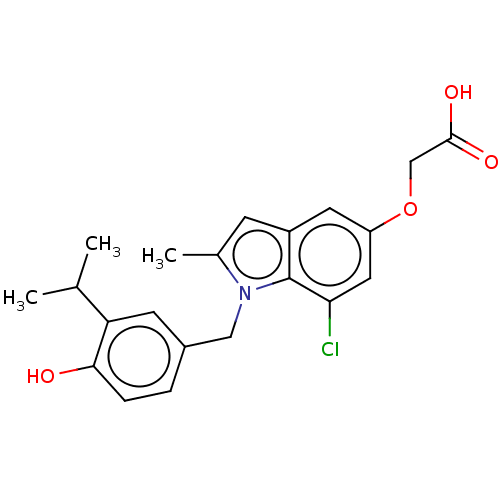
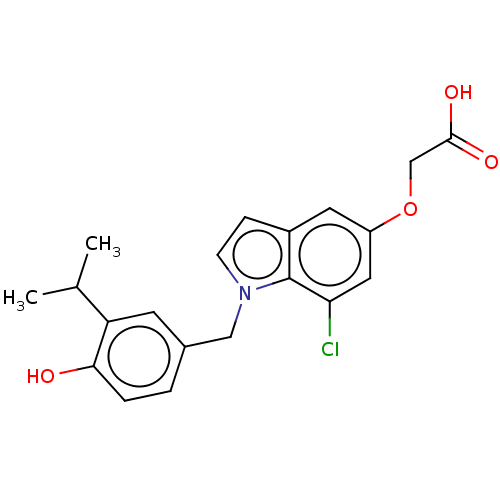
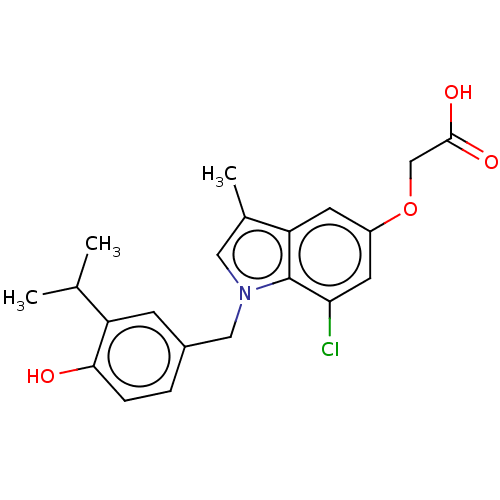
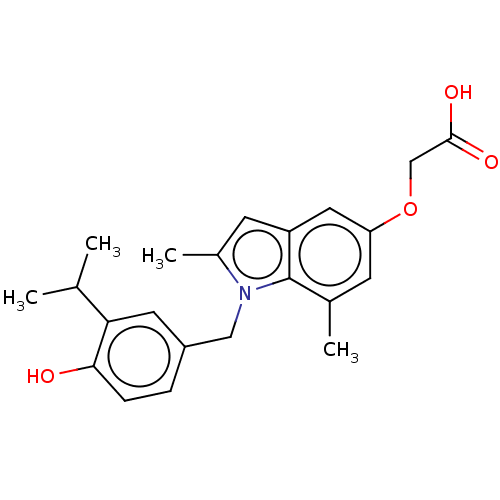
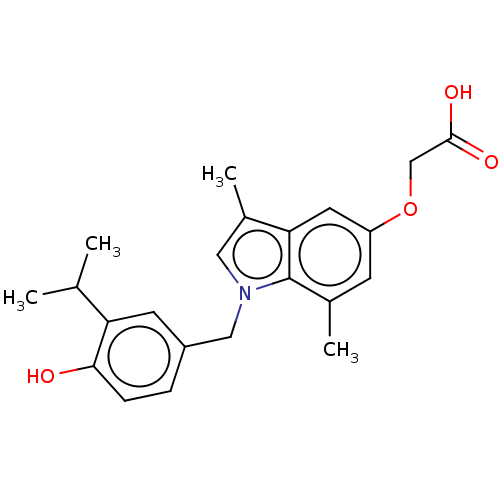
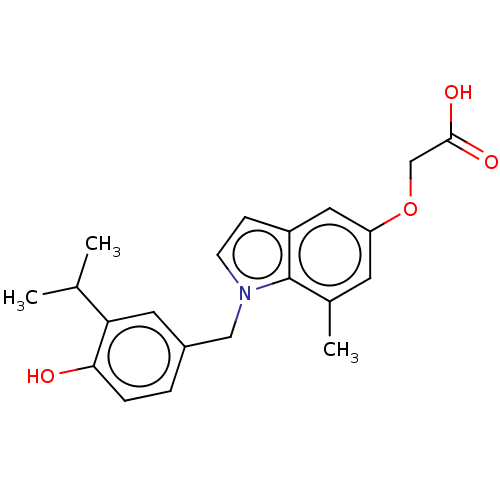
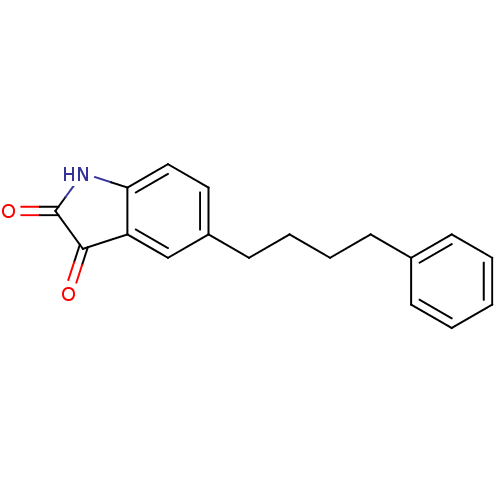
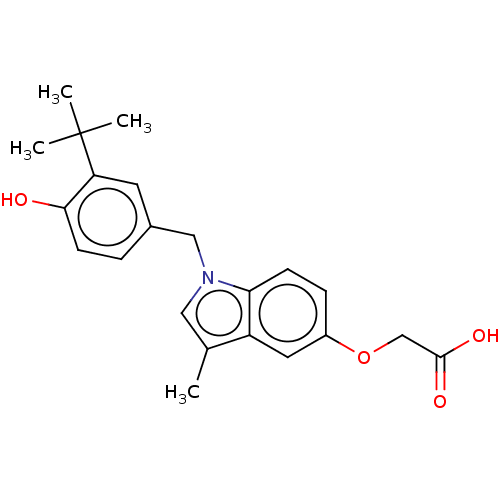
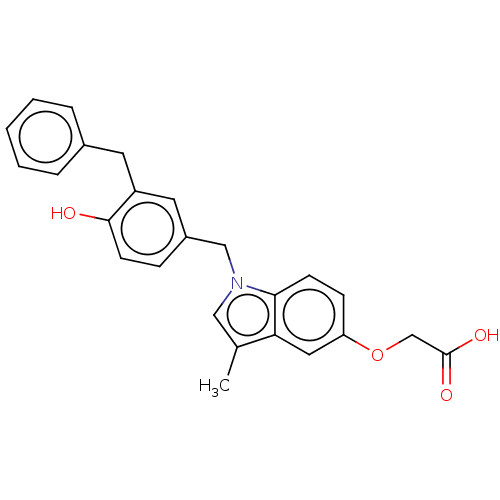
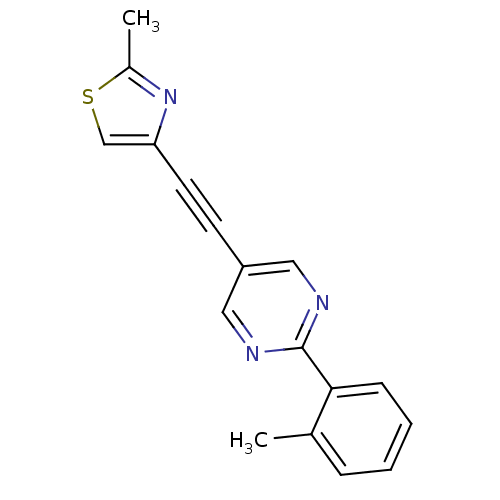
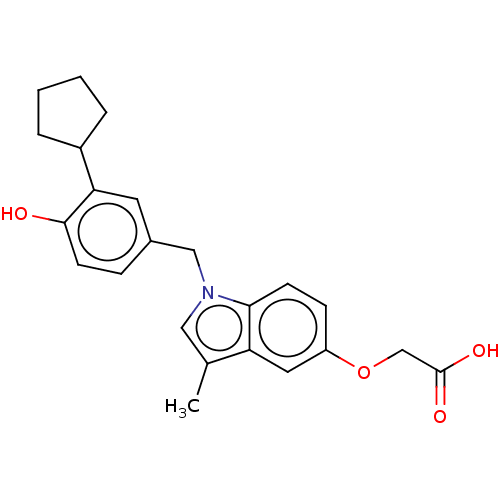
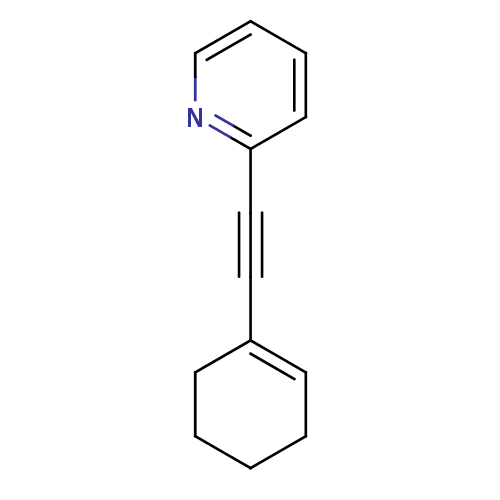
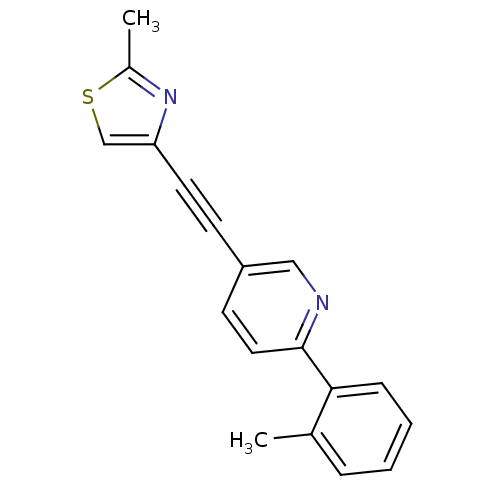
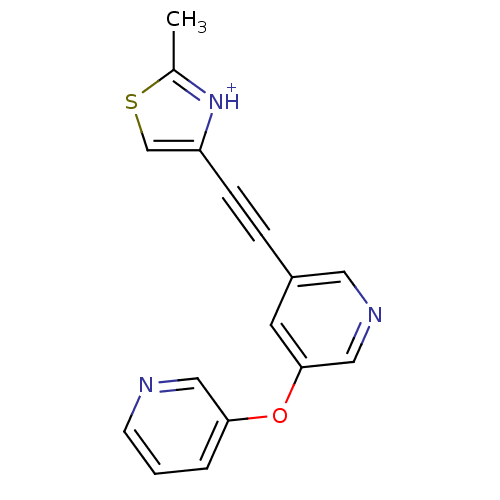
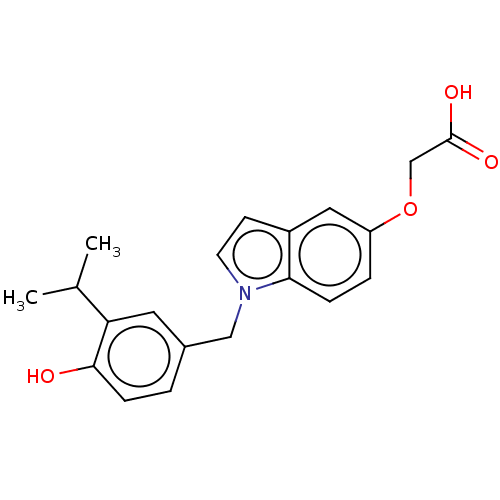
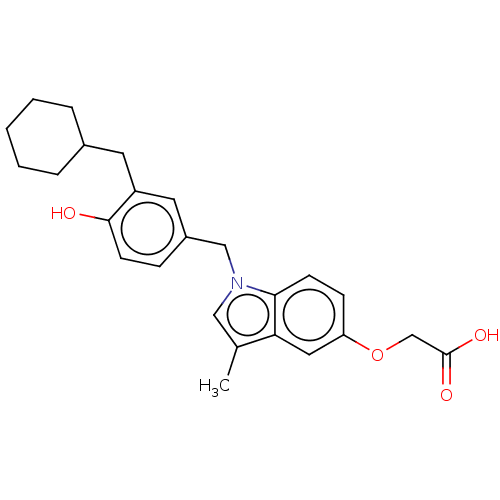
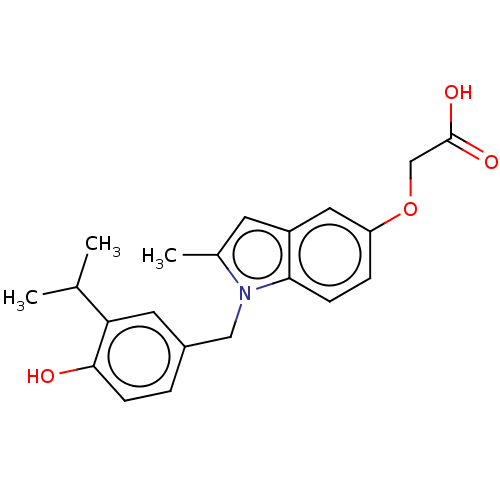
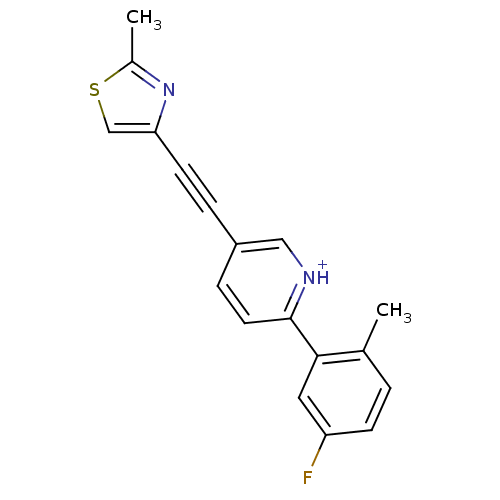
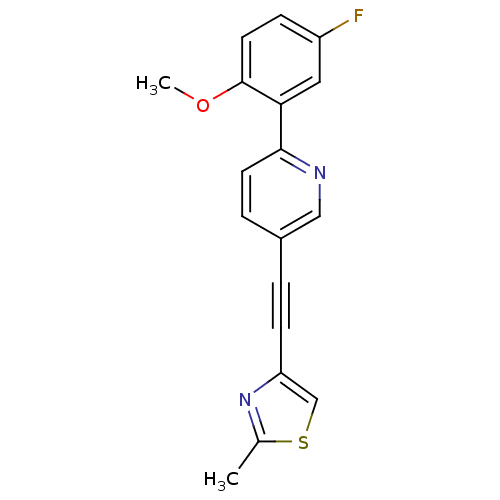
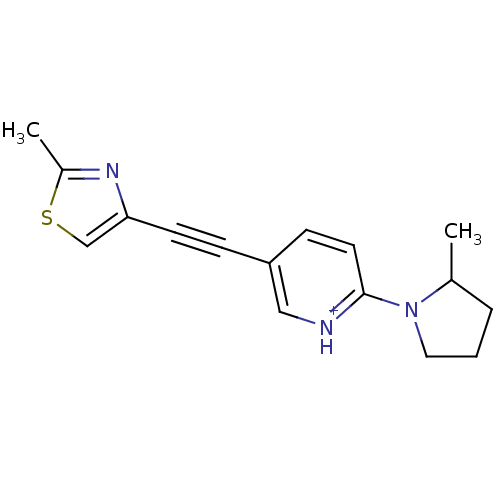
Affinity DataKi: 0.0400nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRbeta expressed in human Hela cell lysate measured after overnight incubation by competi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0700nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRbeta expressed in human Hela cell lysate measured after overnight incubation by competi...More data for this Ligand-Target Pair
