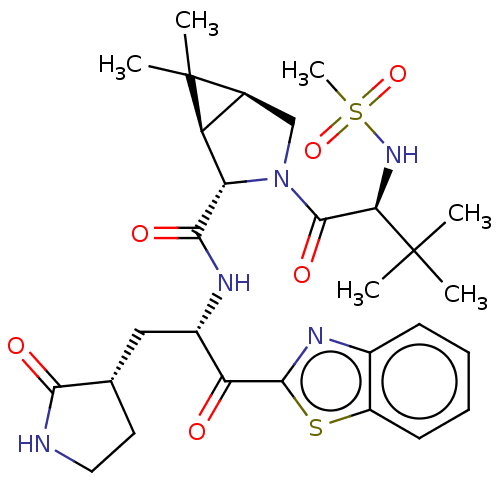
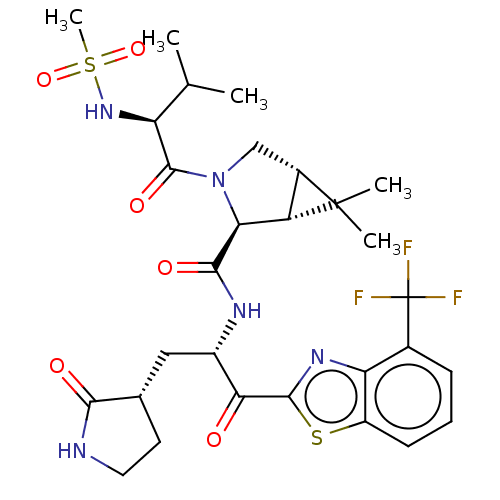
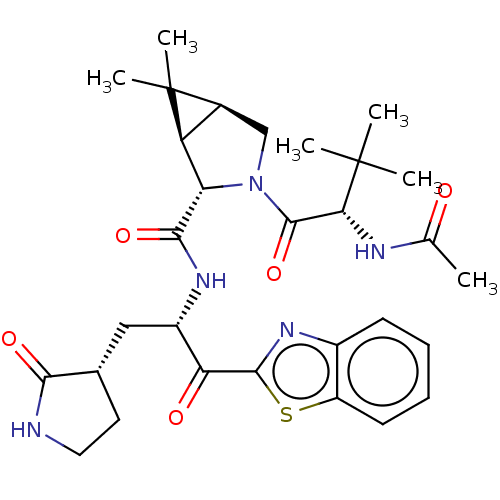
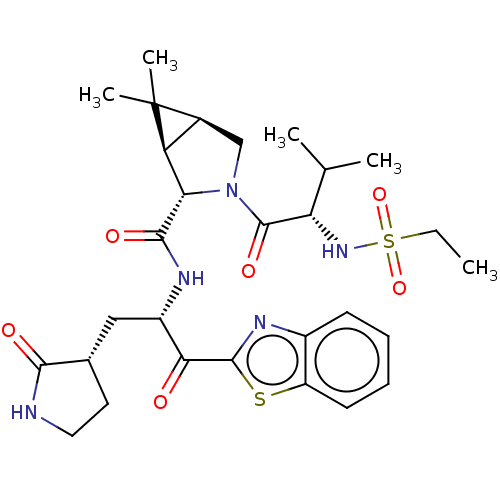
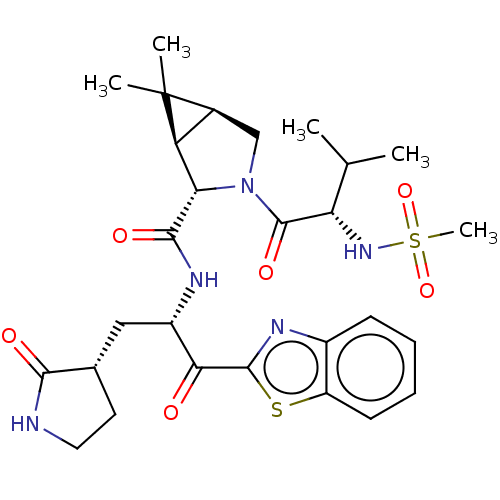
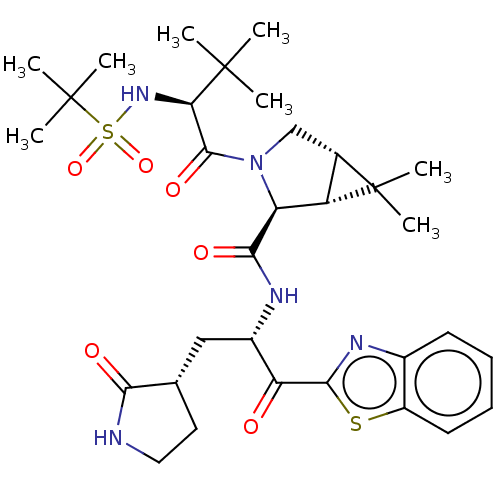
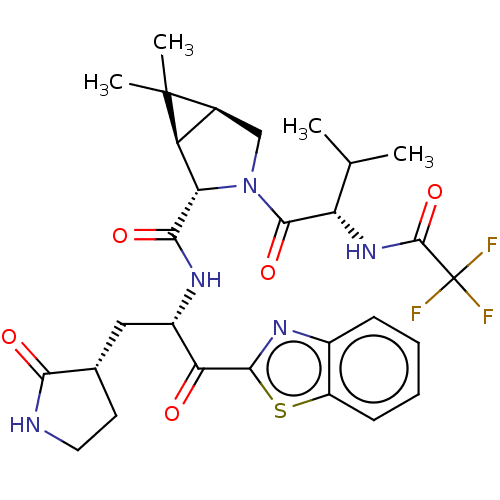
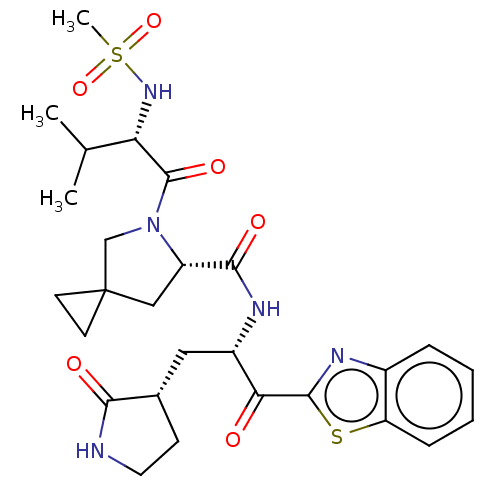
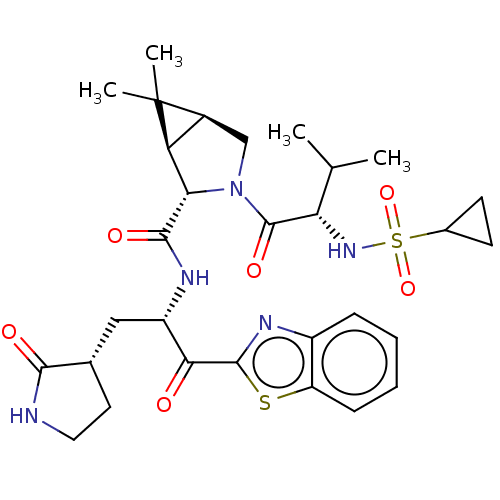
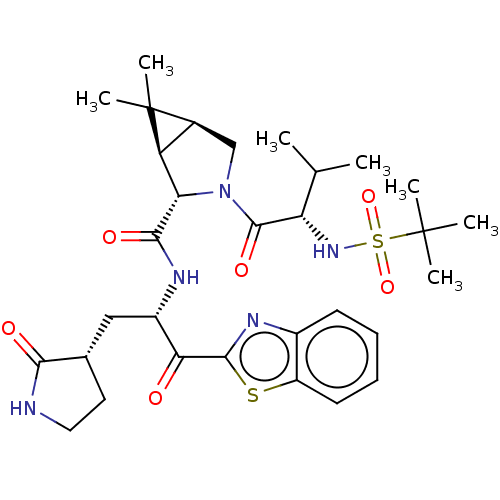
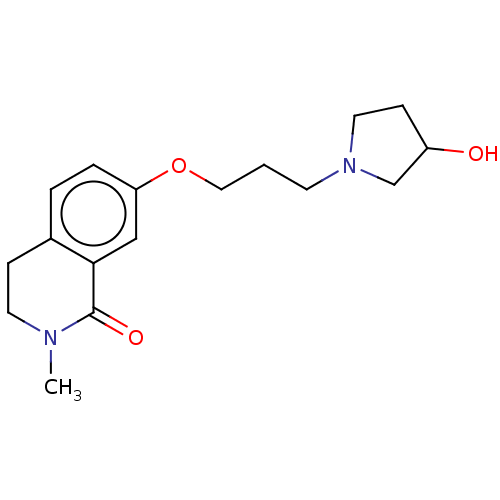
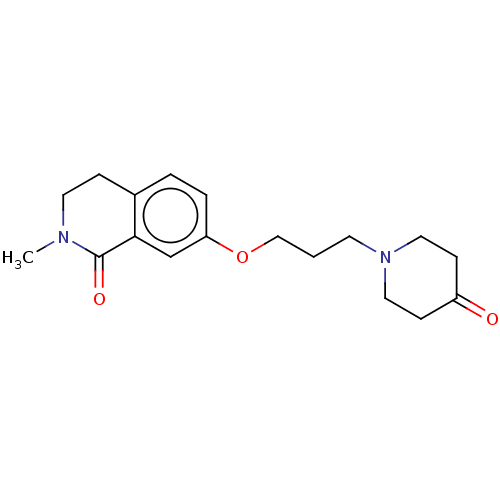
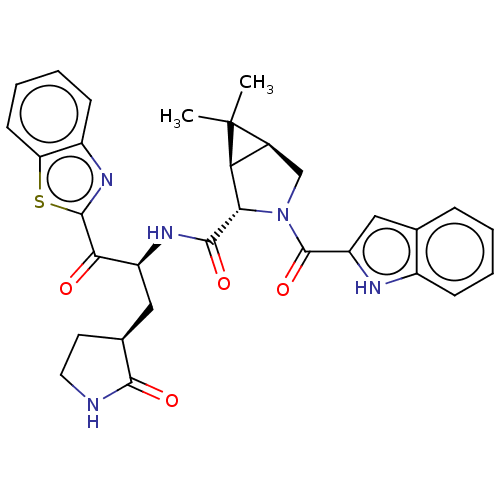
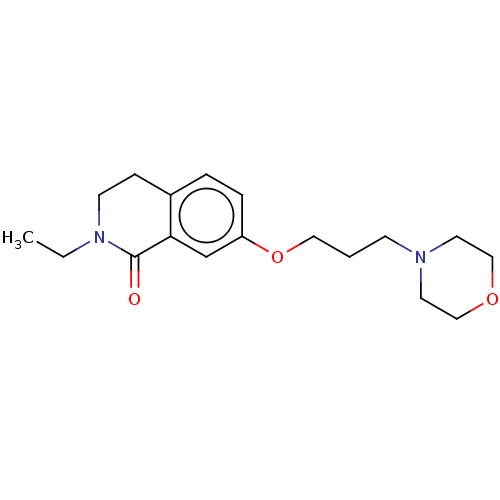
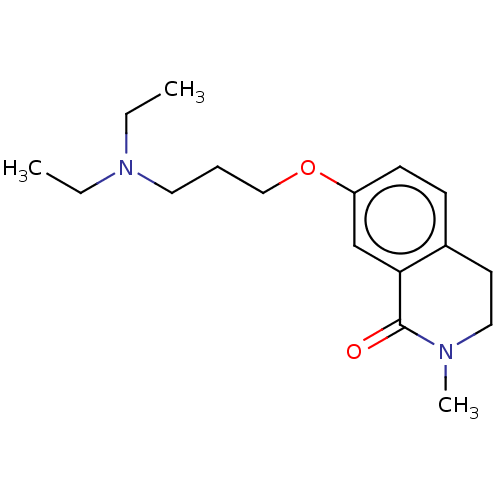
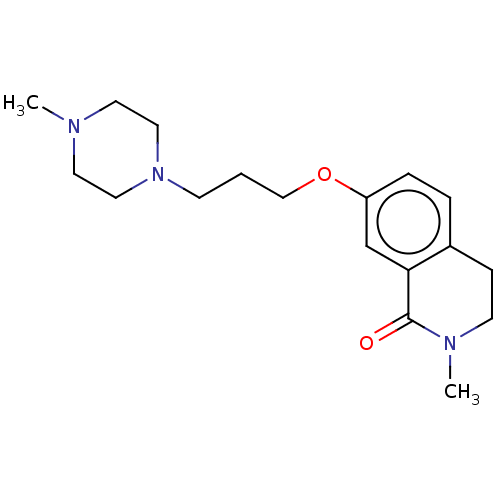
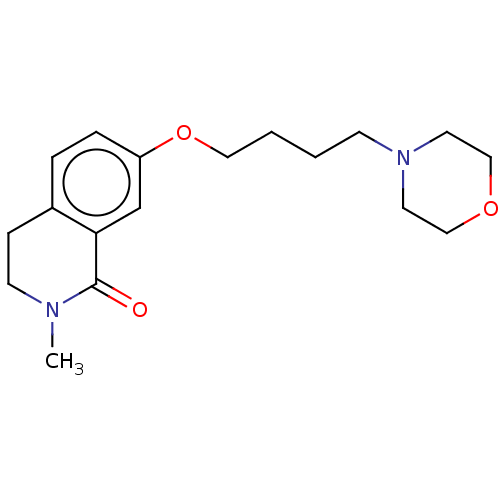
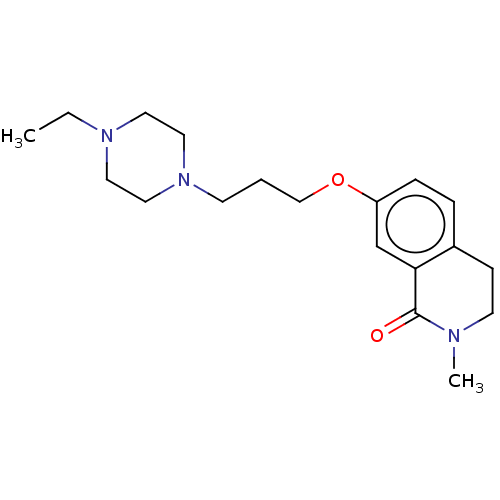
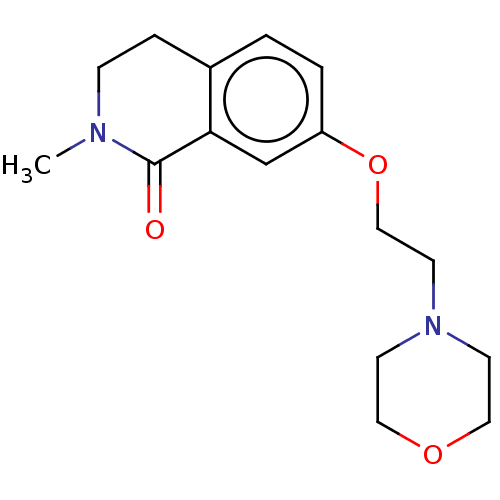
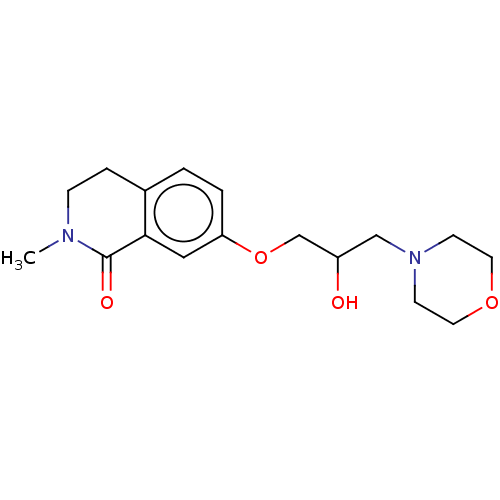
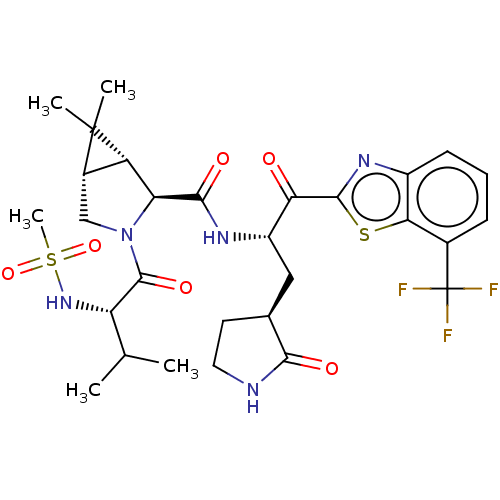
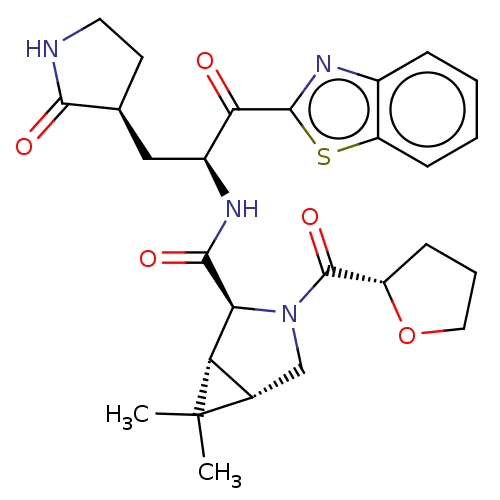
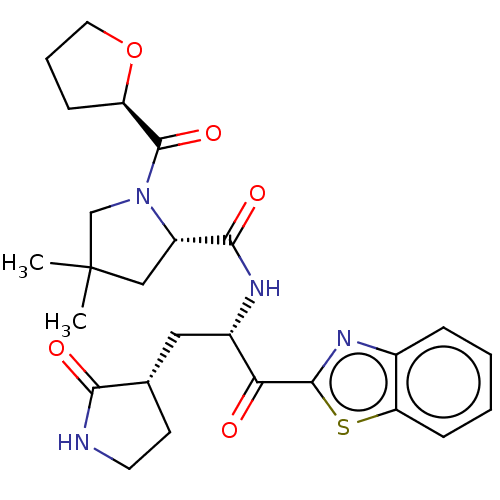
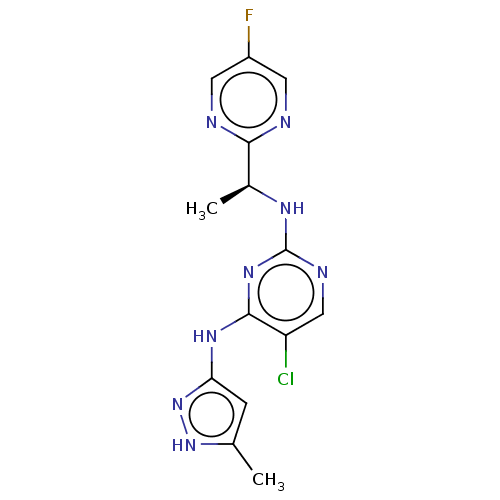
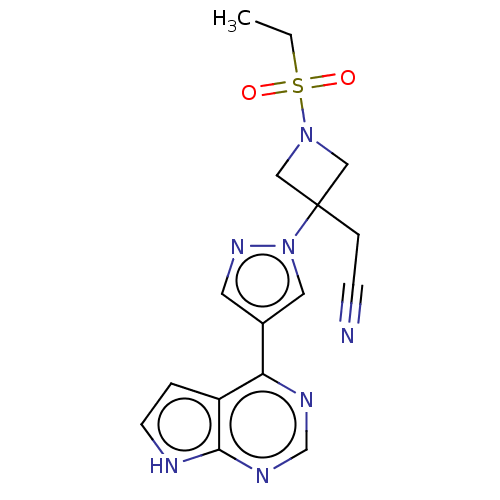
Affinity DataKi: 1.55nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
