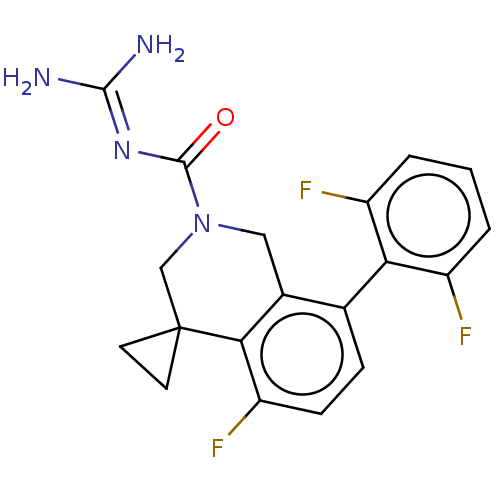
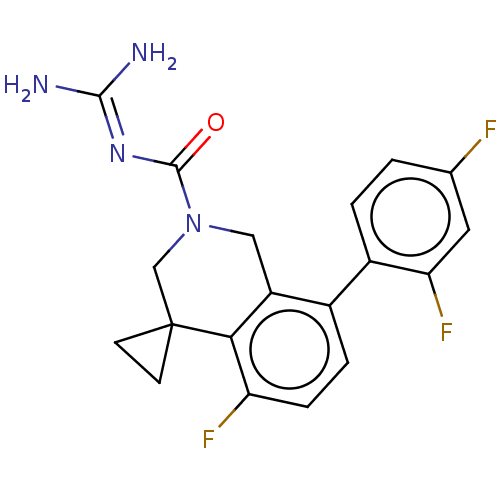
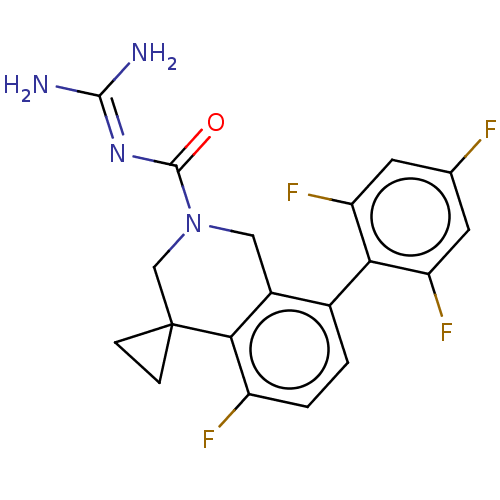
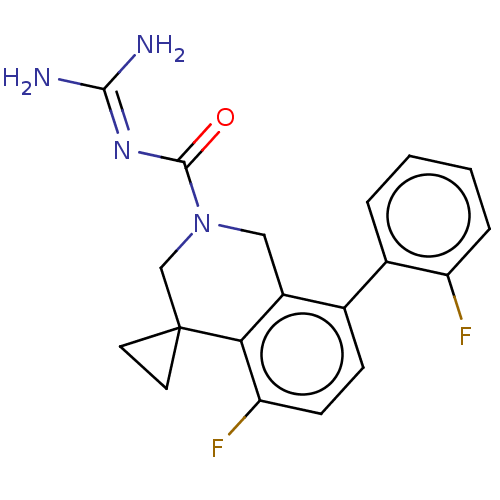
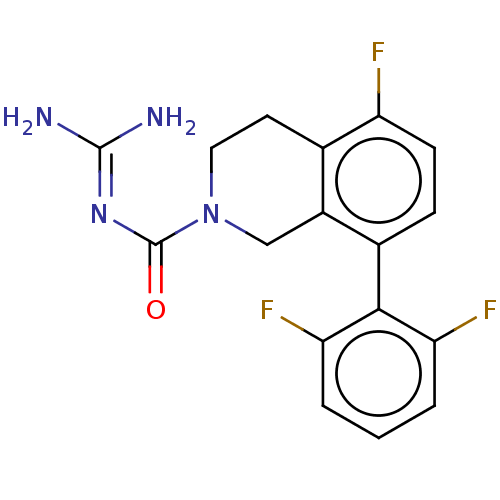
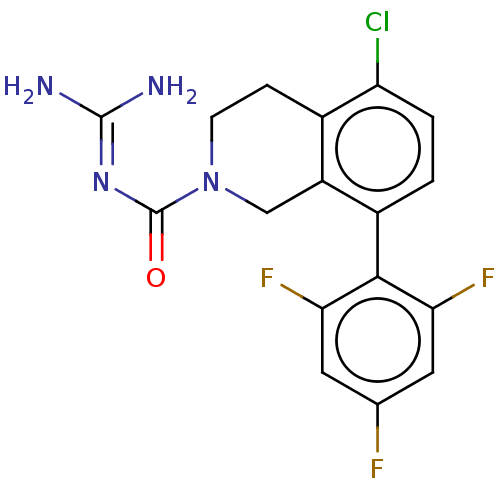
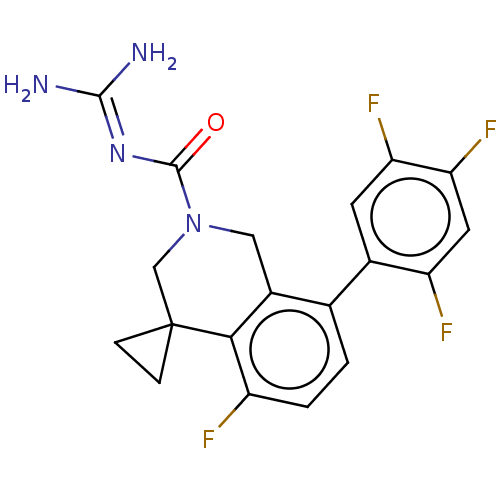
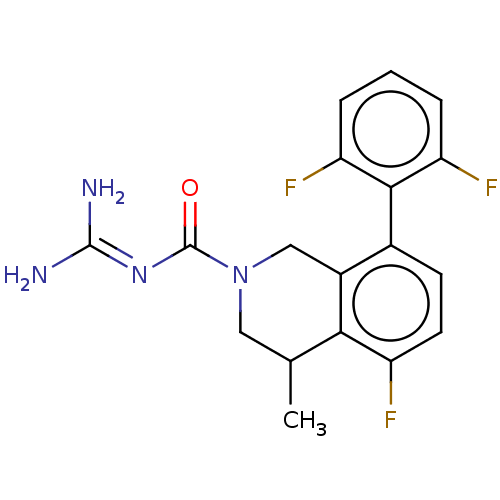
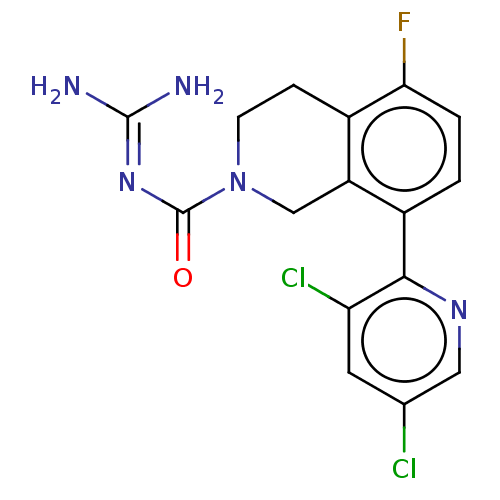
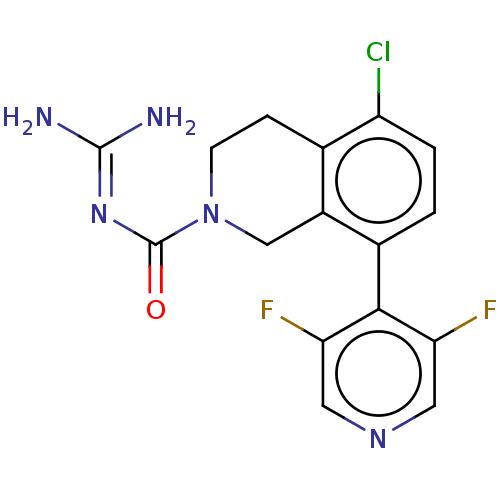
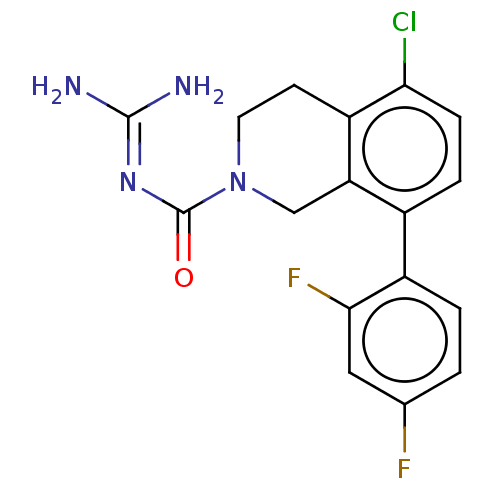
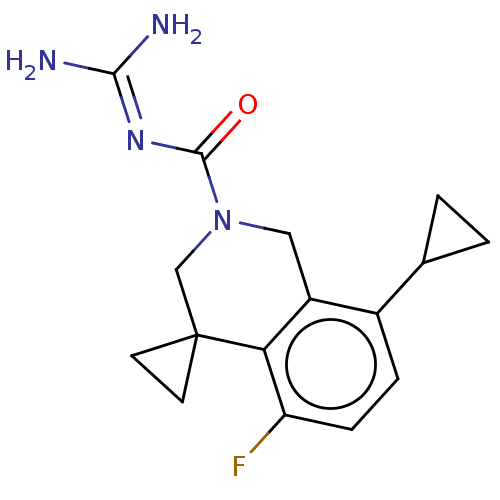
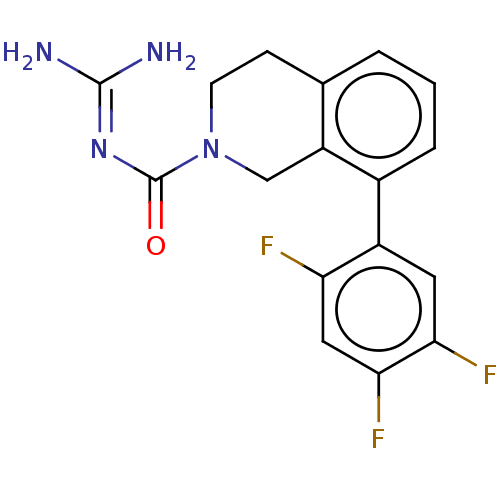
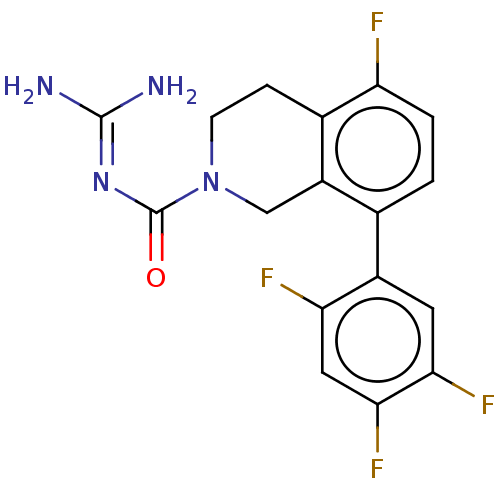
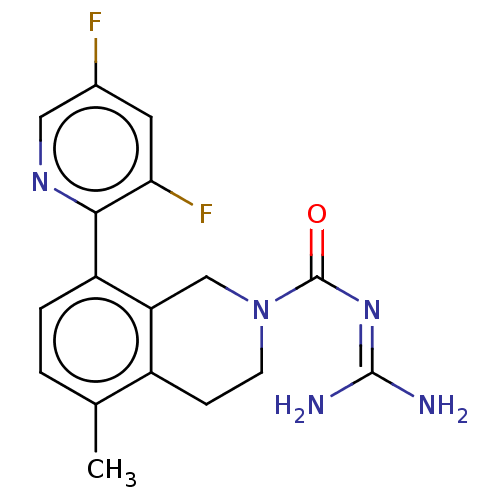
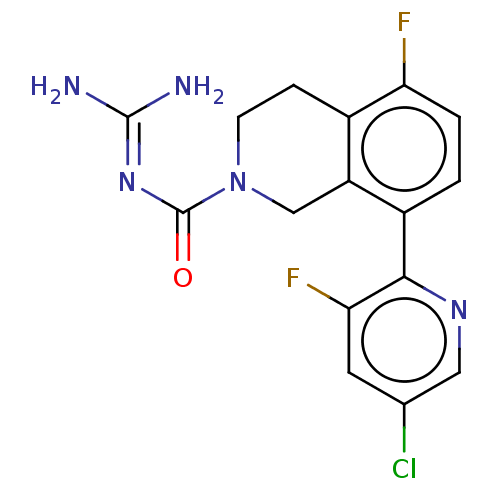
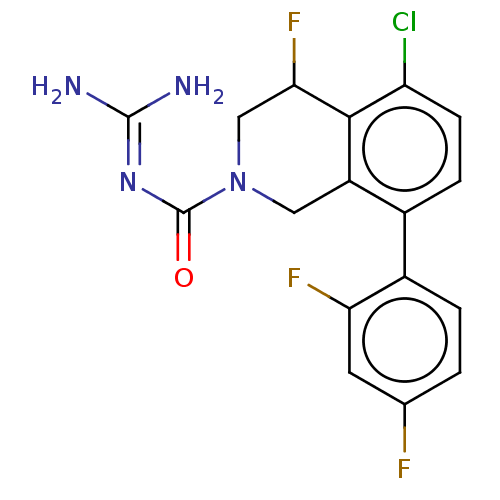
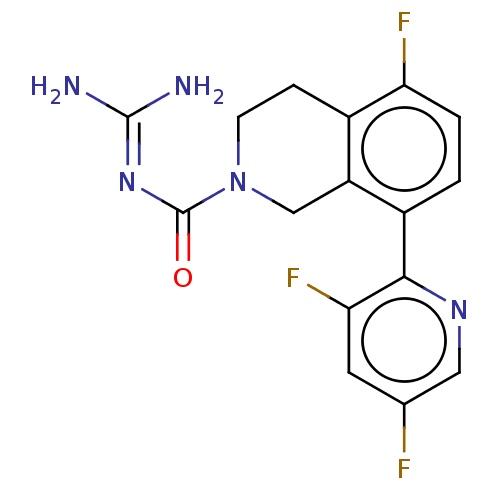