Affinity DataKi: 0.0900nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRbeta expressed in human Hela cell lysate measured after overnight incubation by competi...More data for this Ligand-Target Pair
Affinity DataKi: 0.140nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRbeta expressed in human Hela cell lysate measured after overnight incubation by competi...More data for this Ligand-Target Pair
Affinity DataKi: 0.180nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRbeta expressed in human Hela cell lysate measured after overnight incubation by competi...More data for this Ligand-Target Pair
Affinity DataKi: 0.260nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRbeta expressed in human Hela cell lysate measured after overnight incubation by competi...More data for this Ligand-Target Pair
TargetAmine oxidase [flavin-containing] B(Homo sapiens (Human))
North-West University
Curated by ChEMBL
North-West University
Curated by ChEMBL
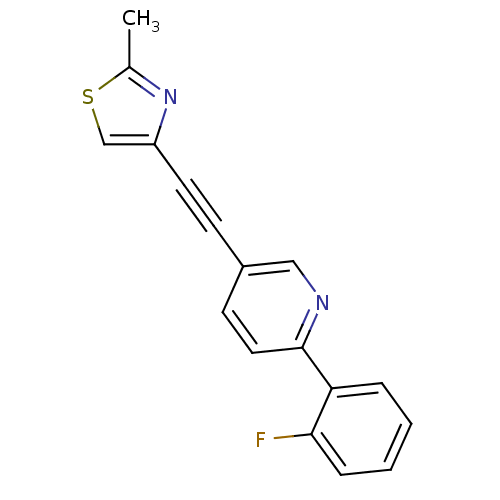
Affinity DataKi: 0.280nMAssay Description:Inhibition of human recombinant MAOB expressed in insect cells by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRbeta expressed in human Hela cell lysate measured after overnight incubation by competi...More data for this Ligand-Target Pair
