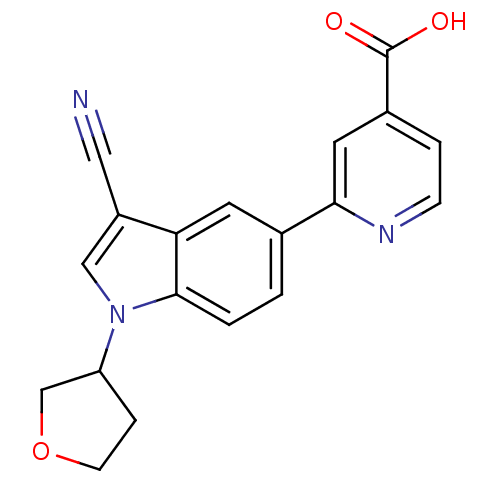
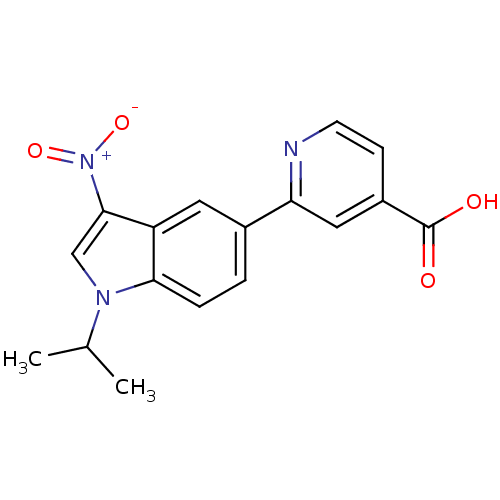
Report error Found 841 with Last Name = 'ho' and Initial = 'es'
Affinity DataKi: 0.5nMAssay Description:Displacement of [3H]-U69593 from kappa opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioli...More data for this Ligand-Target Pair
Affinity DataKi: 0.900nMAssay Description:Displacement of [3H]DAMGO from mu opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioligand ...More data for this Ligand-Target Pair
Affinity DataKi: 4.80nMAssay Description:Non-competitive mixed-type inhibition of SDS-activated human erythrocytes 20S proteasome chymotrypsin like activity using Suc-LLVY-AMC as substrate m...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Non-competitive mixed-type inhibition of SDS-activated human erythrocytes 20S proteasome chymotrypsin like activity using Suc-LLVY-AMC as substrate m...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]DAMGO from mu opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioligand ...More data for this Ligand-Target Pair
