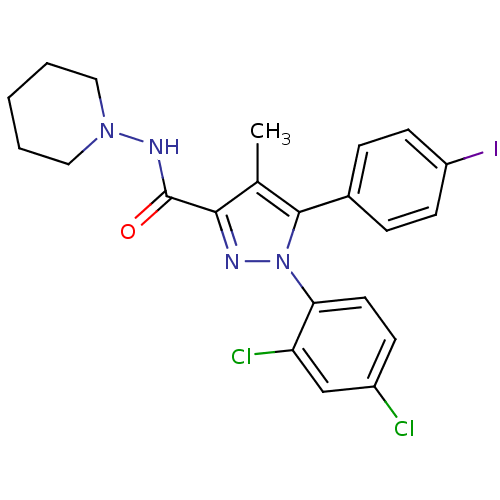
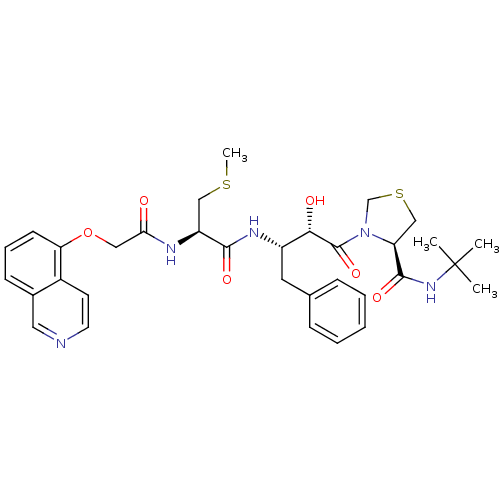
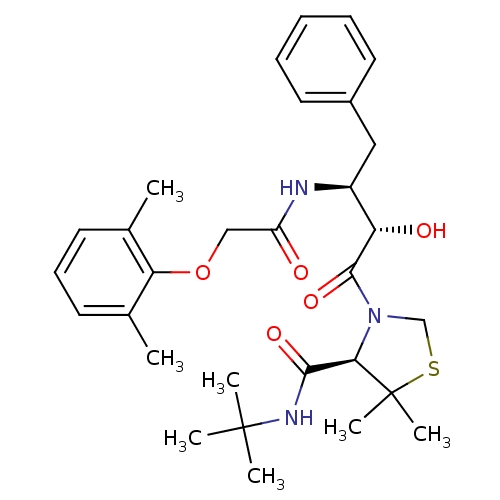
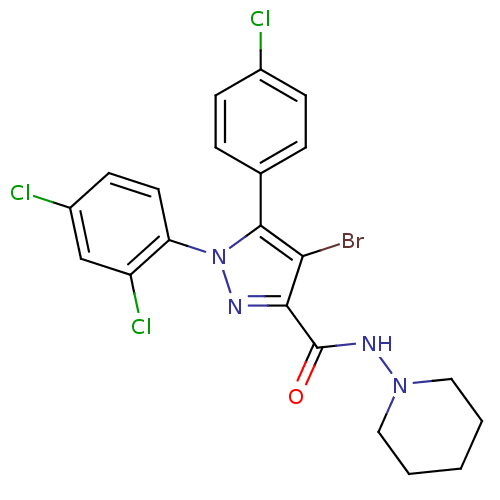
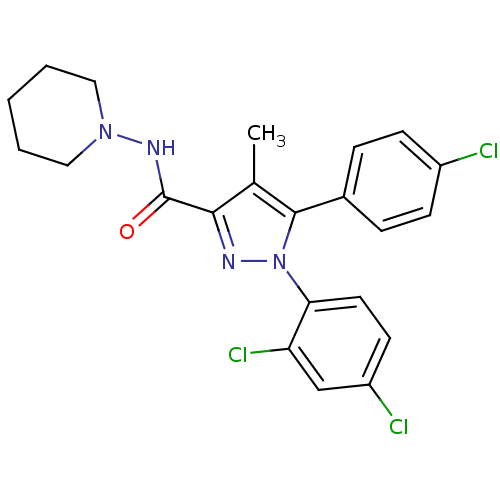
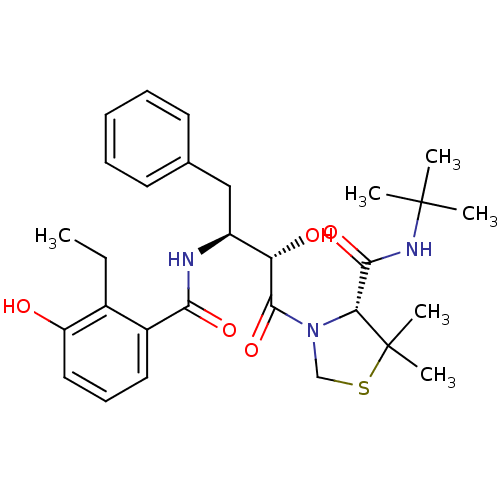
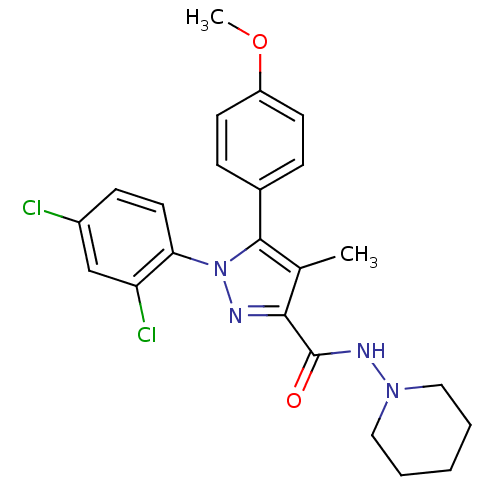
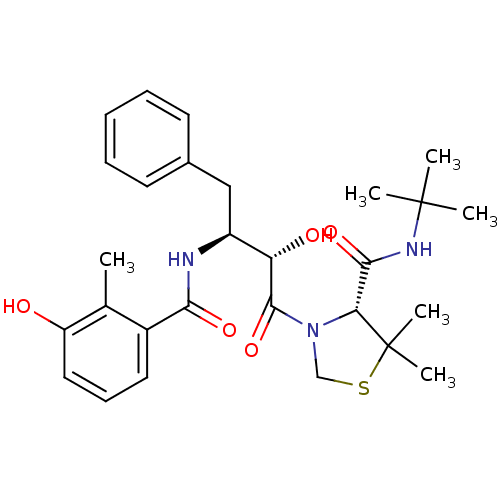
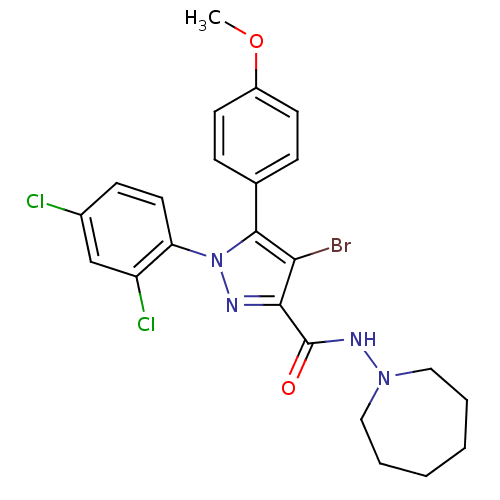
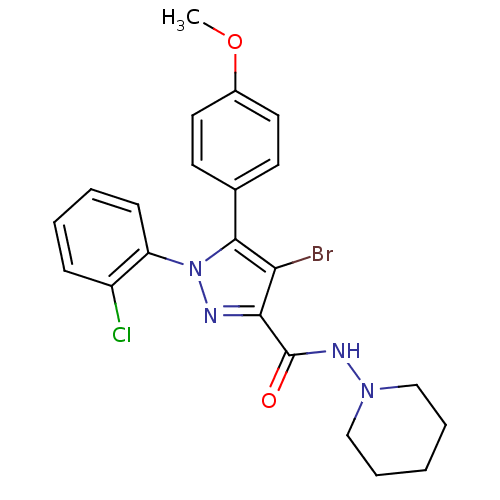
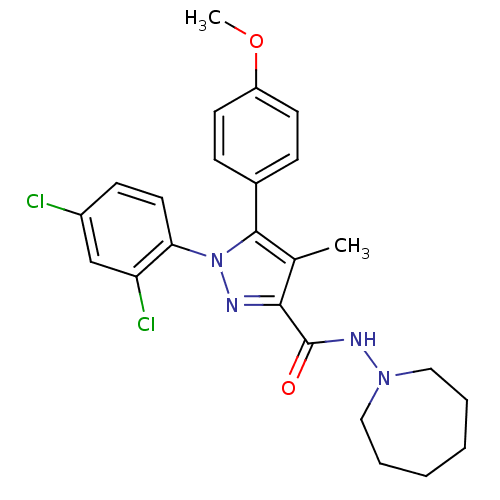
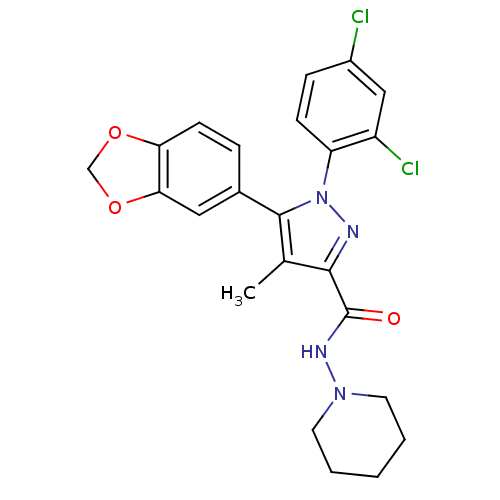
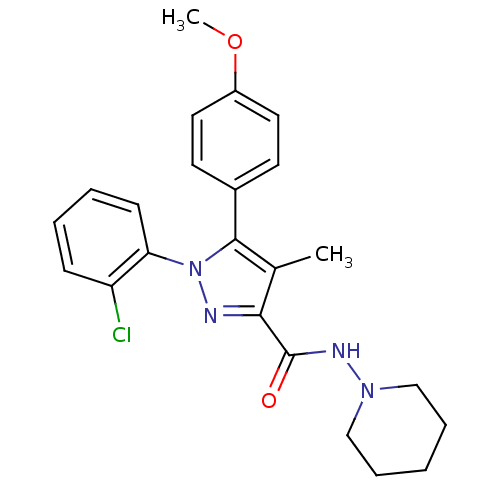
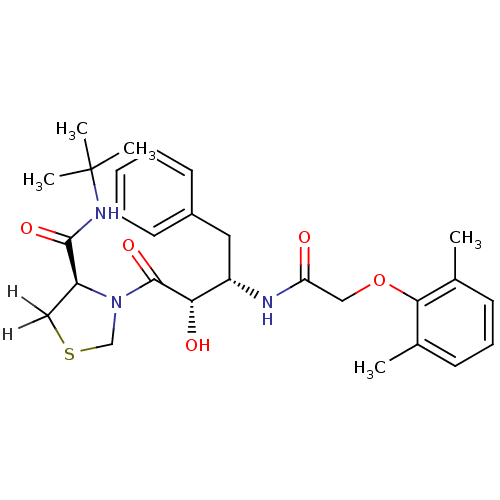
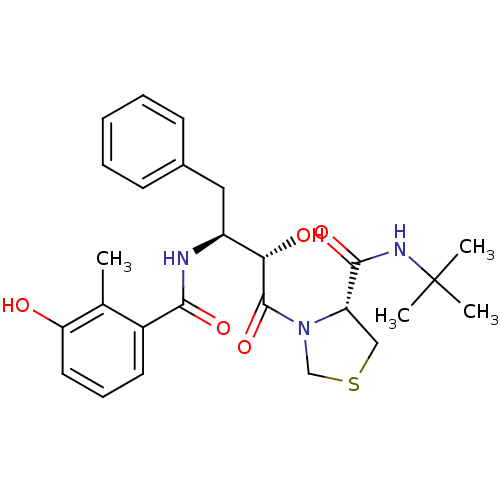
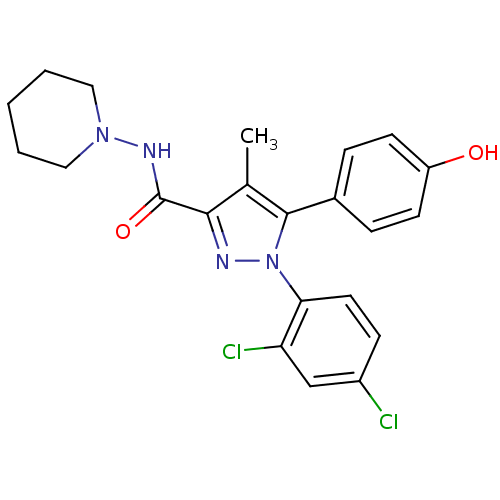
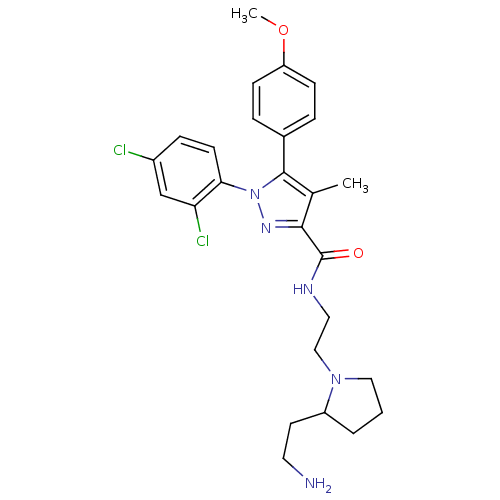
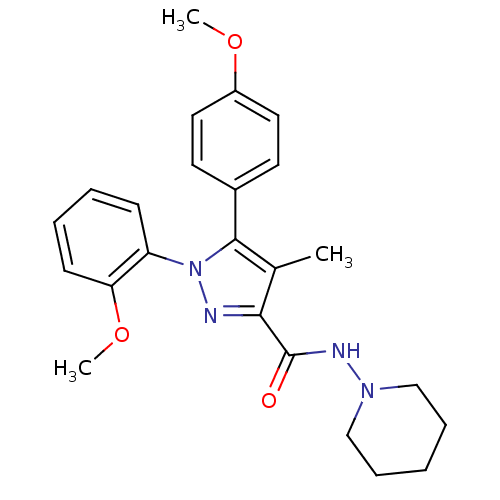
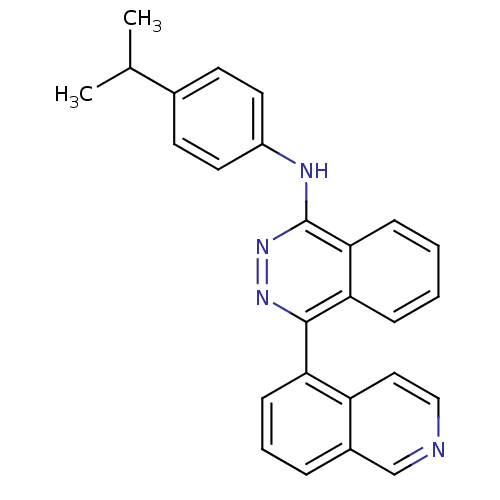
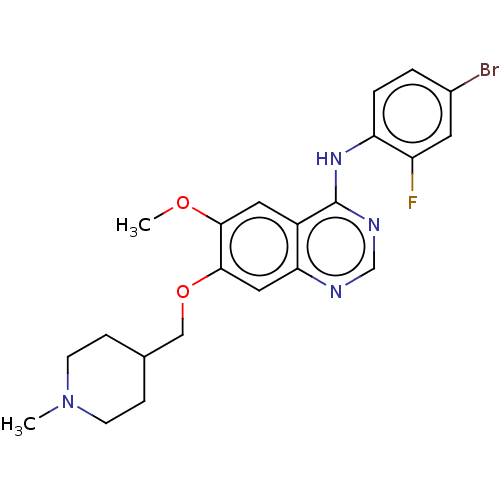
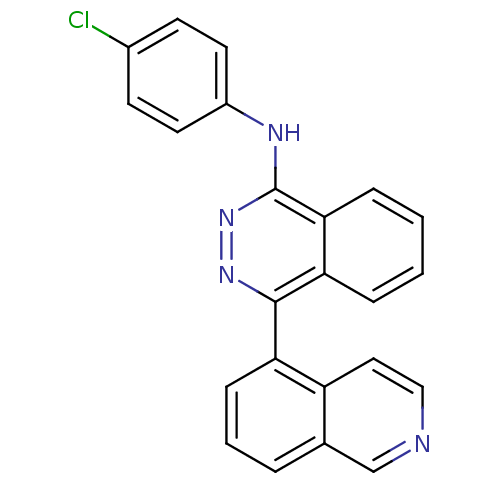
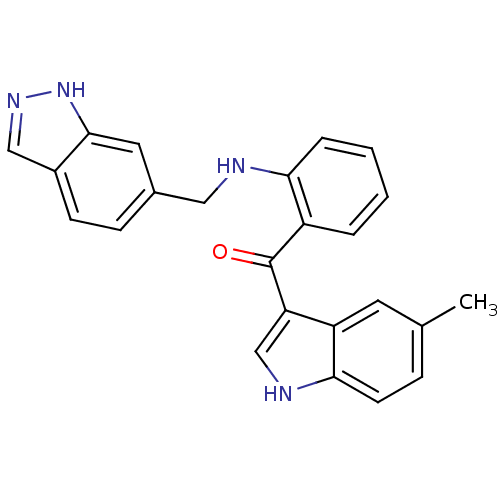