Affinity DataKi: 0.350nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRalpha expressed in human Hela cell lysate measured after overnight incubation by compet...More data for this Ligand-Target Pair
Affinity DataKi: 0.380nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRbeta expressed in human Hela cell lysate measured after overnight incubation by competi...More data for this Ligand-Target Pair
Affinity DataKi: 0.380nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRalpha expressed in human Hela cell lysate measured after overnight incubation by compet...More data for this Ligand-Target Pair
Affinity DataKi: 0.510nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRalpha expressed in human Hela cell lysate measured after overnight incubation by compet...More data for this Ligand-Target Pair
Affinity DataKi: 0.540nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRbeta expressed in human Hela cell lysate measured after overnight incubation by competi...More data for this Ligand-Target Pair
Affinity DataKi: 0.600nM ΔG°: -52.6kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 0.640nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRbeta expressed in human Hela cell lysate measured after overnight incubation by competi...More data for this Ligand-Target Pair
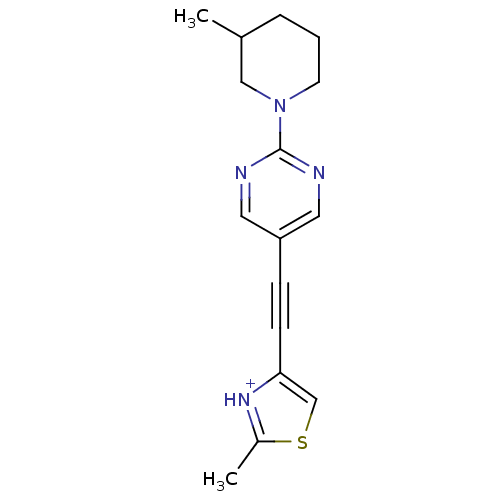
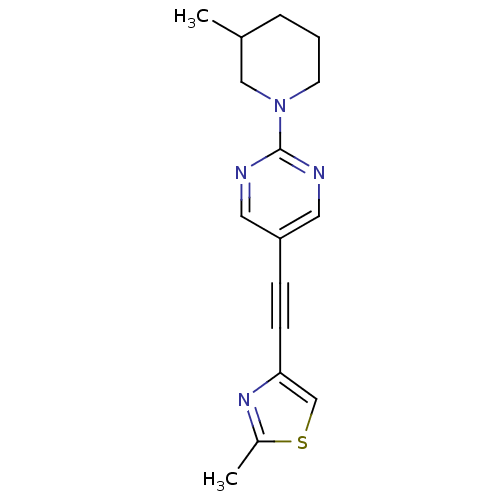
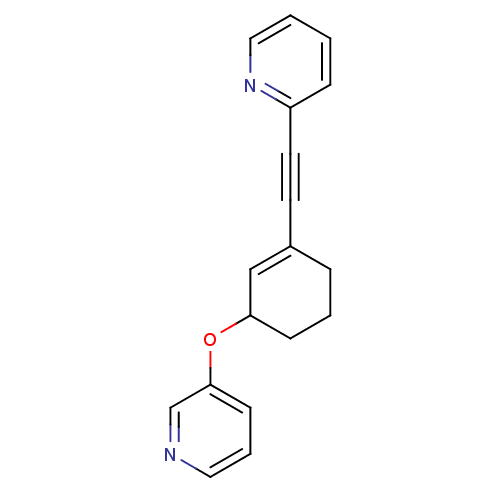
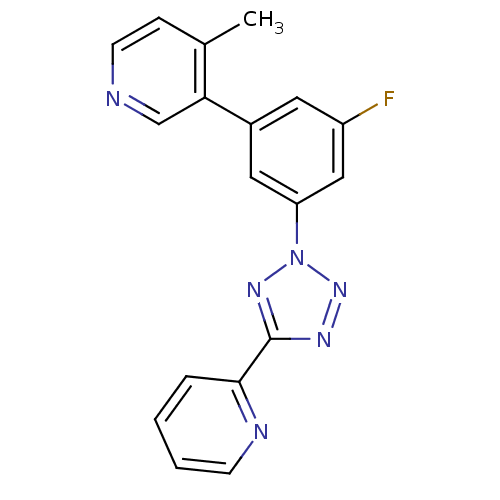
Affinity DataKi: 0.650nMAssay Description:Displacement of [3H]-3-methoxy-5-(pyridin-2-ylethynyl)pyridine from mGlu5 receptor of rat cortical membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.650nM ΔG°: -52.4kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 0.700nM ΔG°: -52.3kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 0.800nM ΔG°: -51.9kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 0.830nMAssay Description:Displacement of [3H]-3-methoxy-5-(pyridin-2-ylethynyl)pyridine from mGlu5 receptor of rat cortical membraneMore data for this Ligand-Target Pair
Affinity DataKi: 0.900nM ΔG°: -51.6kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 0.900nM ΔG°: -51.6kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 0.900nM ΔG°: -51.6kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -51.4kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -51.4kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -51.4kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -51.4kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -51.4kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRbeta expressed in human Hela cell lysate measured after overnight incubation by competi...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRbeta expressed in human Hela cell lysate measured after overnight incubation by competi...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRbeta expressed in human Hela cell lysate measured after overnight incubation by competi...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nM ΔG°: -51.1kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Displacement of [3H]-3-methoxy-5-(pyridin-2-ylethynyl)pyridine from mGlu5 receptor of rat cortical membraneMore data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Displacement of [125I]-L-3,5,3'-triiodothyronine from human TRalpha expressed in human Hela cell lysate measured after overnight incubation by compet...More data for this Ligand-Target Pair
Affinity DataKi: 1.20nM ΔG°: -50.9kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nM ΔG°: -50.5kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nM ΔG°: -50.5kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nM ΔG°: -50.4kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nM ΔG°: -50.4kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nM ΔG°: -50.4kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nM ΔG°: -50.4kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nM ΔG°: -50.2kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nM ΔG°: -50.2kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Displacement of [3H]-3-methoxy-5-(pyridin-2-ylethynyl)pyridine from mGlu5 receptor of rat cortical membraneMore data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Binding affinity towards Metabotropic glutamate receptor was determined by displacing [3H]-3-methoxy-5-(pyridin-2-ylethynyl)pyridine from rat cortica...More data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -49.7kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -49.7kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -49.7kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -49.7kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -49.7kJ/moleT: 2°CAssay Description:Binding assays were performed as described in Schaffhauser H, Richards J G, Cartmell J, Chaboz S, Kemp J A, Klingelschmidt A, Messer J, Stadler H, Wo...More data for this Ligand-Target Pair