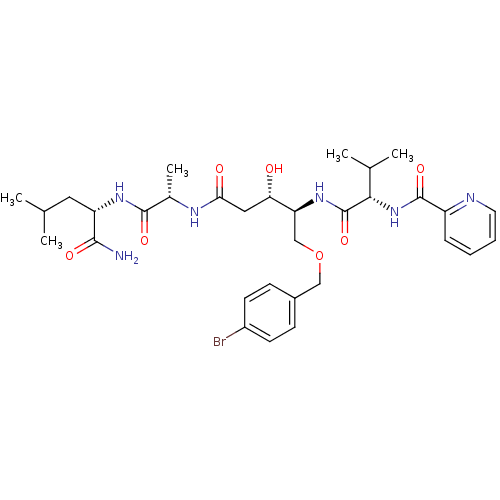
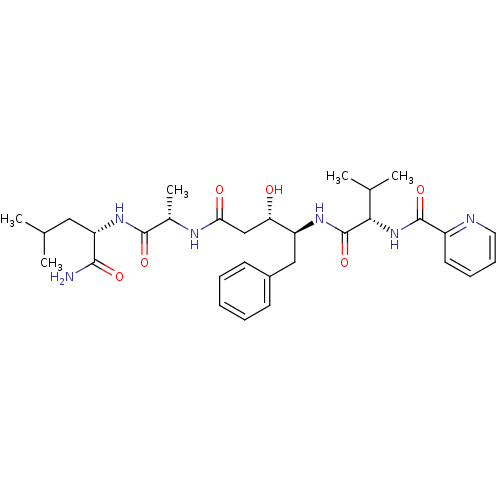
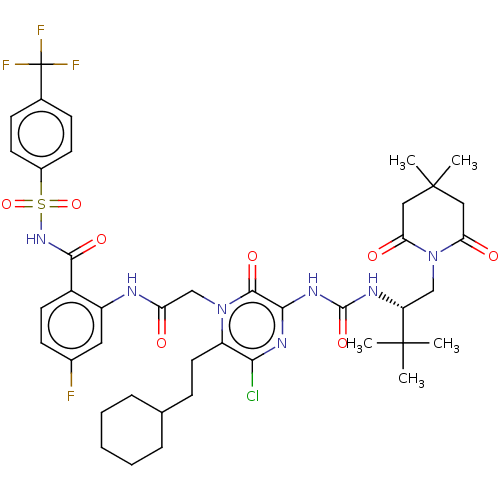
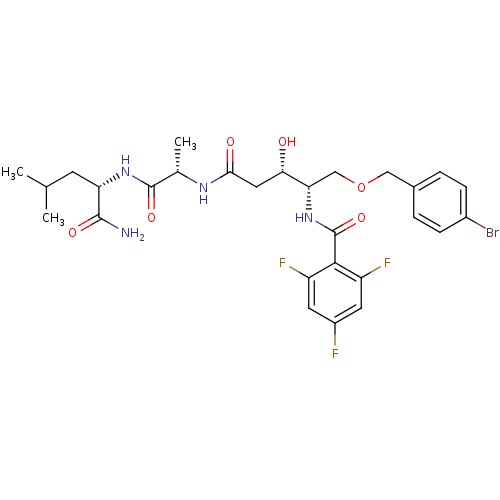
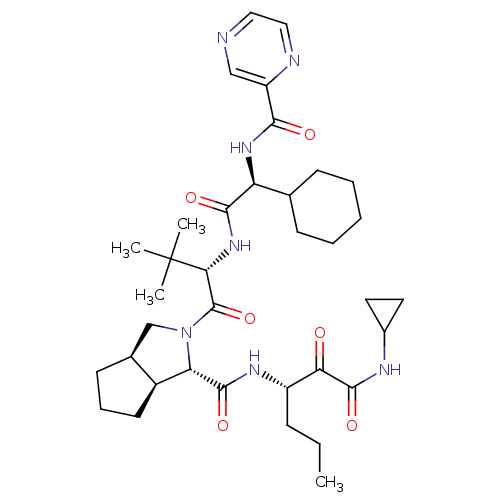
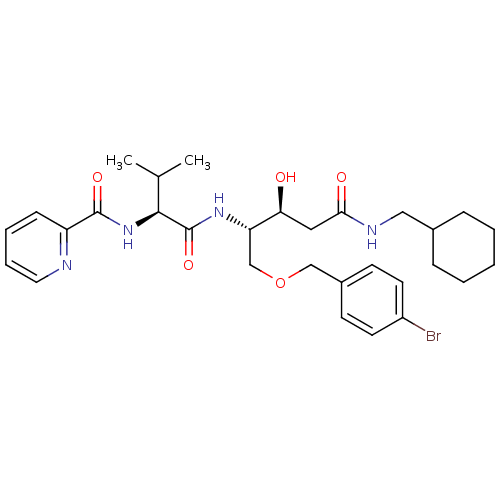
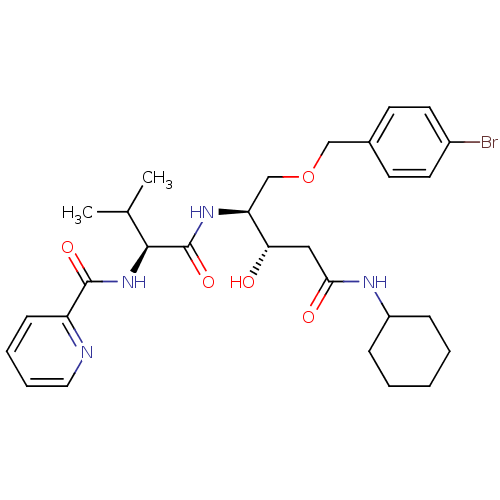
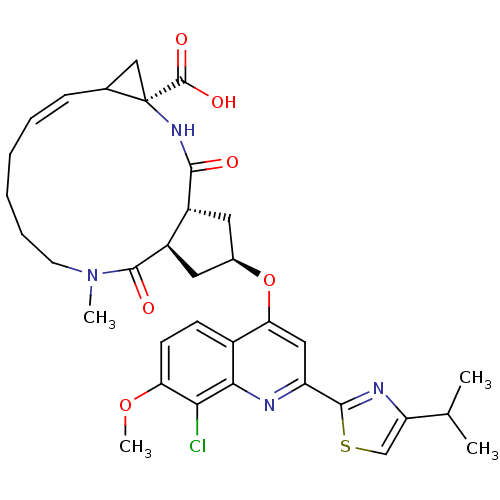
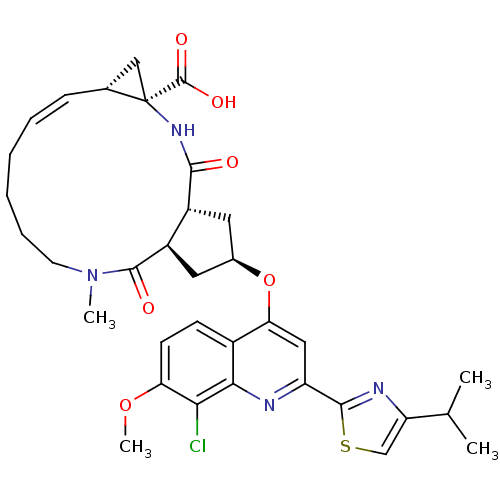
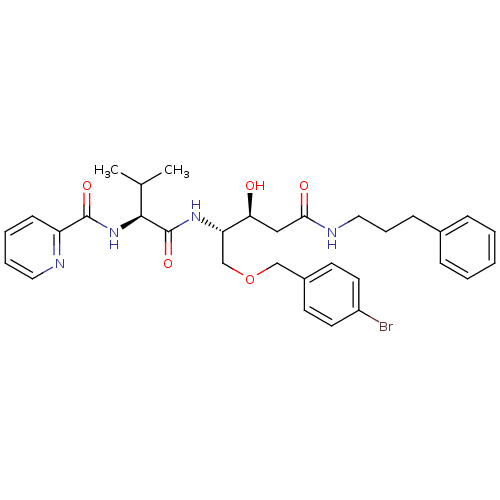
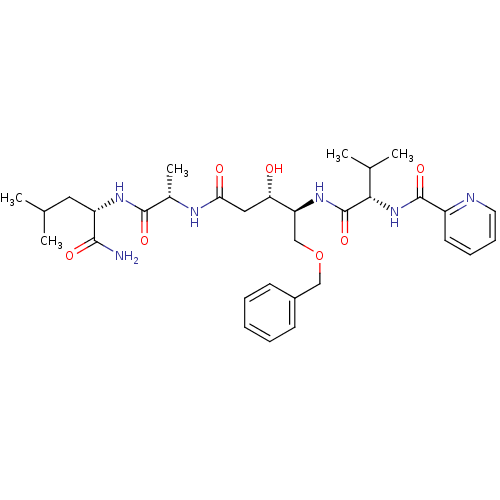
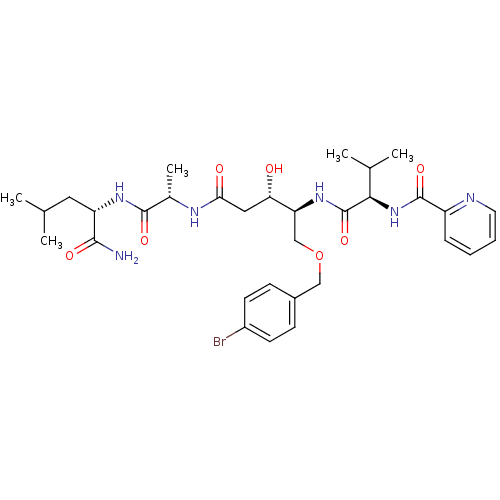
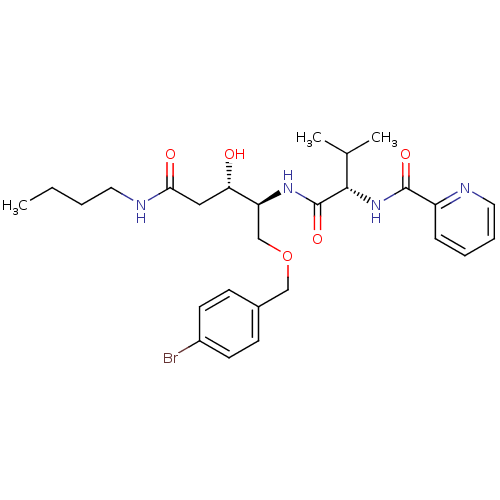
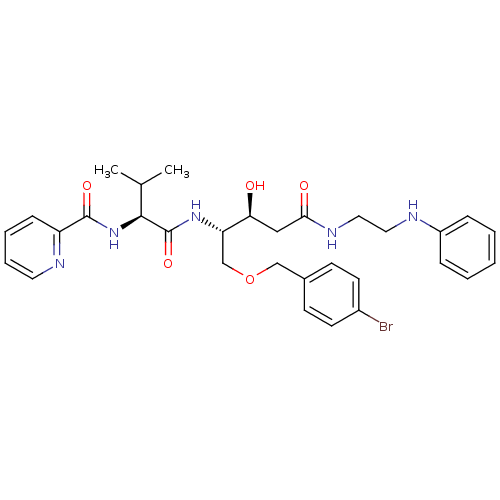
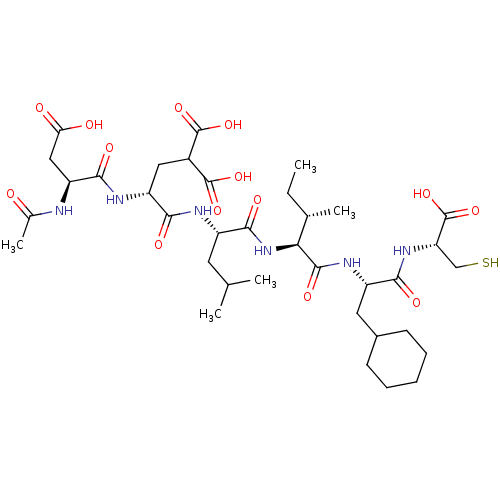
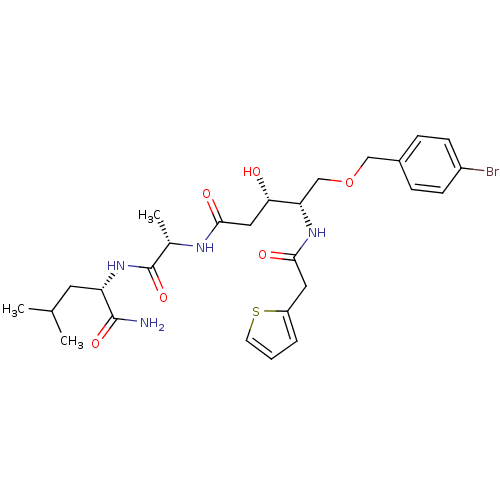
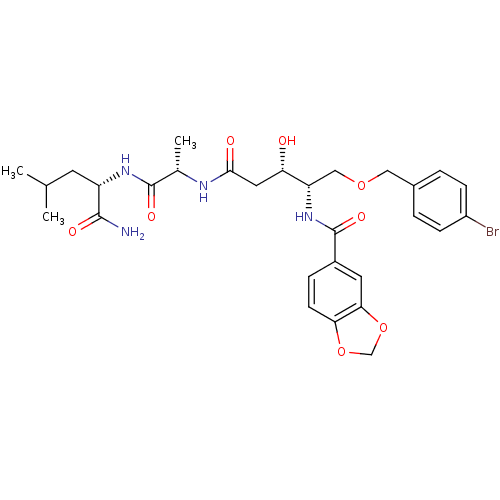
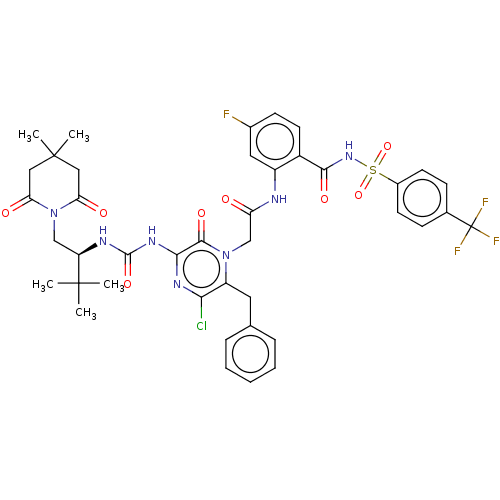
