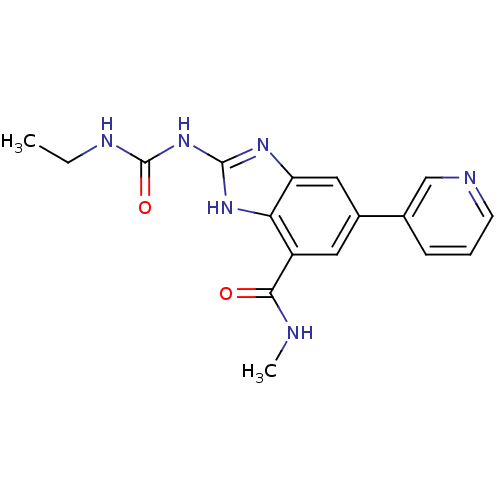
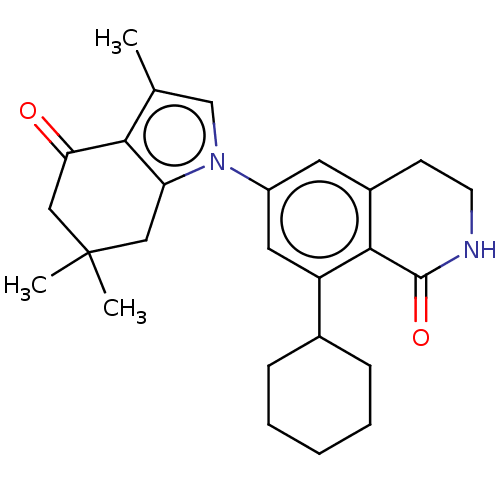
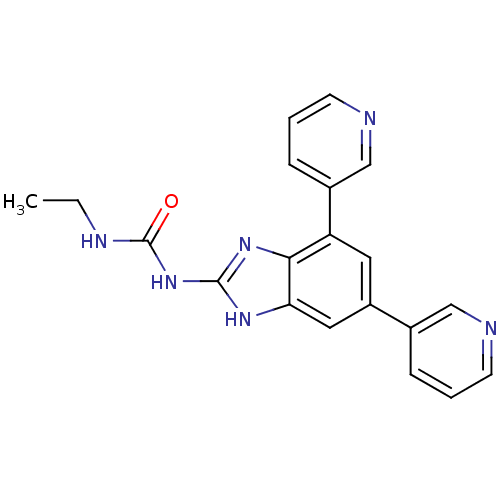
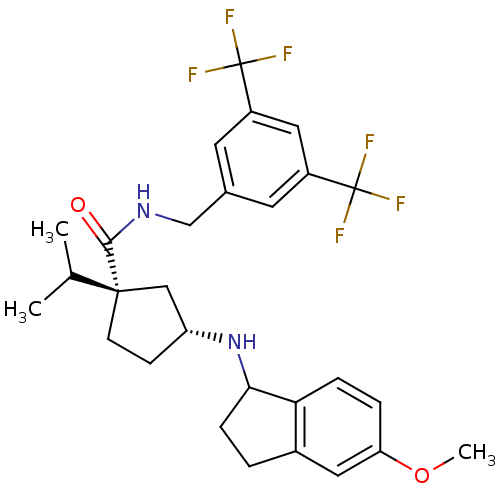
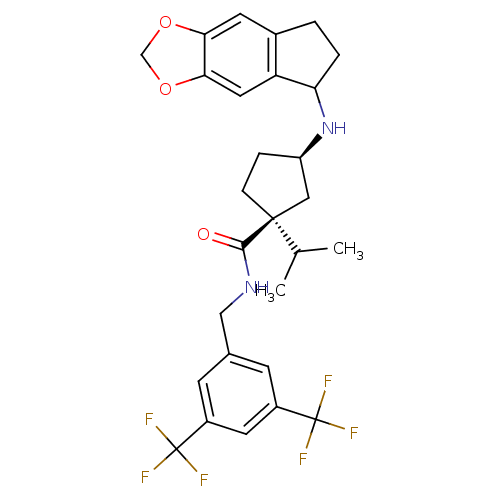
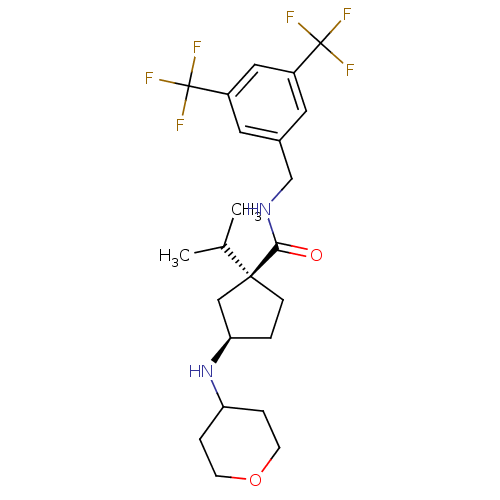
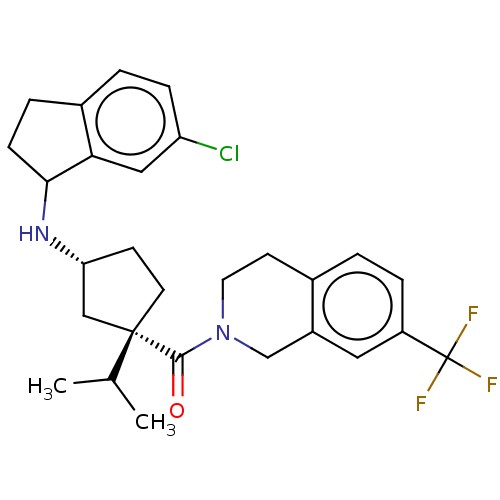
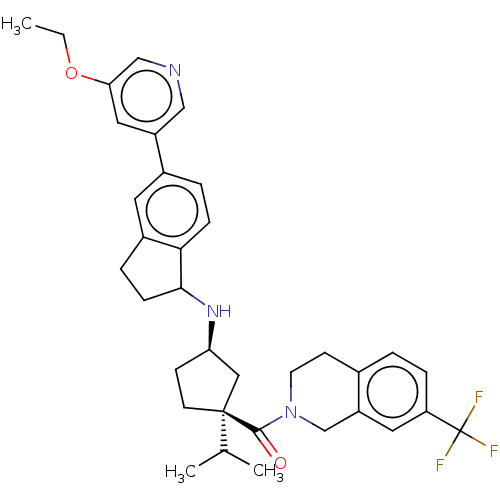
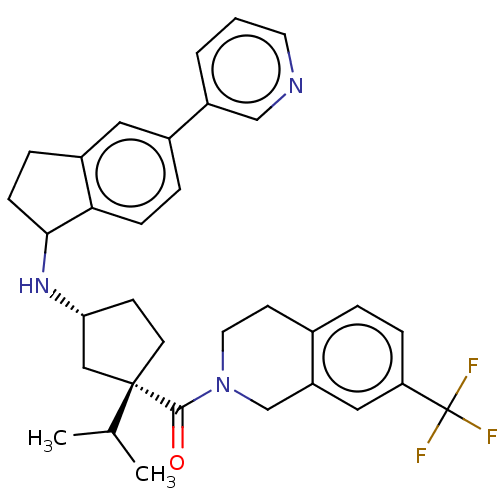
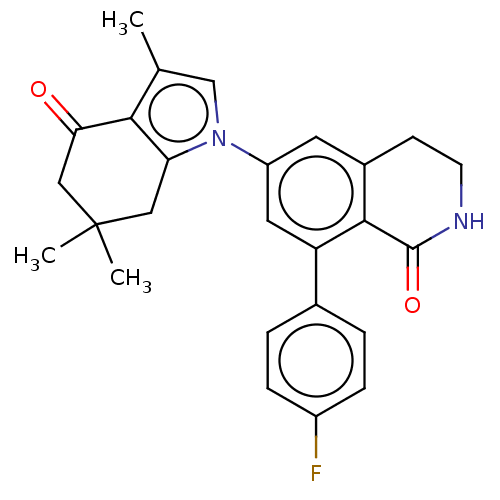
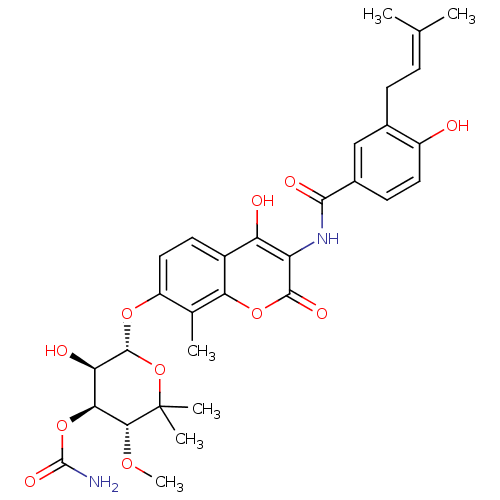
Affinity DataKi: 0.550nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 2.30nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 2.40nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90alpha (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90beta (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.40nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 