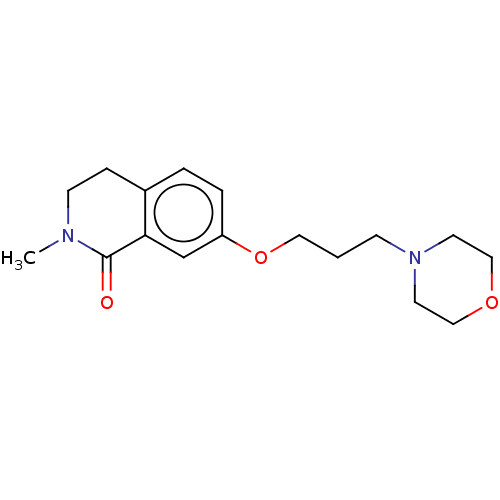
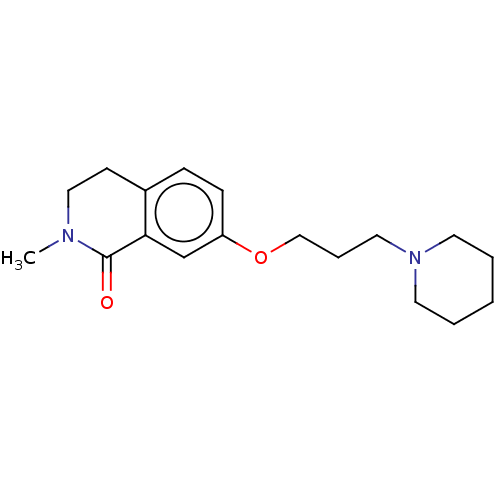
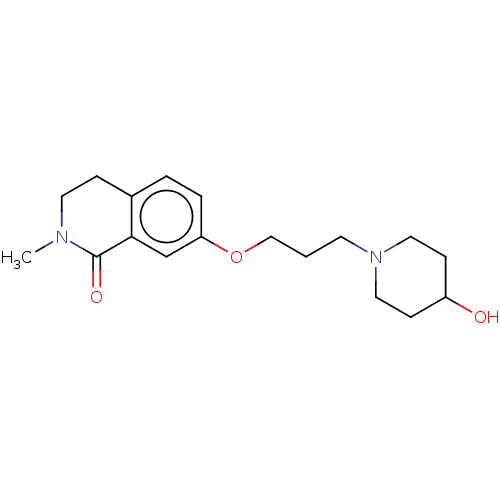
Affinity DataKi: 3.60nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
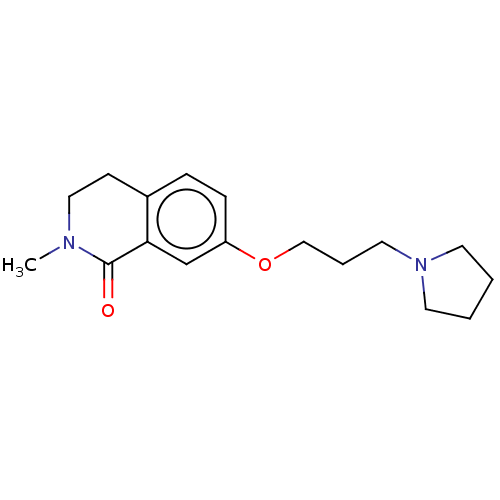
Affinity DataKi: 6.02nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
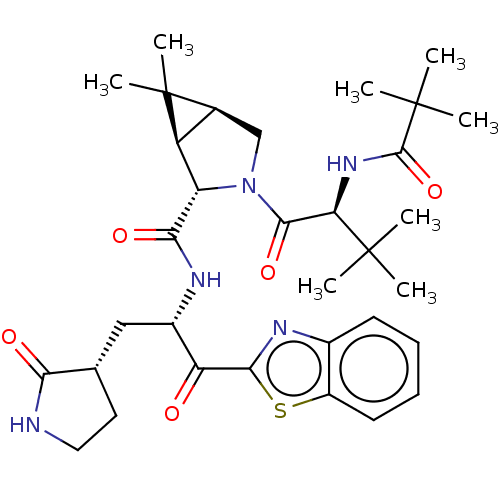
Affinity DataKi: 6.14nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
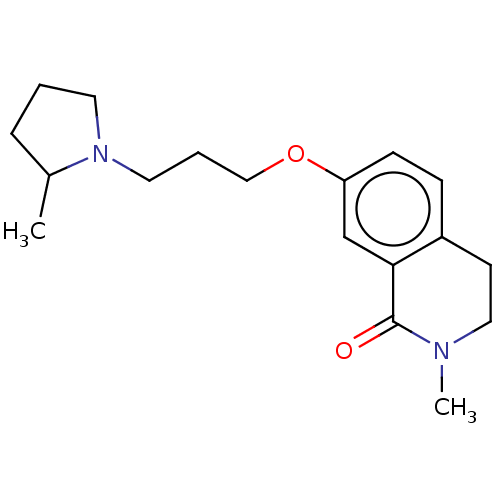
Affinity DataKi: 6.5nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
Affinity DataKi: 7.74nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 9.77nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 23nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 26nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 32nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
Affinity DataKi: 34nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 35nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 39nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
