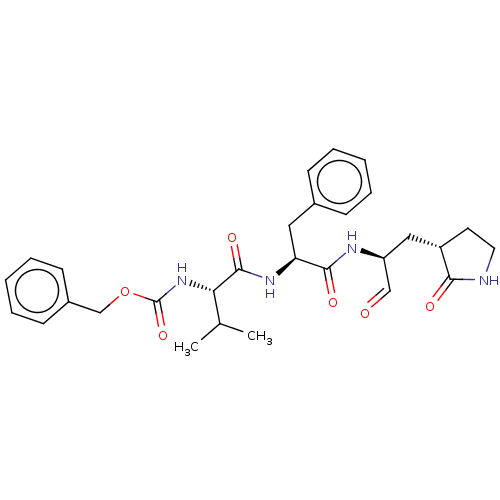
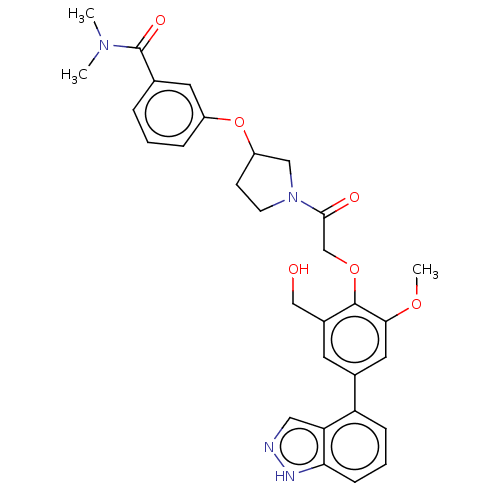
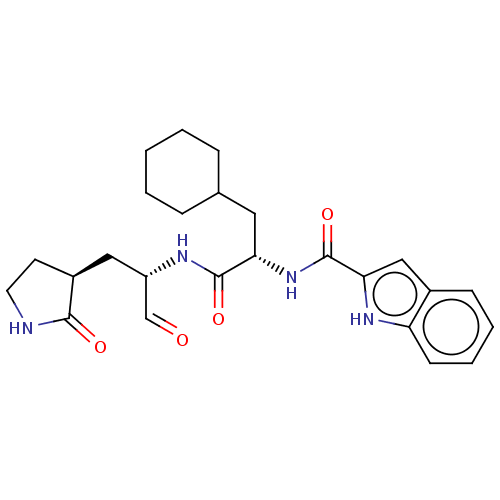
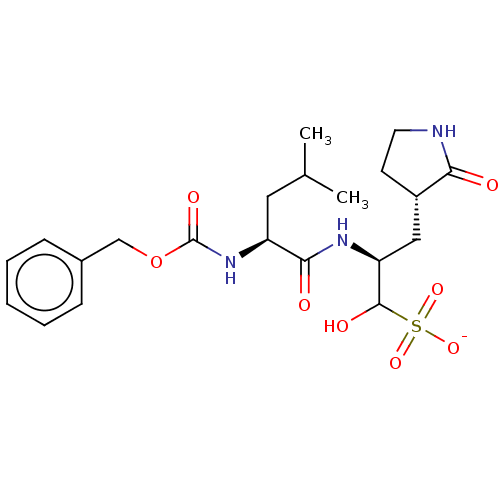
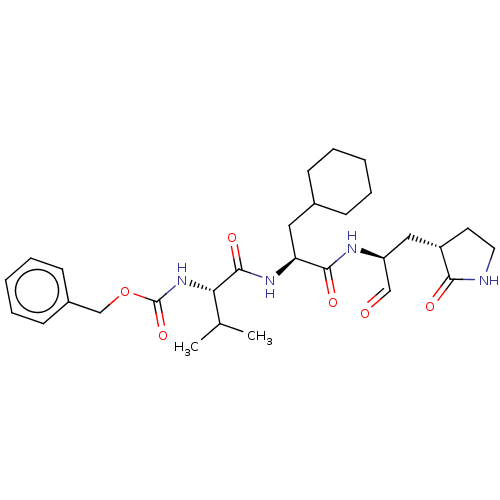
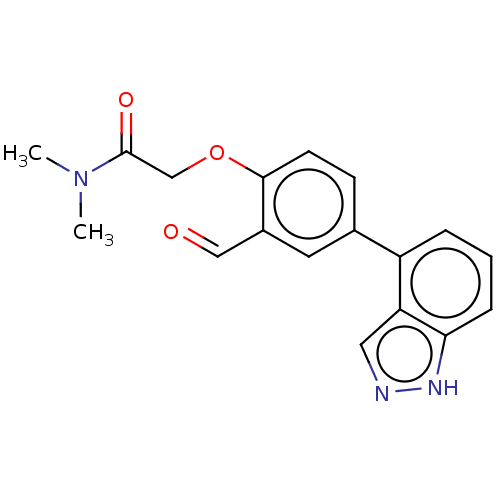
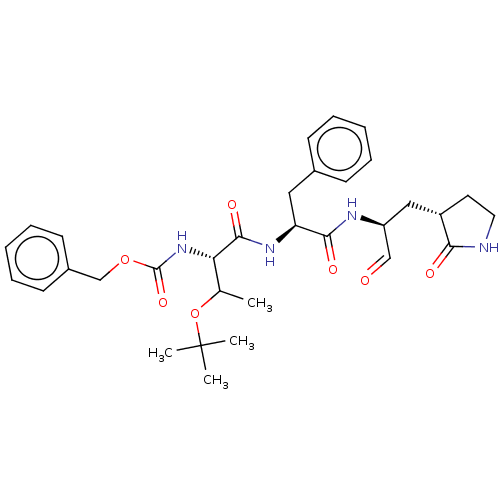
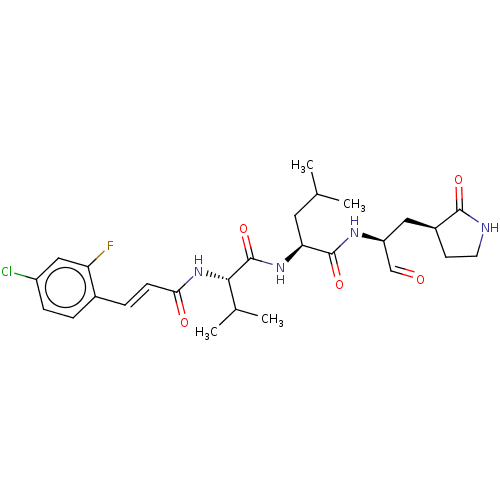
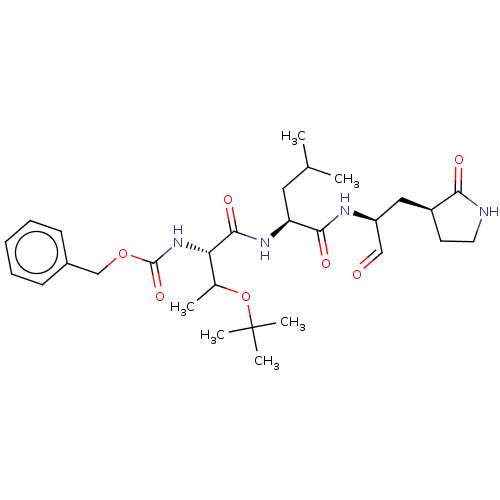
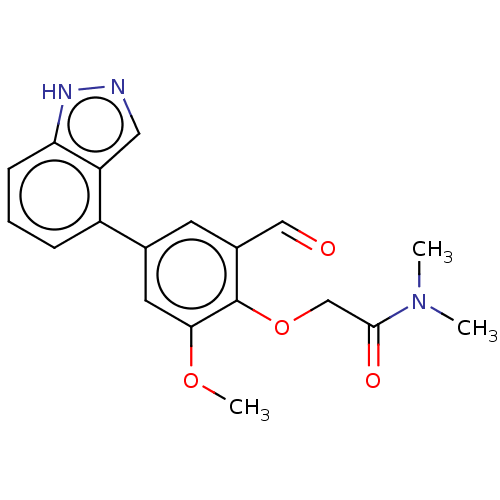
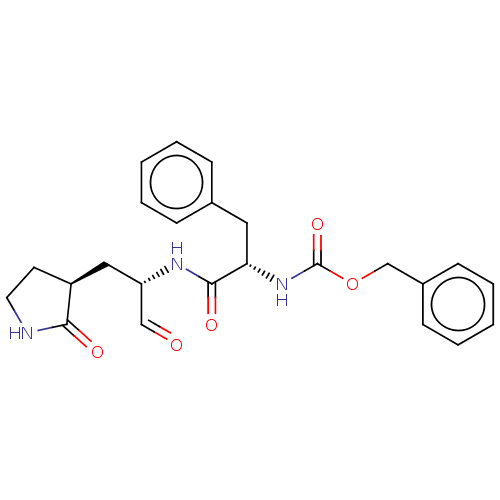
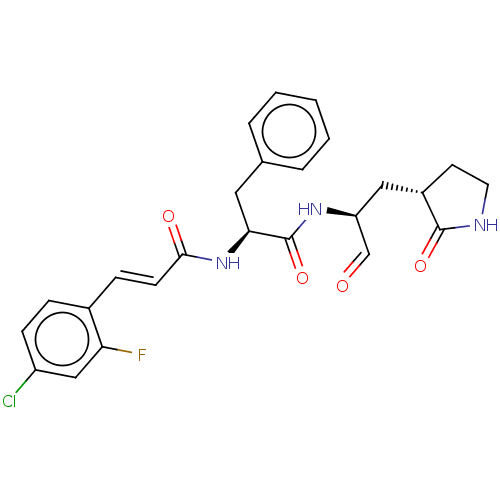
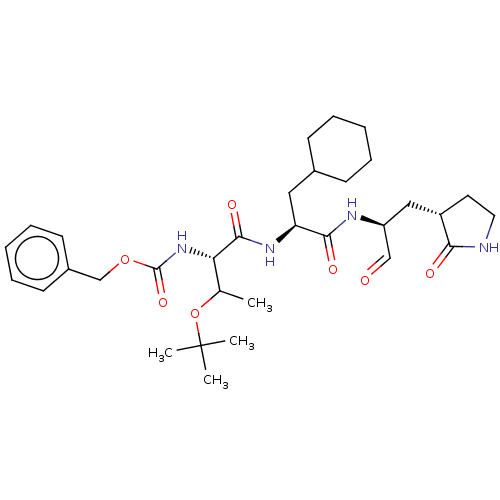
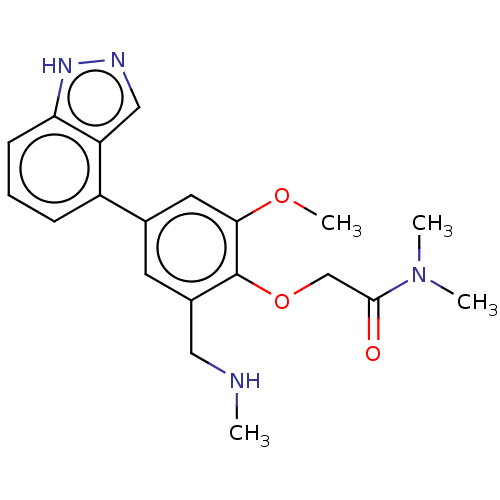
Affinity DataKi: 0.0400nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 0.110nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 0.130nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 0.170nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 0.230nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 0.240nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 0.270nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 0.310nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 0.490nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 0.580nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 0.600nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of N-terminal 6-histine tagged Bacillus anthracis LF 263-C terminal catalytic domain using MCA-KKVYPYPME-Dap(Dnp)-NH2 as substrate after 4...More data for this Ligand-Target Pair