3.60nMAssay Description:Displacement of [125I]-CCL2 from human CCR2 receptor expressed in human U2OS cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 3.90nMAssay Description:Displacement of [125I]-CCL2 from human CCR2 receptor expressed in human U2OS cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Binding affinity to human N-terminal polyHis-tagged GRP94 (L69 to N337) expressed in Escherichia coli BL21(DE3) after 3 hrs by fluorescence polarizat...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: <4nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 4.40nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 4.5nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 4.60nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 4.60nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 4.90nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Binding affinity to human N-terminal polyHis-tagged GRP94 (L69 to N337) expressed in Escherichia coli BL21(DE3) after 3 hrs by fluorescence polarizat...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90beta (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90alpha (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90beta (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90alpha (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 6.10nMAssay Description:Displacement of [125I]-CCL2 from human CCR2 receptor expressed in human U2OS cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 6.30nMAssay Description:Displacement of [125I]-CCL2 from human CCR2 receptor expressed in human U2OS cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 6.80nMAssay Description:Displacement of [125I]-CCL2 from human CCR2 receptor expressed in human U2OS cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 7nM ΔG°: -47.3kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 7.20nMAssay Description:Displacement of [125I]-CCL2 from human CCR2 receptor expressed in human U2OS cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 7.20nMAssay Description:Displacement of [125I]-CCL2 from human CCR2 receptor expressed in human U2OS cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 7.90nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 8nM ΔG°: -47.0kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Displacement of 0.1 nM [125I]CCL2 from human CCR2 expressing human U2OS cell membrane by scintillation spectrometryMore