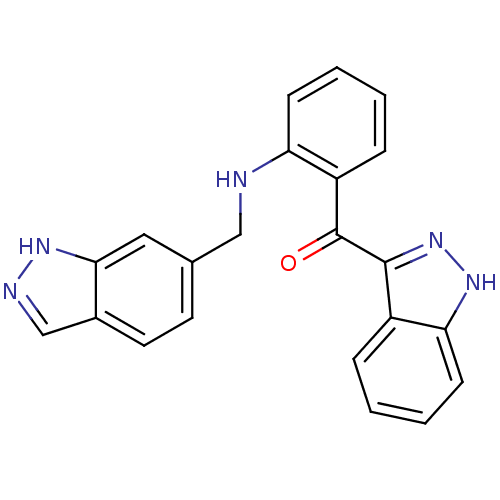
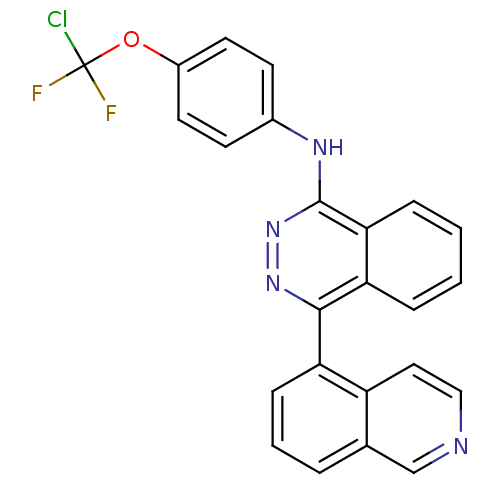
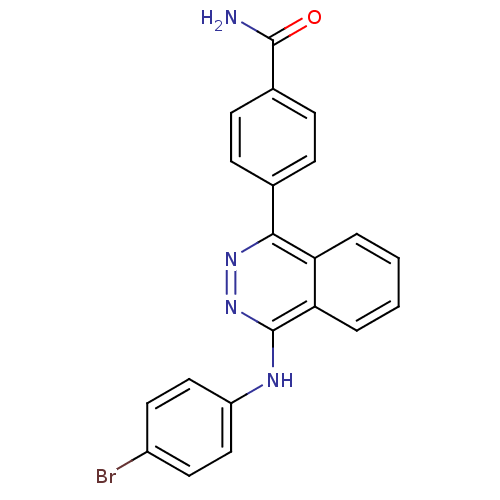
Affinity DataKi: 0.0880nM ΔG°: -59.7kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
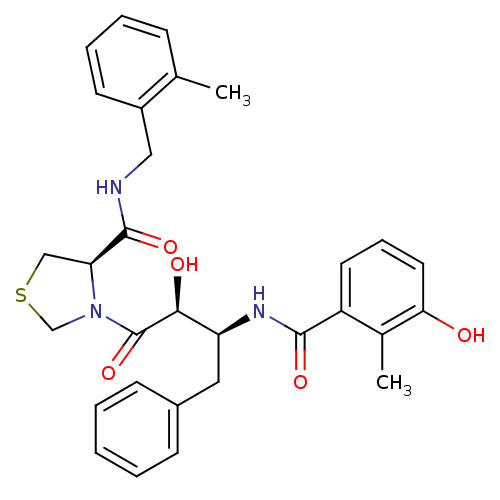
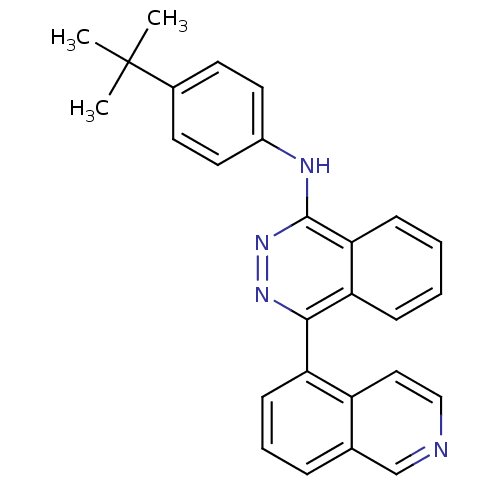
Affinity DataKi: 0.160nM ΔG°: -58.2kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
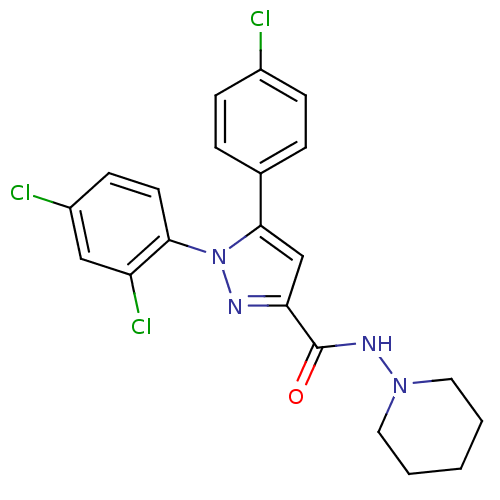
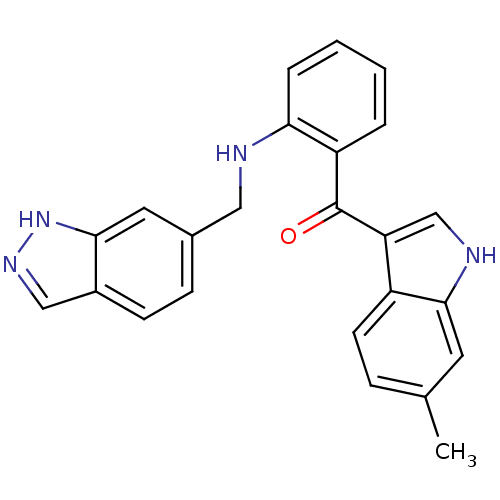
Affinity DataKi: 0.290nM ΔG°: -56.6kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
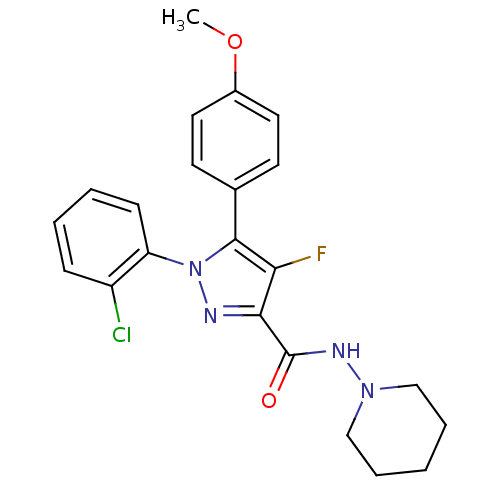
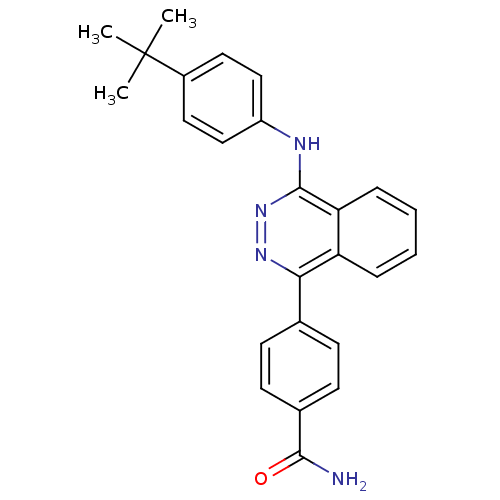
Affinity DataKi: 0.330nM ΔG°: -56.3kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 0.600nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