Affinity DataKi: 49nMAssay Description:Displacement of [3H]-U69593 from kappa opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioli...More data for this Ligand-Target Pair
Affinity DataKi: 54nMAssay Description:Displacement of [3H]DAMGO from mu opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioligand ...More data for this Ligand-Target Pair
Affinity DataKi: 100nMAssay Description:Displacement of [3H]DAMGO from mu opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioligand ...More data for this Ligand-Target Pair
Affinity DataKi: 120nMAssay Description:Displacement of [3H]DAMGO from mu opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioligand ...More data for this Ligand-Target Pair
Affinity DataKi: 220nMAssay Description:Displacement of [3H]DAMGO from mu opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioligand ...More data for this Ligand-Target Pair
Affinity DataKi: 300nMAssay Description:Displacement of [3H]DAMGO from mu opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioligand ...More data for this Ligand-Target Pair
Affinity DataKi: 330nMAssay Description:Displacement of [3H]-U69593 from kappa opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioli...More data for this Ligand-Target Pair
Affinity DataKi: 330nMAssay Description:Displacement of [3H]DAMGO from mu opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioligand ...More data for this Ligand-Target Pair
Affinity DataKi: 580nMAssay Description:Displacement of [3H]-U69593 from kappa opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioli...More data for this Ligand-Target Pair
Affinity DataKi: 580nMAssay Description:Displacement of [3H]-U69593 from kappa opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioli...More data for this Ligand-Target Pair
Affinity DataKi: 1.17E+3nMAssay Description:Displacement of [3H]-U69593 from kappa opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioli...More data for this Ligand-Target Pair
Affinity DataKi: 1.67E+3nMAssay Description:Displacement of [3H]-U69593 from kappa opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioli...More data for this Ligand-Target Pair
Affinity DataKi: 2.25E+3nMAssay Description:Displacement of [3H]-U69593 from kappa opioid receptor (unknown origin) assessed as inhibition constant incubated for 180 mins by competitive radioli...More data for this Ligand-Target Pair