TargetHistamine H3 receptor(Rattus norvegicus (rat))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 68nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from histamine H3 receptor in rat cerebral cortex homogenates incubated for 60 mins by Cheng and Prusoff...More data for this Ligand-Target Pair
Affinity DataKi: 99nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 112nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
Affinity DataKi: 137nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: 139nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 165nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
TargetHistamine H3 receptor(Rattus norvegicus (rat))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 169nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from histamine H3 receptor in rat cerebral cortex homogenates incubated for 60 mins by Cheng and Prusoff...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 176nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
TargetHistamine H3 receptor(Rattus norvegicus (rat))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 189nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from histamine H3 receptor in rat cerebral cortex homogenates incubated for 60 mins by Cheng and Prusoff...More data for this Ligand-Target Pair
Affinity DataKi: 206nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 252nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
TargetHistamine H3 receptor(Rattus norvegicus (rat))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 279nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from histamine H3 receptor in rat cerebral cortex homogenates incubated for 60 mins by Cheng and Prusoff...More data for this Ligand-Target Pair
TargetHistamine H3 receptor(Rattus norvegicus (rat))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 426nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from histamine H3 receptor in rat cerebral cortex homogenates incubated for 60 mins by Cheng and Prusoff...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 542nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 706nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 856nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
TargetHistamine H3 receptor(Rattus norvegicus (rat))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 908nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from histamine H3 receptor in rat cerebral cortex homogenates incubated for 60 mins by Cheng and Prusoff...More data for this Ligand-Target Pair
TargetHistamine H3 receptor(Rattus norvegicus (rat))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 1.03E+3nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from histamine H3 receptor in rat cerebral cortex homogenates incubated for 60 mins by Cheng and Prusoff...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 1.10E+3nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: 1.21E+3nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
TargetHistamine H1 receptor(Cavia porcellus (domestic guinea pig))
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Jiangsu Marine Resources Development Research Institute
