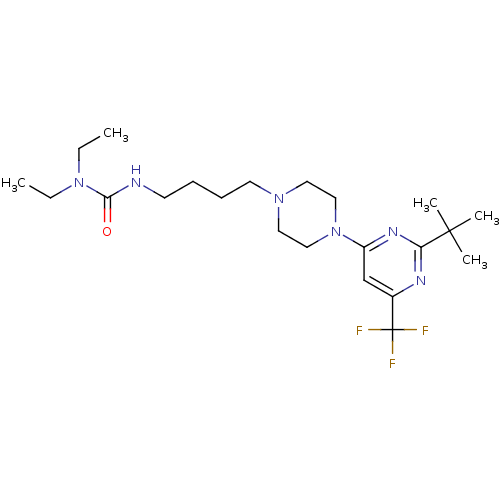
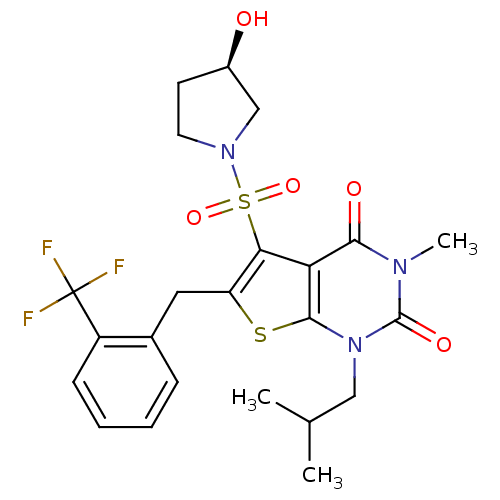
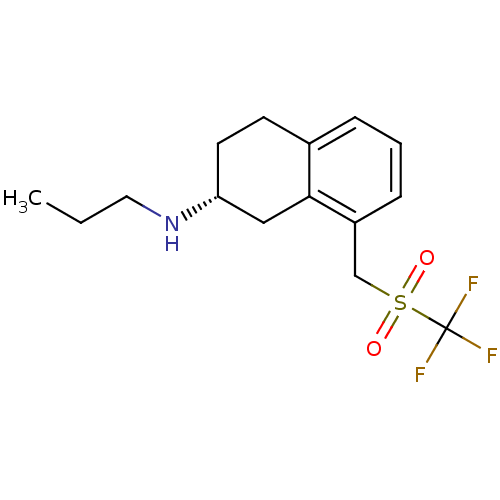
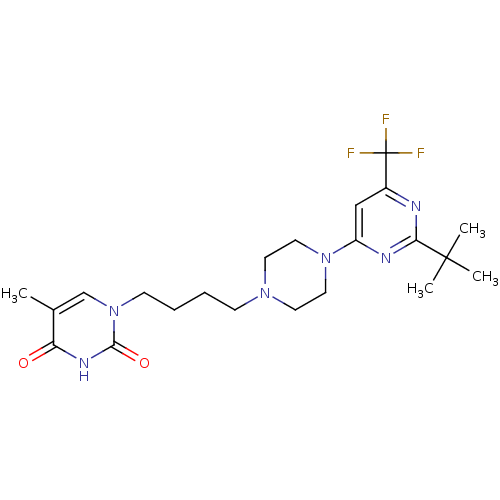
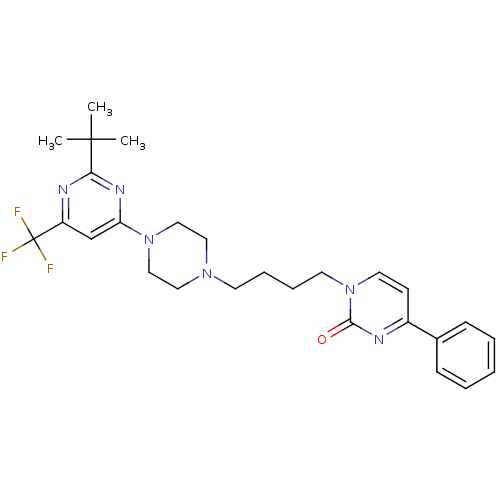
TargetNeuronal acetylcholine receptor subunit alpha-4(Rattus norvegicus (Rat))
National Institute On Drug Abuse
Curated by PDSP Ki Database
National Institute On Drug Abuse
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(RAT)
National Institute On Drug Abuse
Curated by PDSP Ki Database
National Institute On Drug Abuse
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(RAT)
National Institute On Drug Abuse
Curated by PDSP Ki Database
National Institute On Drug Abuse
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(RAT)
National Institute On Drug Abuse
Curated by PDSP Ki Database
National Institute On Drug Abuse
Curated by PDSP Ki Database
TargetNeuronal acetylcholine receptor subunit alpha-4(Rattus norvegicus (Rat))
National Institute On Drug Abuse
Curated by PDSP Ki Database
National Institute On Drug Abuse
Curated by PDSP Ki Database
TargetNeuronal acetylcholine receptor subunit alpha-4(Rattus norvegicus (Rat))
National Institute On Drug Abuse
Curated by PDSP Ki Database
National Institute On Drug Abuse
Curated by PDSP Ki Database
TargetNeuronal acetylcholine receptor subunit alpha-4(Rattus norvegicus (Rat))
National Institute On Drug Abuse
Curated by PDSP Ki Database
National Institute On Drug Abuse
Curated by PDSP Ki Database
Affinity DataKi: 0.0300nMAssay Description:Proteolytic activity of SARS-CoV-2 Coronavirus 3CL protease is measured using a continuous fluorescence resonance energy transfer assay. The SARS-CoV...More data for this Ligand-Target Pair
Affinity DataKi: 0.0300nMAssay Description:Proteolytic activity of SARS-CoV-2 Coronavirus 3CL protease is measured using a continuous fluorescence resonance energy transfer assay. The SARS-CoV...More data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3(Rattus norvegicus (Rat))
National Institute On Drug Abuse
Curated by PDSP Ki Database
National Institute On Drug Abuse
Curated by PDSP Ki Database