Affinity DataKi: 4.30E+3nMAssay Description:TP_TRANSPORTER: inhibition of PAH uptake in OAT1-expressing CHO cellsMore data for this Ligand-Target Pair
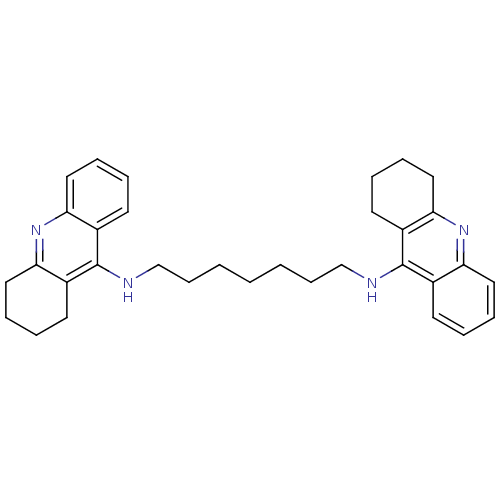
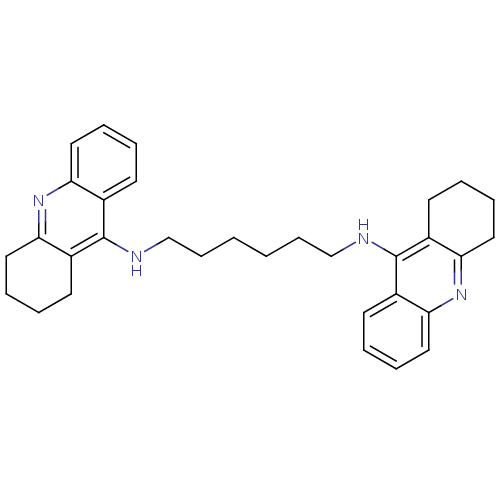
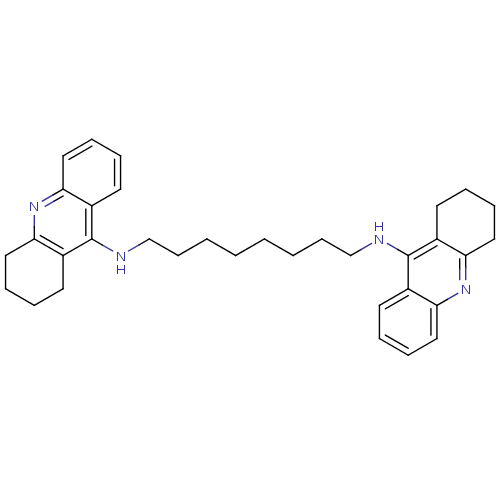
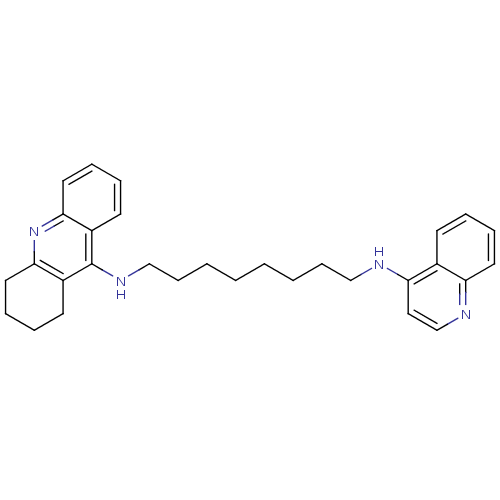
Affinity DataIC50: 1.5nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 3.40nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 3.80nMpH: 7.4 T: 37°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 3.80nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
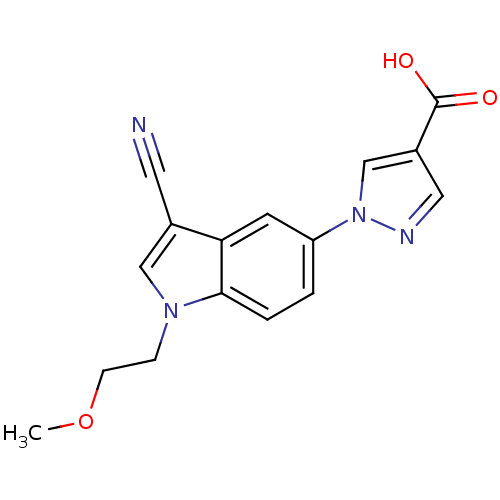
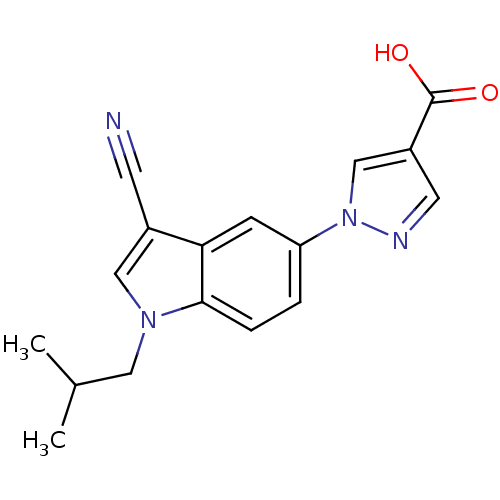
Affinity DataIC50: 4nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 4.5nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 4.70nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 4.80nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 5.20nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 5.40nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
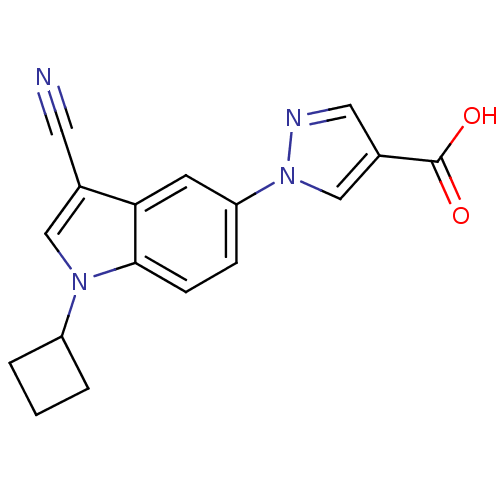