Affinity DataKi: 0.740nM ΔG°: -54.2kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 1.40nM ΔG°: -52.6kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 1.40nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 1.80nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
Affinity DataKi: 2.24nM ΔG°: -51.4kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 4.10nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
Affinity DataKi: 5.14nM ΔG°: -49.2kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 5.20nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 6.20nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 7nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 7.10nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 8nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
Affinity DataKi: 8.91nM ΔG°: -47.8kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 9nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 18.2nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
Affinity DataKi: 21.7nM ΔG°: -45.5kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
Affinity DataKi: 24.9nM ΔG°: -45.1kJ/molepH: 6.0 T: 2°CAssay Description:Sensitivity of HIV-1 protease activity to protease inhibitors was determined by a peptide substrate cleavage assay. Substrates and cleavage fragments...More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 37.8nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 104nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 380nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
TargetCannabinoid receptor 1(Rattus norvegicus (rat))
National Institute On Drug Abuse
Curated by ChEMBL
National Institute On Drug Abuse
Curated by ChEMBL
Affinity DataKi: 440nMAssay Description:Binding affinity was determined by using a competition assay with [125 I]- AM251 against rat cannabinoid receptor 1More data for this Ligand-Target Pair
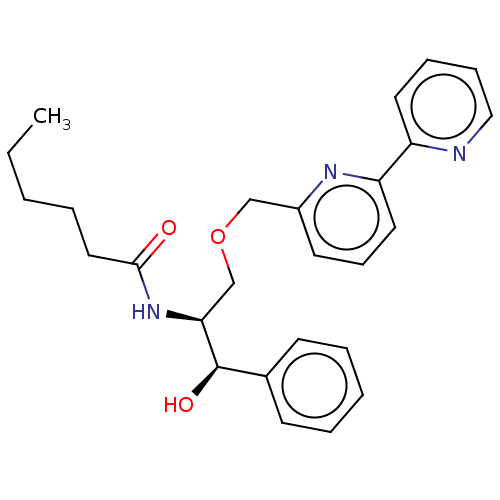
