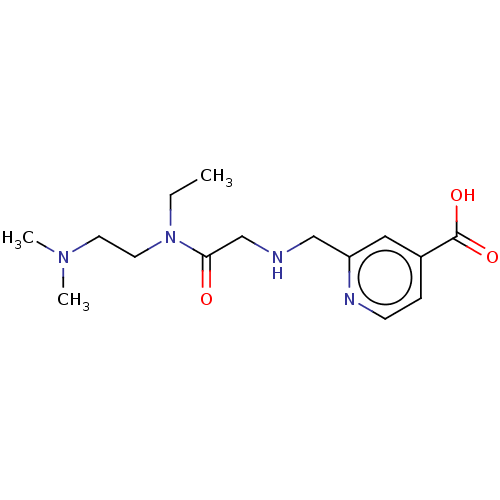
BDBM191598 2-(((2-((2-(dimethylamino)ethyl)(ethyl)amino)-2-oxoethyl)amino)methyl)isonicotinic acid (KDM5-C49)::KDM5-C49::KDOAM-20
SMILES CCN(CCN(C)C)C(=O)CNCc1cc(ccn1)C(O)=O
InChI Key InChIKey=RGGMVQDDHDCRDM-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 47 hits for monomerid = 191598
Found 47 hits for monomerid = 191598
Affinity DataIC50: 4.80E+3nMT: 2°CAssay Description:For inhibition assays, powders of inhibitor compounds were dissolved in 100% (v/v) DMSO to give a 20 or 50 mM concentration and stored at -20 �C unti...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of KDM6B (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMT: 2°CAssay Description:For inhibition assays, powders of inhibitor compounds were dissolved in 100% (v/v) DMSO to give a 20 or 50 mM concentration and stored at -20 �C unti...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMT: 2°CAssay Description:For inhibition assays, powders of inhibitor compounds were dissolved in 100% (v/v) DMSO to give a 20 or 50 mM concentration and stored at -20 �C unti...More data for this Ligand-Target Pair
Affinity DataKd: 52nMAssay Description:ITC experiments were carried out at an enzyme [E] concentration of 25-50 μM with 0.25-1 mM compound [I] concentration in the KDM5 storage buffer...More data for this Ligand-Target Pair
Affinity DataKd: 80nMAssay Description:ITC experiments were carried out at an enzyme [E] concentration of 25-50 μM with 0.25-1 mM compound [I] concentration in the KDM5 storage buffer...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMpH: 7.2 T: 2°CAssay Description:Reactions were performed in 50 mM HEPES [pH 7.2 for KDM5 or pH 7.5 for KDM4A(1-350)] containing 0.01% BSA and 0.01% Tween-20. To each well, 3 μL...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMpH: 7.2 T: 2°CAssay Description:Reactions were performed in 50 mM HEPES [pH 7.2 for KDM5 or pH 7.5 for KDM4A(1-350)] containing 0.01% BSA and 0.01% Tween-20. To each well, 3 μL...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Demethylase reactions and AlphaScreen assays were performed as described (Sayegh et al., 2013), with a few exceptions. All demethylase buffers contai...More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Demethylase reactions and AlphaScreen assays were performed as described (Sayegh et al., 2013), with a few exceptions. All demethylase buffers contai...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Demethylase reactions and AlphaScreen assays were performed as described (Sayegh et al., 2013), with a few exceptions. All demethylase buffers contai...More data for this Ligand-Target Pair
Affinity DataIC50: 59nMpH: 7.2 T: 2°CAssay Description:Reactions were performed in 50 mM HEPES [pH 7.2 for KDM5 or pH 7.5 for KDM4A(1-350)] containing 0.01% BSA and 0.01% Tween-20. To each well, 3 μL...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 780nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMpH: 7.5Assay Description:The demethylase AlphaScreen assay was performed in 384-well plate format using white proxiplates (PerkinElmer), and transfer of compound (100 nl) was...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 600nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:We previously described 4-carboxy-2-triazolopyridines as selective KDM2 inhibitors (England et al., 2014). In efforts to find alternative scaffolds b...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of recombinant human C-terminal FLAG-tagged KDM2B (1 to 650 residues) expressed in baculovirus infected sf9 cells using biotin-H3K36me2 as...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+3nMAssay Description:Inhibition of recombinant KDM3B (842 to 1761 residues) (unknown origin) expressed in baculovirus infected sf9 cells using biotin-H3K9me as substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: <100nMAssay Description:Inhibition of recombinant human N-terminal His-tagged KDM4A (1 to 350 residues) expressed in Escherichia coli using methylated biotin-H3K9 substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: <100nMAssay Description:Inhibition of KDM4C (1 to 349 residues) (unknown origin) expressed in Escherichia coli using biotinylated histone H3 as substrate preincubated for 10...More data for this Ligand-Target Pair
Affinity DataIC50: <100nMAssay Description:Inhibition of KDM5B (1 to 809 residues) (unknown origin) expressed in Escherichia coli using biotin-H3K4me3 as substrate preincubated for 10 mins fol...More data for this Ligand-Target Pair
Affinity DataIC50: <100nMAssay Description:Inhibition of recombinant human N-terminal FLAG-tagged KDM5C (2 to 1560 residues) expressed in baculovirus infected sf9 cells using biotin-H3K4me3 as...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+3nMAssay Description:Inhibition of KDM6A (919 to 1401 residues) (unknown origin) expressed in Escherichia coli using biotinylated histone H3 as substrate preincubated for...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+3nMAssay Description:Inhibition of human C-terminal FLAG-tagged KDM6B (1043 residues) expressed in baculovirus infected sf9 cells using H3K27me3 as substrate preincubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of KDM5C in human U2OS cells assessed as reduction in demethylation of H3K4me3 after 20 hrs by Hoechst 33342 staining based immunofluoresc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of KDM5B in human U2OS cells assessed as reduction in demethylation of H3K4me3 after 20 hrs by Hoechst 33342 staining based immunofluoresc...More data for this Ligand-Target Pair
Affinity DataIC50: <100nMAssay Description:Inhibition of human N-terminal GST-tagged KDM4B (1 to 500 residues) expressed in baculovirus infected sf9 cells using biotinylated histone H3 as subs...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of N-terminal his6-tagged human KDM5B (1 to 769 residues) expressed in sf9 insect cells incubated for 20 mins by Alphascreen assayMore data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Inhibition of N-terminal his6-tagged human KDM5A (1 to 797 residues) expressed in sf9 insect cells incubated for 20 mins by Alphascreen assayMore data for this Ligand-Target Pair
Affinity DataIC50: 59nMAssay Description:Inhibition of N-terminal his6-tagged human KDM5C (1 to 839 residues) expressed in sf9 insect cells incubated for 20 mins by Alphascreen assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of KDM5B (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMT: 2°CAssay Description:For inhibition assays, powders of inhibitor compounds were dissolved in 100% (v/v) DMSO to give a 20 or 50 mM concentration and stored at -20 �C unti...More data for this Ligand-Target Pair