Affinity DataIC50: 6nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.20nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.20nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.70nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.90nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 7.80nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 8.5nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 8.80nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 10.1nMAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Xanthine oxidase originated from bovine milk was incubated for 3 min with test compounds, and the initial velocity of uric acid formation was determi...More data for this Ligand-Target Pair
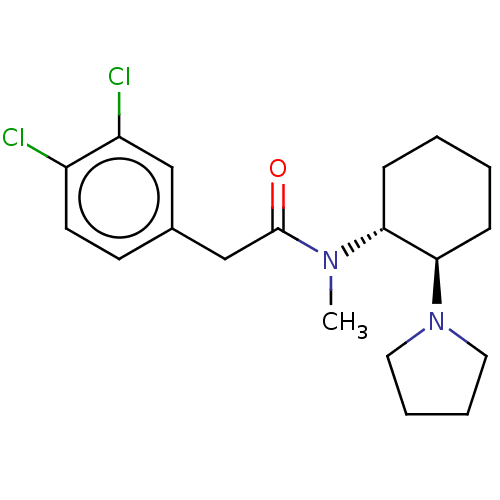
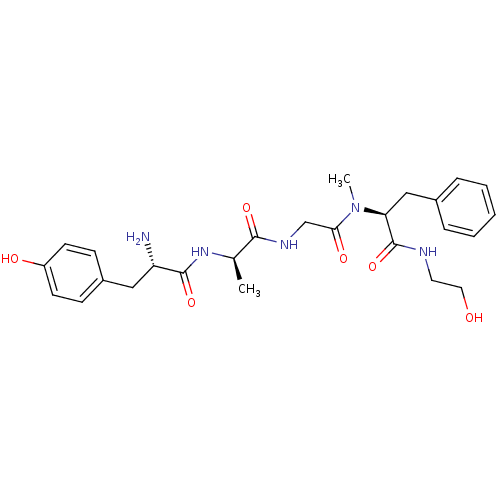
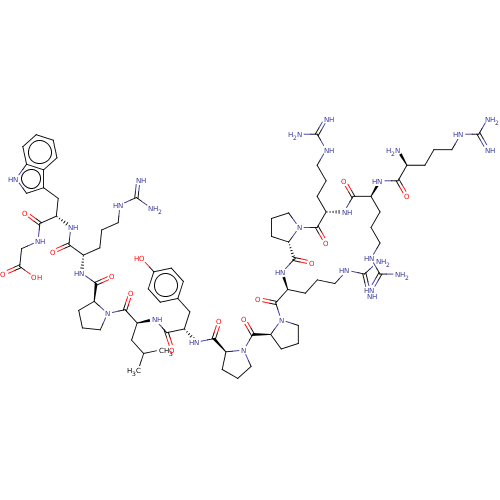
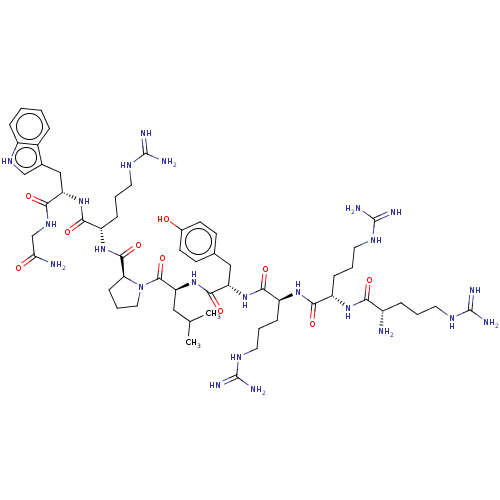
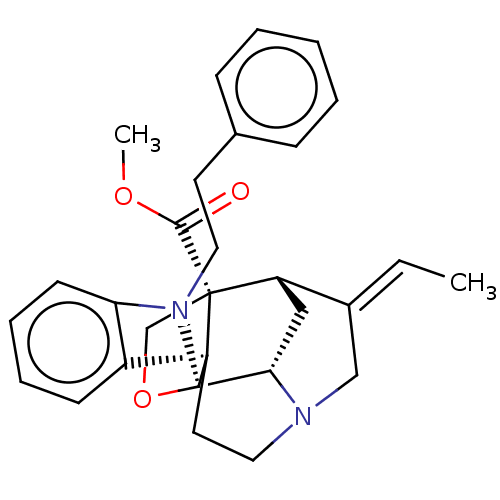
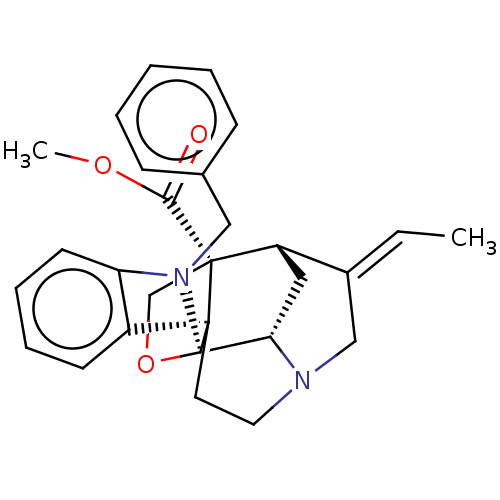
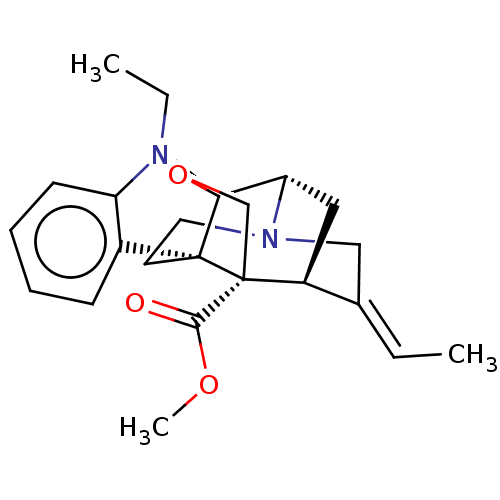
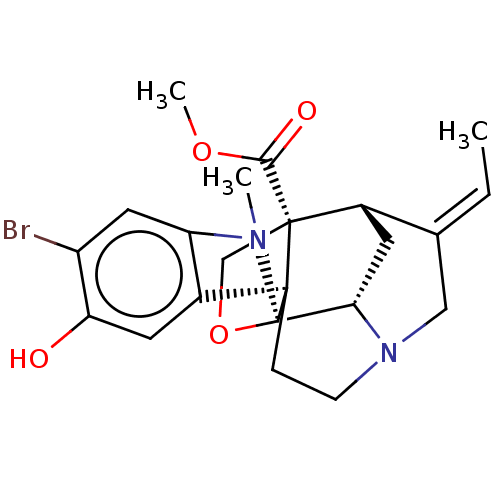
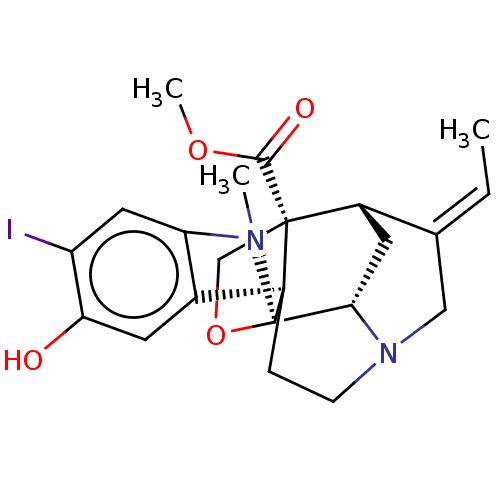
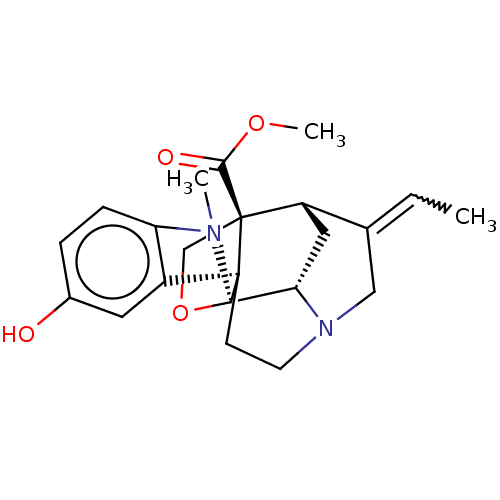
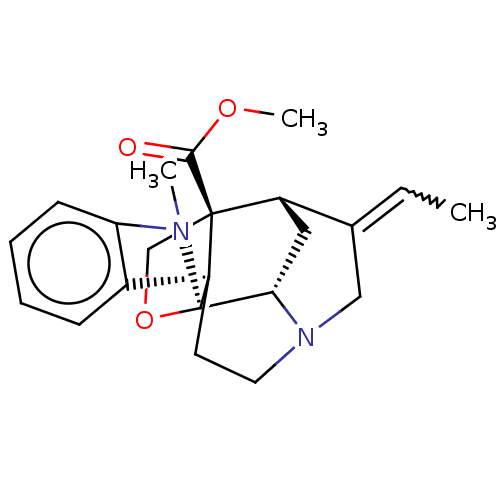
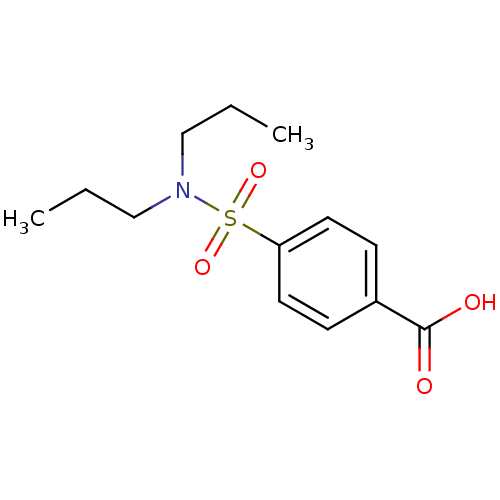
 3D Structure (crystal)
3D Structure (crystal)