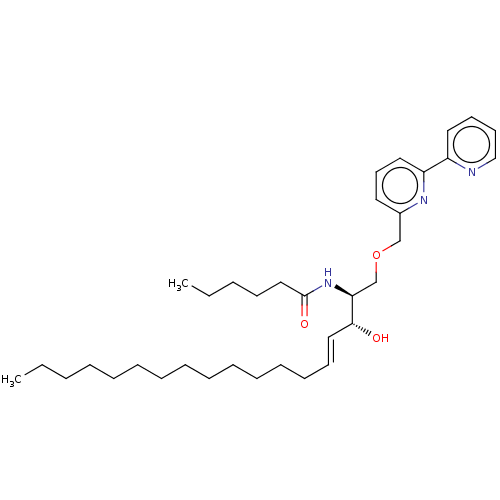
Affinity DataKi: 1.30E+3nM ΔG°: -34.9kJ/mole IC50: 800nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
Affinity DataKi: 2.80E+3nM ΔG°: -33.0kJ/mole IC50: 900nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
Affinity DataKi: 5.20E+3nM ΔG°: -31.4kJ/mole IC50: 1.20E+3nMpH: 7.5 T: 2°CAssay Description:Briefly, Bc-SMase (50 ng/ml) was treated at 37 °C for 60 min with various compounds that were solubilized in dimethylacetoamide. The reaction mix...More data for this Ligand-Target Pair
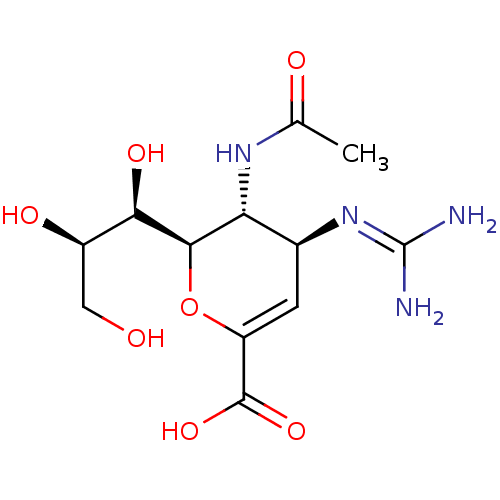
Affinity DataKi: 1.70E+4nMAssay Description:Inhibition of human neuraminidase 2More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+5nMAssay Description:Inhibition of human neuraminidase 2More data for this Ligand-Target Pair
Affinity DataKi: 3.30E+5nMAssay Description:Inhibition of human neuraminidase 2More data for this Ligand-Target Pair