Curated by ChEMBL
Affinity DataKi: >2.00E+3nMAssay Description:Displacement of [3H]-mepyramine from histamine H1 receptor in guinea pig cerebellum homogenates incubated for 60 mins by Cheng and Prusoff equation a...More data for this Ligand-Target Pair
Affinity DataKi: 6.33E+3nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataKi: >1.08E+4nMAssay Description:The proteolytic activity of the main protease, 3CLpro, of SARS-CoV-2 was monitored using a continuous fluorescence resonance energy transfer (FRET) a...More data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMpH: 7.5Assay Description:A kinase selectivity panel which measures substrate peptide phosphorylation was set-up for recombinant human Jak1 (aa 866-1154), Jak2 (aa808-1132), J...More data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:The assays were performed in 384-well, low volume microtiter assay plates in a final reaction volume of 9 ul. Dose-response curves were generated by ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:The assays were performed in 384-well, low volume microtiter assay plates in a final reaction volume of 9 ul. Dose-response curves were generated by ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.700nMpH: 7.5Assay Description:A kinase selectivity panel which measures substrate peptide phosphorylation was set-up for recombinant human Jak1 (aa 866-1154), Jak2 (aa808-1132), J...More data for this Ligand-Target Pair
Affinity DataIC50: 0.700nMpH: 7.5Assay Description:A kinase selectivity panel which measures substrate peptide phosphorylation was set-up for recombinant human Jak1 (aa 866-1154), Jak2 (aa808-1132), J...More data for this Ligand-Target Pair
Affinity DataIC50: 0.700nMAssay Description:The assays were performed in 384-well, low volume microtiter assay plates in a final reaction volume of 9 ul. Dose-response curves were generated by ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.700nMAssay Description:The assays were performed in 384-well, low volume microtiter assay plates in a final reaction volume of 9 ul. Dose-response curves were generated by ...More data for this Ligand-Target Pair