Affinity DataKi: 8.30nM IC50: 8.5nMAssay Description:The assays were carried out with 20 nM enzyme (except for MPI3, for which 10 nM enzyme was used) and 10 μM substrate at 37oC with continuous sha...More data for this Ligand-Target Pair
Affinity DataKi: 15nM IC50: 15nMAssay Description:The assays were carried out with 20 nM enzyme (except for MPI3, for which 10 nM enzyme was used) and 10 μM substrate at 37oC with continuous sha...More data for this Ligand-Target Pair
Affinity DataKi: 24nMAssay Description:To determine the Ki values of active compounds, 25 nM Mpro was mixed with increasing concentrations of compounds (from 4 nM to 4,000 nM with twofold ...More data for this Ligand-Target Pair
Affinity DataKi: 30nM IC50: 31nMAssay Description:The assays were carried out with 20 nM enzyme (except for MPI3, for which 10 nM enzyme was used) and 10 μM substrate at 37oC with continuous sha...More data for this Ligand-Target Pair
Affinity DataKi: 30nM IC50: 31nMAssay Description:The assays were carried out with 20 nM enzyme (except for MPI3, for which 10 nM enzyme was used) and 10 μM substrate at 37oC with continuous sha...More data for this Ligand-Target Pair
Affinity DataKi: 32nM IC50: 33nMAssay Description:The assays were carried out with 20 nM enzyme (except for MPI3, for which 10 nM enzyme was used) and 10 μM substrate at 37oC with continuous sha...More data for this Ligand-Target Pair