Affinity DataKi: 0.0500nM ΔG°: -59.8kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.0890nMAssay Description:Inhibition of HCV genotype-1a NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -58.0kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -58.0kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -58.0kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.25nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.25nM ΔG°: -55.7kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -55.3kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nM ΔG°: -54.0kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 0.560nM ΔG°: -52.3kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of HCV genotype-1a full length NS3 protease R155K mutant preincubated for 10 mins followed by Ac-DED(Edans)EEAbuj-[COO]ASK(Dabcyl)-NH2 add...More data for this Ligand-Target Pair
Affinity DataKi: 2.20nM ΔG°: -48.9kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 3.40nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 3.90nMAssay Description:Inhibition of full-length HCV NS3 protease D168V mutantMore data for this Ligand-Target Pair
Affinity DataKi: 3.90nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 4.10nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 4.90nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 5nM ΔG°: -48.2kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 5.40nMAssay Description:Inhibition of HCV genotype-3a full length NS3 protease preincubated for 10 mins followed by Ac-DED(Edans)EEAbuj-[COO]ASK(Dabcyl)-NH2 addition in pres...More data for this Ligand-Target Pair
Affinity DataKi: 5.40nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 5.40nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 6.40nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 6.70nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 8.10nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 8.90nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Inhibition of HCV genotype-1a full length NS3 protease preincubated for 10 mins followed by Ac-DED(Edans)EEAbuj-[COO]ASK(Dabcyl)-NH2 addition in pres...More data for this Ligand-Target Pair
Affinity DataKi: 9.40nM ΔG°: -46.6kJ/molepH: 7.5 T: 2°CAssay Description:The inhibition of full-length hepatitis C NS3 protease enzyme was measured essentially as described in Poliakov, 2002 Prot Expression & Purification ...More data for this Ligand-Target Pair