Affinity DataKi: 0.0955nMAssay Description:Displacement of [3H]5-[(3-hydroxypropyl)thio]-3-methyl-1-[2-(methyl-t)propyl-2,3-t2]-6-(1-naphthalenylmethyl)-1H-pyrrolo[3,4-d]pyrimidine-2,4(3H,6H)-...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:Proteolytic activity of SARS-CoV-2 Coronavirus 3CL protease is measured using a continuous fluorescence resonance energy transfer assay. The SARS-CoV...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -56.5kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3(Rattus norvegicus (Rat))
National Institute On Drug Abuse
Curated by PDSP Ki Database
National Institute On Drug Abuse
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(RAT)
National Institute On Drug Abuse
Curated by PDSP Ki Database
National Institute On Drug Abuse
Curated by PDSP Ki Database
TargetNeuronal acetylcholine receptor subunit alpha-4(Rattus norvegicus (Rat))
National Institute On Drug Abuse
Curated by PDSP Ki Database
National Institute On Drug Abuse
Curated by PDSP Ki Database
Affinity DataKi: 0.275nMAssay Description:Displacement of [3H]5-[(3-hydroxypropyl)thio]-3-methyl-1-[2-(methyl-t)propyl-2,3-t2]-6-(1-naphthalenylmethyl)-1H-pyrrolo[3,4-d]pyrimidine-2,4(3H,6H)-...More data for this Ligand-Target Pair
Affinity DataKi: 0.280nM ΔG°: -54.0kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 0.290nM ΔG°: -53.9kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Proteolytic activity of SARS-CoV-2 Coronavirus 3CL protease is measured using a continuous fluorescence resonance energy transfer assay. The SARS-CoV...More data for this Ligand-Target Pair
Affinity DataKi: 0.302nMAssay Description:Displacement of [3H]5-[(3-hydroxypropyl)thio]-3-methyl-1-[2-(methyl-t)propyl-2,3-t2]-6-(1-naphthalenylmethyl)-1H-pyrrolo[3,4-d]pyrimidine-2,4(3H,6H)-...More data for this Ligand-Target Pair
Affinity DataKi: 0.330nMAssay Description:Displacement of [3H]5-[(3-hydroxypropyl)thio]-3-methyl-1-[2-(methyl-t)propyl-2,3-t2]-6-(1-naphthalenylmethyl)-1H-pyrrolo[3,4-d]pyrimidine-2,4(3H,6H)-...More data for this Ligand-Target Pair
Affinity DataKi: 0.330nM ΔG°: -53.6kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 0.331nMAssay Description:Displacement of [3H]5-[(3-hydroxypropyl)thio]-3-methyl-1-[2-(methyl-t)propyl-2,3-t2]-6-(1-naphthalenylmethyl)-1H-pyrrolo[3,4-d]pyrimidine-2,4(3H,6H)-...More data for this Ligand-Target Pair