Affinity DataKi: 42nMAssay Description:To determine the Ki values of active compounds, 25 nM Mpro was mixed with increasing concentrations of compounds (from 4 nM to 4,000 nM with twofold ...More data for this Ligand-Target Pair
Affinity DataKi: 46nM IC50: 47nMAssay Description:The assays were carried out with 20 nM enzyme (except for MPI3, for which 10 nM enzyme was used) and 10 μM substrate at 37oC with continuous sha...More data for this Ligand-Target Pair
Affinity DataKi: 55nM IC50: 56nMAssay Description:The assays were carried out with 20 nM enzyme (except for MPI3, for which 10 nM enzyme was used) and 10 μM substrate at 37oC with continuous sha...More data for this Ligand-Target Pair
Affinity DataKi: 59nM IC50: 60nMAssay Description:The assays were carried out with 20 nM enzyme (except for MPI3, for which 10 nM enzyme was used) and 10 μM substrate at 37oC with continuous sha...More data for this Ligand-Target Pair
Affinity DataKi: 65nMAssay Description:To determine the Ki values of active compounds, 25 nM Mpro was mixed with increasing concentrations of compounds (from 4 nM to 4,000 nM with twofold ...More data for this Ligand-Target Pair
Affinity DataKi: 98nM IC50: 100nMAssay Description:The assays were carried out with 20 nM enzyme (except for MPI3, for which 10 nM enzyme was used) and 10 μM substrate at 37oC with continuous sha...More data for this Ligand-Target Pair
Affinity DataKi: 101nM IC50: 103nMAssay Description:The assays were carried out with 20 nM enzyme (except for MPI3, for which 10 nM enzyme was used) and 10 μM substrate at 37oC with continuous sha...More data for this Ligand-Target Pair
Affinity DataKi: 103nM IC50: 105nMAssay Description:The assays were carried out with 20 nM enzyme (except for MPI3, for which 10 nM enzyme was used) and 10 μM substrate at 37oC with continuous sha...More data for this Ligand-Target Pair
Affinity DataKi: 2.30E+3nMAssay Description:Inhibition of MMP-12 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: 2.40E+3nMAssay Description:To determine the Ki values of active compounds, 25 nM Mpro was mixed with increasing concentrations of compounds (from 4 nM to 4,000 nM with twofold ...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+4nMAssay Description:Inhibition of MMP-9 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: 1.20E+4nMAssay Description:Inhibition of MMP-12 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: 2.10E+4nMAssay Description:Inhibition of MMP-9 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: 3.10E+4nMAssay Description:Inhibition of MMP-3 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: 3.30E+4nMAssay