Affinity DataKi: 9.40nMAssay Description:The aim of this in vitro assay was to measure the inhibition of HCV NS3/4A protease complexes by the compounds of the present invention. This assay p...More data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -45.2kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 11nM ΔG°: -45.0kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 15nM ΔG°: -44.2kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Inhibition of HCV genotype-3a full length NS3 protease preincubated for 10 mins followed by Ac-DED(Edans)EEAbuj-[COO]ASK(Dabcyl)-NH2 addition in pres...More data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 24nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 24nMAssay Description:Inhibition of full-length HCV NS3 protease R155K mutantMore data for this Ligand-Target Pair
Affinity DataKi: 25nM ΔG°: -43.0kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 27nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 29nM ΔG°: -42.6kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 30nM ΔG°: -42.5kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Inhibition of HCV genotype-1a full length NS3 protease R155K mutant preincubated for 10 mins followed by Ac-DED(Edans)EEAbuj-[COO]ASK(Dabcyl)-NH2 add...More data for this Ligand-Target Pair

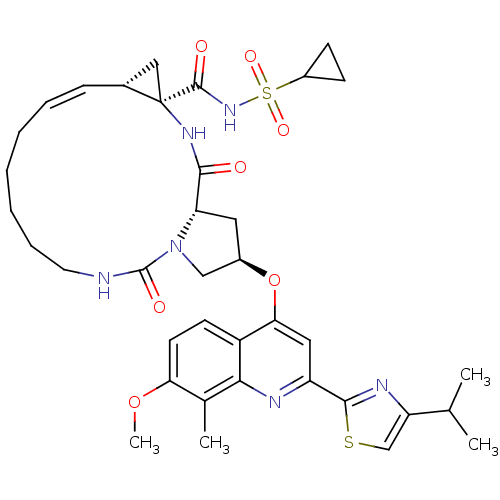
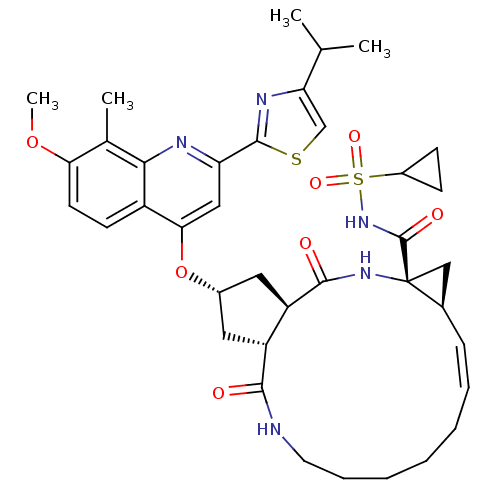
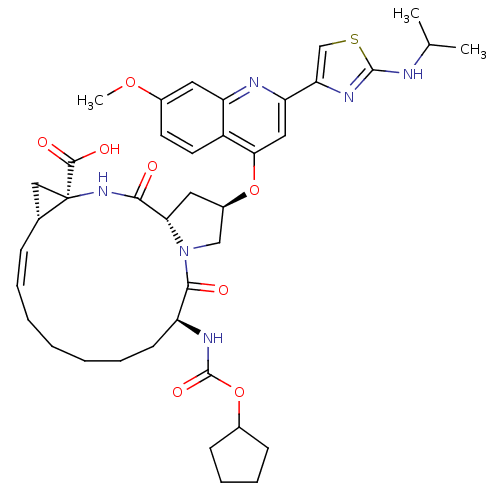
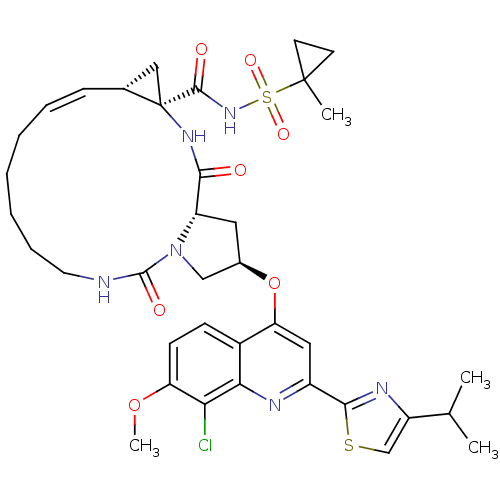
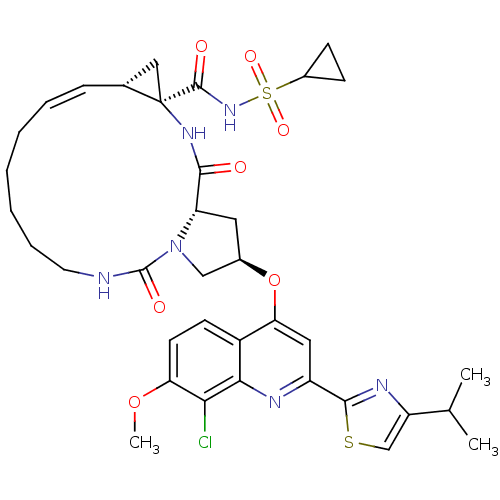
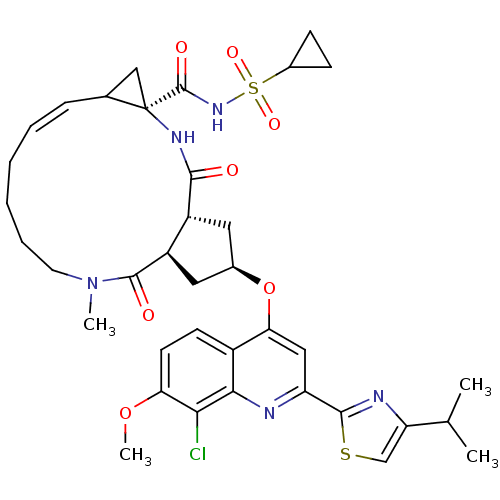
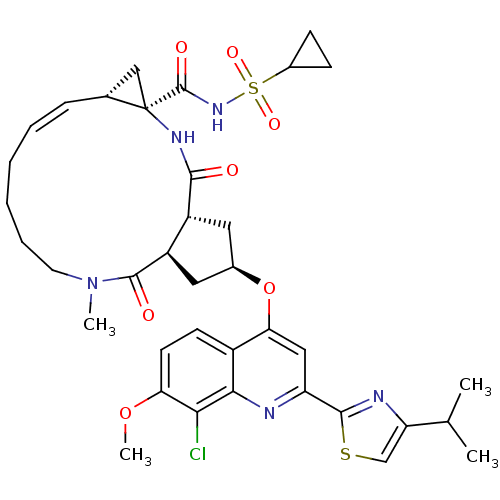
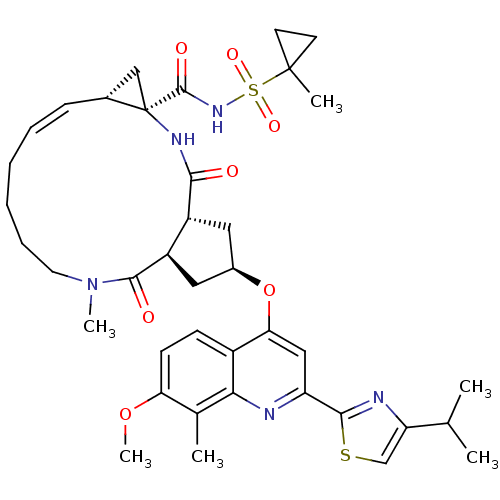
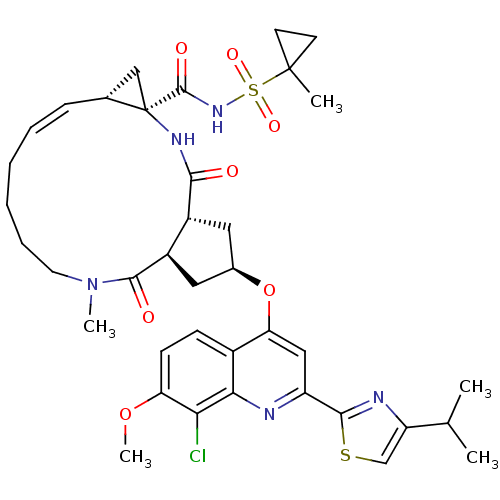
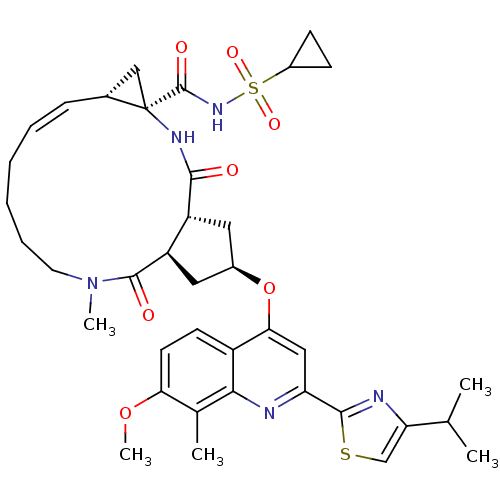
 3D Structure (crystal)
3D Structure (crystal)