Affinity DataKi: 0.350nM ΔG°: -53.4kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 0.355nMAssay Description:Displacement of [3H]5-[(3-hydroxypropyl)thio]-3-methyl-1-[2-(methyl-t)propyl-2,3-t2]-6-(1-naphthalenylmethyl)-1H-pyrrolo[3,4-d]pyrimidine-2,4(3H,6H)-...More data for this Ligand-Target Pair
Affinity DataKi: 0.355nMAssay Description:Displacement of [3H]5-[(3-hydroxypropyl)thio]-3-methyl-1-[2-(methyl-t)propyl-2,3-t2]-6-(1-naphthalenylmethyl)-1H-pyrrolo[3,4-d]pyrimidine-2,4(3H,6H)-...More data for this Ligand-Target Pair
Affinity DataKi: 0.420nM ΔG°: -53.0kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:In vitro binding affinity towards 5-hydroxytryptamine 1A receptor by using [3H]-8-OH-DPAT as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Proteolytic activity of SARS-CoV-2 Coronavirus 3CL protease is measured using a continuous fluorescence resonance energy transfer assay. The SARS-CoV...More data for this Ligand-Target Pair
Affinity DataKi: 0.680nM ΔG°: -51.8kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 0.708nMAssay Description:Displacement of [3H]5-[(3-hydroxypropyl)thio]-3-methyl-1-[2-(methyl-t)propyl-2,3-t2]-6-(1-naphthalenylmethyl)-1H-pyrrolo[3,4-d]pyrimidine-2,4(3H,6H)-...More data for this Ligand-Target Pair
Affinity DataKi: 0.790nM ΔG°: -51.4kJ/molepH: 7.4 T: 2°CAssay Description:A radioligand-binding assay was developed using scintillation proximity assay (SPA) technology. The wheat germ agglutinin SPA beads (Amersham) (0.2 m...More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Binding affinity to human cloned dopamine D3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Binding affinity to cloned human dopamine D3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Proteolytic activity of SARS-CoV-2 Coronavirus 3CL protease is measured using a continuous fluorescence resonance energy transfer assay. The SARS-CoV...More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Proteolytic activity of SARS-CoV-2 Coronavirus 3CL protease is measured using a continuous fluorescence resonance energy transfer assay. The SARS-CoV...More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:In vitro binding affinity towards 5-hydroxytryptamine 1A receptor by using [3H]-8-OH-DPAT as radioligandMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-4(Rattus norvegicus (Rat))