data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90beta (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -46.4kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -46.4kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -46.4kJ/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair

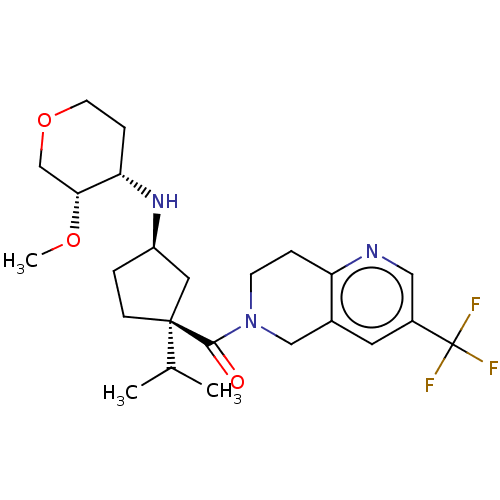
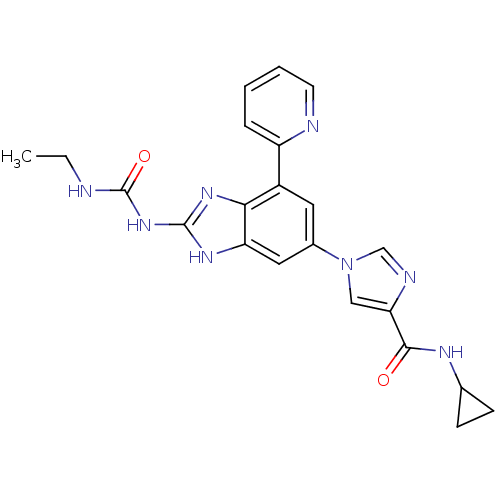
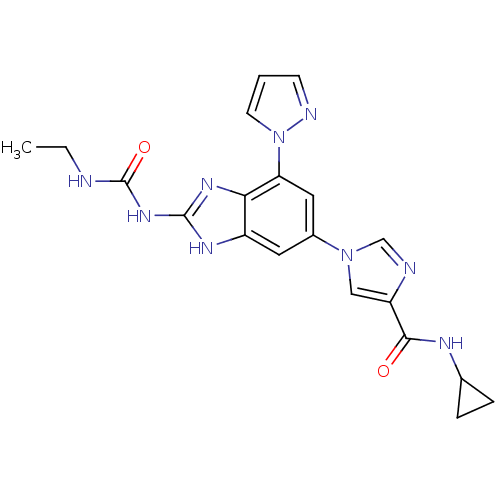
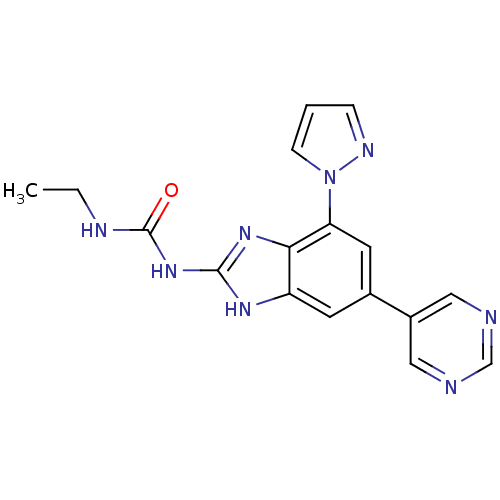
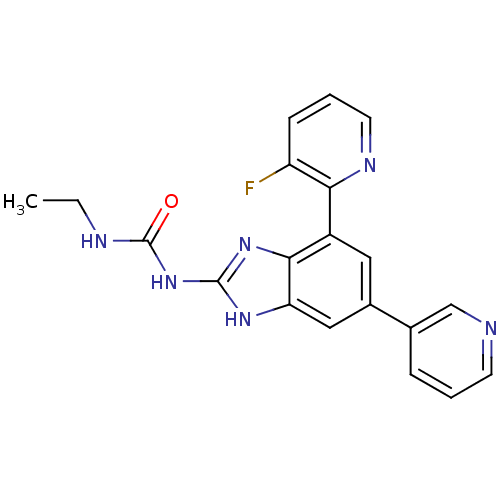
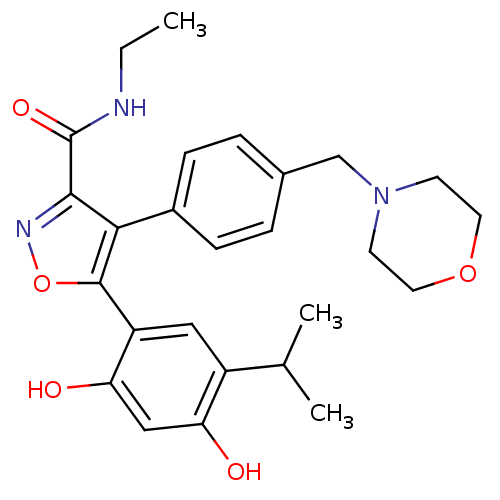
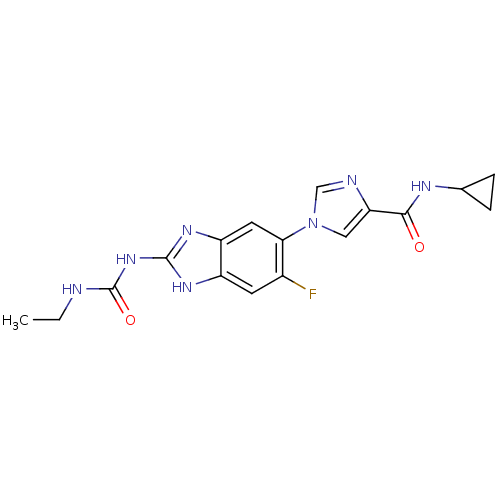
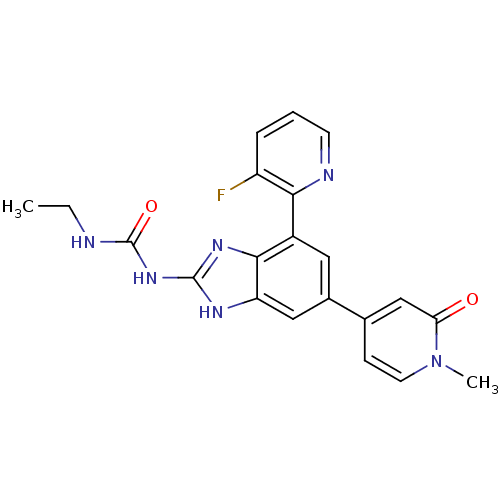
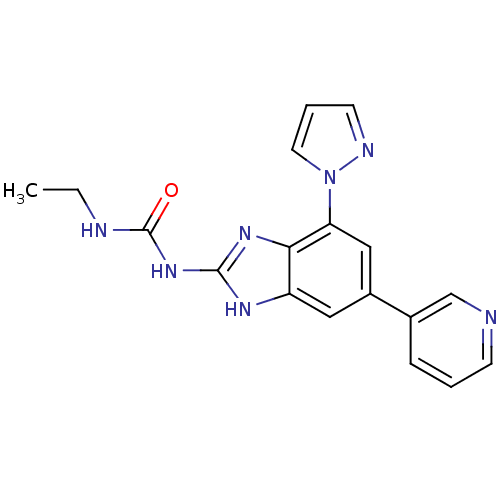

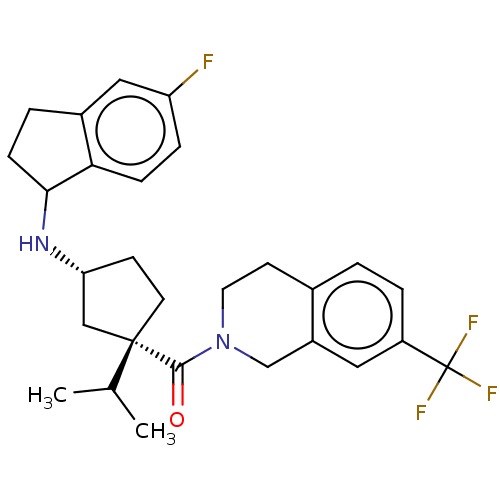
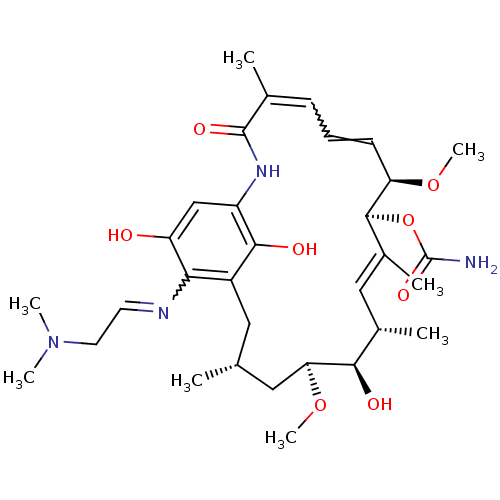
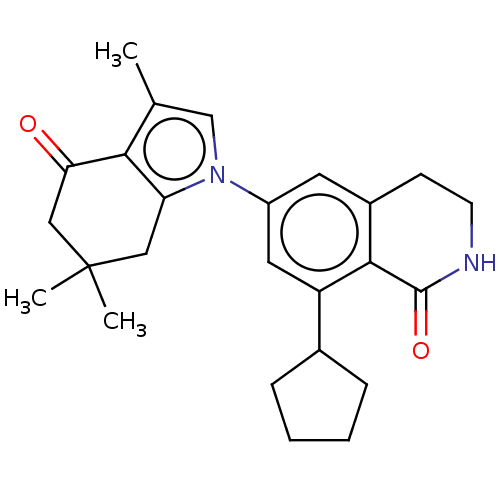
 3D Structure (crystal)
3D Structure (crystal)