Affinity DataKi: 7.40E+5nMAssay Description:Inhibition of human neuraminidase 2More data for this Ligand-Target Pair
Affinity DataKi: 8.80E+5nMAssay Description:Inhibition of human neuraminidase 2More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+6nMAssay Description:Inhibition of human neuraminidase 2More data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 17nMAssay Description:Inhibition of VEGFR2 by HTRF assayMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 17nMAssay Description:Inhibition of VEGFR2 (unknown origin) by HTRF methodMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 45nMAssay Description:Inhibition of VEGFR2 (unknown origin) by HTRF methodMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 48nMAssay Description:Inhibition of VEGFR2 (unknown origin) by HTRF methodMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 48nMAssay Description:Inhibition of VEGFR2 by HTRF assayMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 52nMAssay Description:Inhibition of KDR (unknown origin) autophosphorylation by cell based assayMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 74nMAssay Description:Inhibition of KDR (unknown origin) autophosphorylation by cell based assayMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 75nMAssay Description:Inhibition of VEGFR2 by HTRF assayMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 75nMAssay Description:Inhibition of VEGFR2 (unknown origin) by HTRF methodMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 76nMAssay Description:Inhibition of KDR (unknown origin) autophosphorylation by cell based assayMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 78nMAssay Description:Inhibition of VEGFR2 (unknown origin) by HTRF methodMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 78nMAssay