Description:Inhibition of MMP-1 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: 4.70E+4nMAssay Description:Inhibition of MMP-1 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:Inhibition of MMP-12 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:Inhibition of MMP-12 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:Inhibition of MMP-12 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:Inhibition of MMP-12 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:Inhibition of MMP-12 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:Inhibition of MMP-9 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:Inhibition of MMP-9 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:Inhibition of MMP-9 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:Inhibition of MMP-9 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:Inhibition of MMP-9 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:Inhibition of MMP-3 using OmniMMP as substrateMore data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:Inhibition of MMP-3 using OmniMMP as substrateMore data for this Ligand-Target Pair


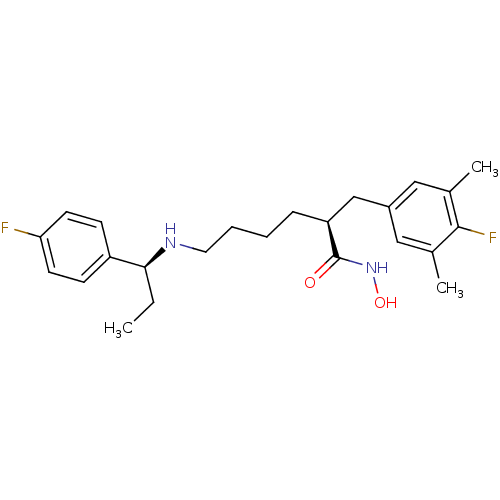
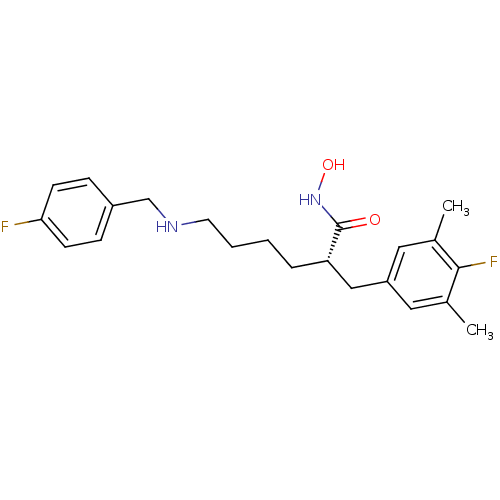
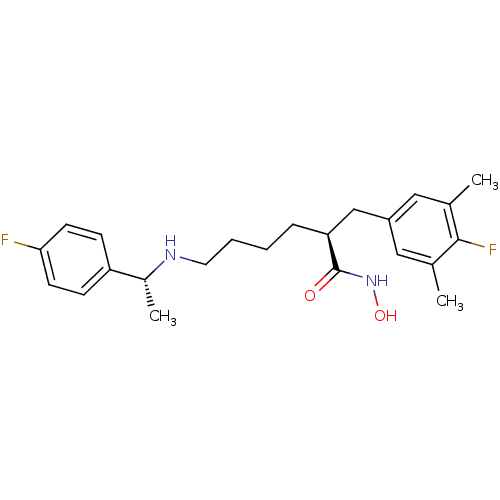
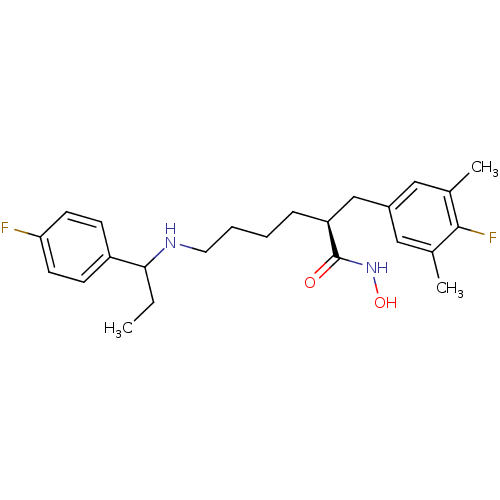

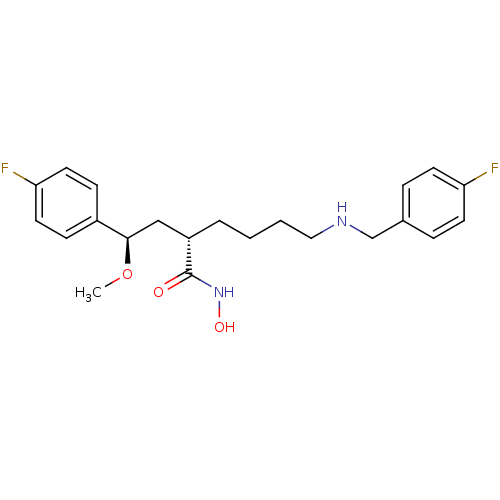
 3D Structure (crystal)
3D Structure (crystal)