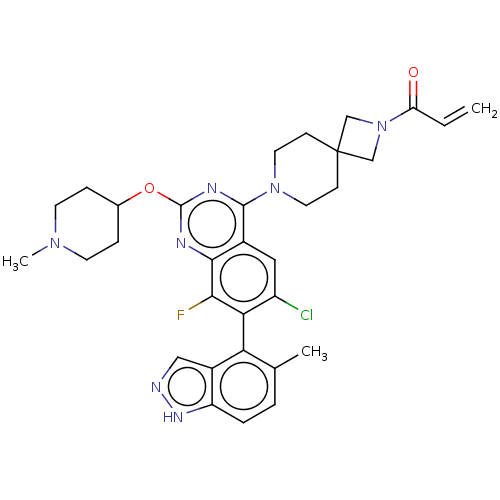
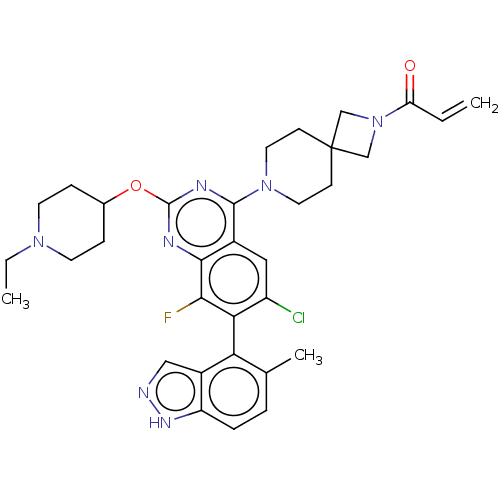
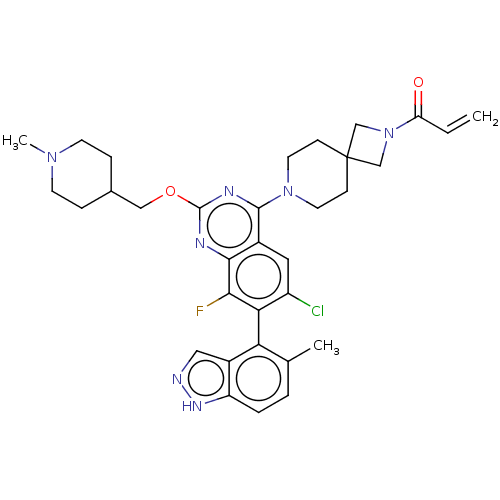
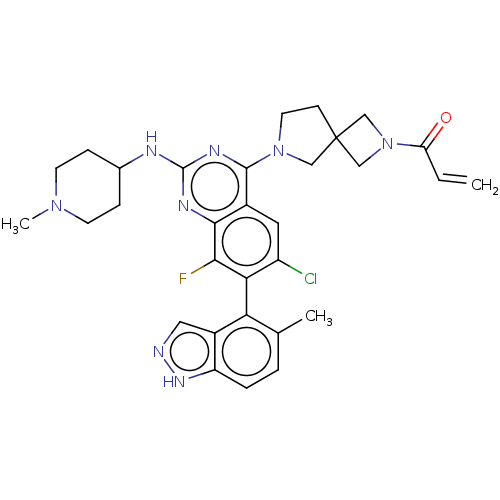
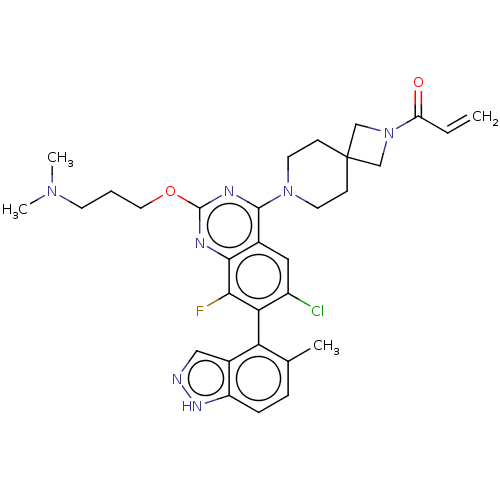
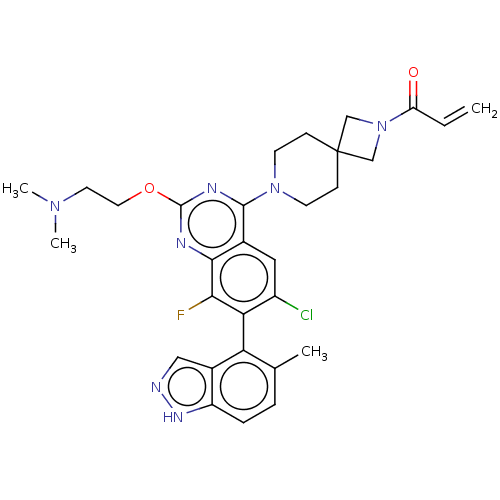
Affinity DataKi: 0.680nM ΔG°: -54.4kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 0.75nM ΔG°: -54.2kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 0.850nM ΔG°: -53.9kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 0.880nM ΔG°: -53.8kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 0.950nM ΔG°: -53.6kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nM ΔG°: -53.2kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nM ΔG°: -53.2kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 1.30nM ΔG°: -52.8kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nM ΔG°: -52.2kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 1.90nM ΔG°: -51.8kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 2.5nM ΔG°: -51.1kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 2.70nM ΔG°: -50.9kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 2.70nM ΔG°: -50.9kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 3.60nM ΔG°: -50.1kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 3.70nM ΔG°: -50.1kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 3.90nM ΔG°: -49.9kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 4.20nM ΔG°: -49.7kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 4.30nM ΔG°: -49.7kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 6.60nM ΔG°: -48.6kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataKi: 6.90nM ΔG°: -48.5kJ/molepH: 7.4 T: 2°CAssay Description:A test compound and 150 uM of a DMSO solution of 5-carboxamide tryptamine (5-CT) were added to a 96-well plate at 2 ul/well and suspended in the incu...More data for this Ligand-Target Pair
Affinity DataIC50: 70nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 77nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 260nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 340nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 350nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 370nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 470nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 650nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 770nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 940nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+3nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+4nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+4nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair
Affinity DataIC50: >3.00E+4nMAssay Description:Inhibition of biotinylated Avi-tagged GDP-bound KRAS G12C mutant (1 to 185 residues) (unknown origin) assessed as inhibition of KRAS G12C mutant/SOS/...More data for this Ligand-Target Pair

 3D Structure (crystal)
3D Structure (crystal)