Description:Inhibition of Vascular endothelial growth factor receptor 2 kinase phosphorylation of pGAT-biotin peptideMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 79nMAssay Description:Inhibition of VEGFR2 by HTRF assayMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 79nMAssay Description:Inhibition of VEGFR2 (unknown origin) by HTRF methodMore data for this Ligand-Target Pair
TargetVascular endothelial growth factor receptor 2(Homo sapiens (Human))
Imclone Systems
Curated by ChEMBL
Imclone Systems
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibition of Vascular endothelial growth factor receptor 2 kinase phosphorylation of pGAT-biotin peptideMore data for this Ligand-Target Pair

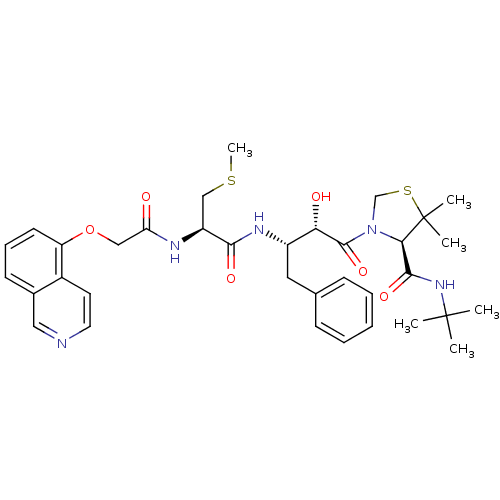
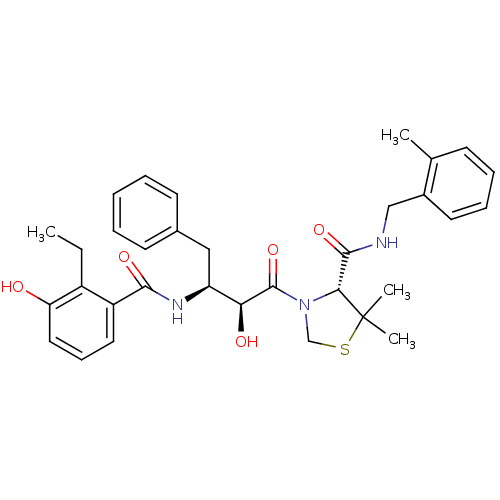
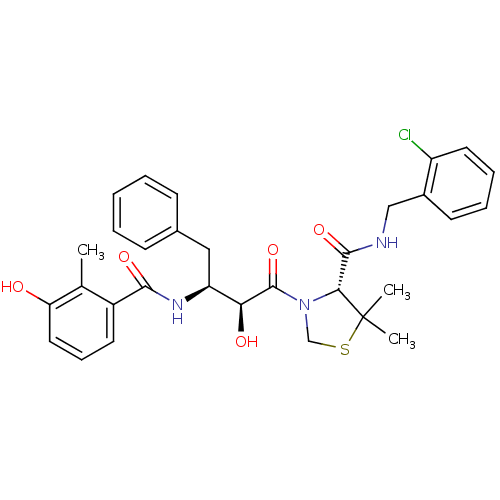
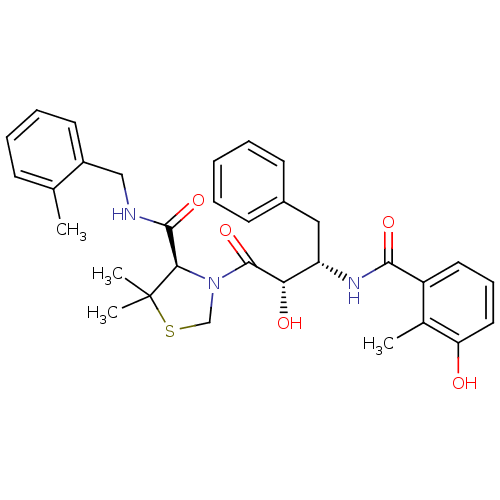
 3D Structure (crystal)
3D Structure (crystal)