National Institute On Drug Abuse
Curated by PDSP Ki Database
National Institute On Drug Abuse
Curated by PDSP Ki Database
Affinity DataKi: 0.900nMAssay Description:Binding affinity to cloned human dopamine D3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.900nMAssay Description:Proteolytic activity of SARS-CoV-2 Coronavirus 3CL protease is measured using a continuous fluorescence resonance energy transfer assay. The SARS-CoV...More data for this Ligand-Target Pair
TargetNicotinic acetylcholine receptor(RAT)
National Institute On Drug Abuse
Curated by PDSP Ki Database
National Institute On Drug Abuse
Curated by PDSP Ki Database
Affinity DataKi: 1.10nM ΔG°: -50.6kJ/molepH: 7.8 T: 2°CAssay Description:Jurkat cell membranes were incubated with [3H]-labeled ligand in the absence or presence of increasing concentrations of test compound. The reagents ...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Binding affinity to cloned human dopamine D3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 1.17nMAssay Description:Displacement of [3H]5-[(3-hydroxypropyl)thio]-3-methyl-1-[2-(methyl-t)propyl-2,3-t2]-6-(1-naphthalenylmethyl)-1H-pyrrolo[3,4-d]pyrimidine-2,4(3H,6H)-...More data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Proteolytic activity of SARS-CoV-2 Coronavirus 3CL protease is measured using a continuous fluorescence resonance energy transfer assay. The SARS-CoV...More data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:In vitro binding affinity towards 5-hydroxytryptamine 1A receptor by using [3H]-8-OH-DPAT as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Binding affinity to human cloned dopamine D3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Binding affinity to human cloned dopamine D3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Binding affinity to cloned human dopamine D3 receptorMore data for this Ligand-Target Pair

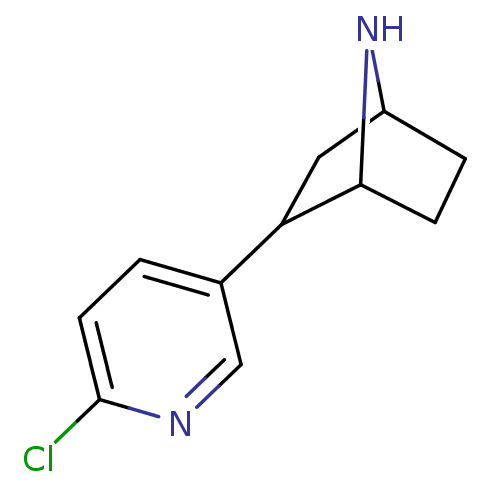
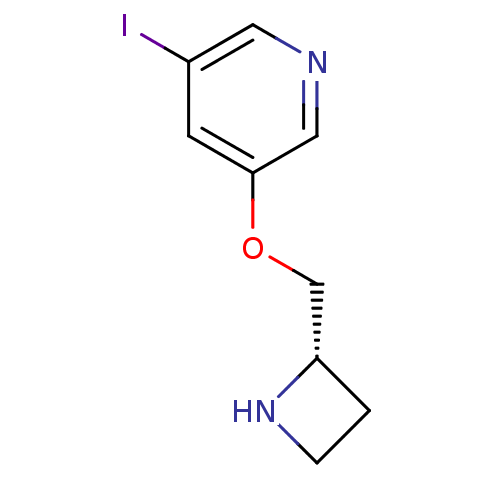
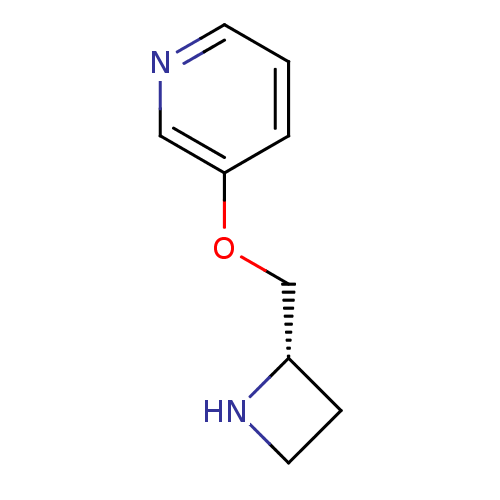
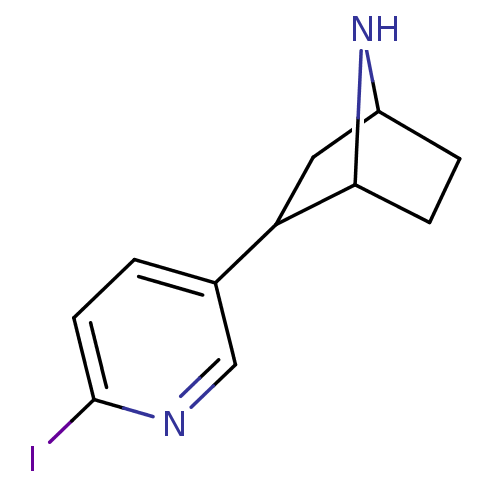
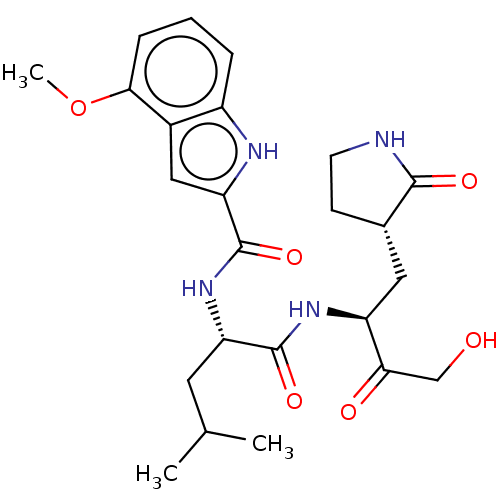
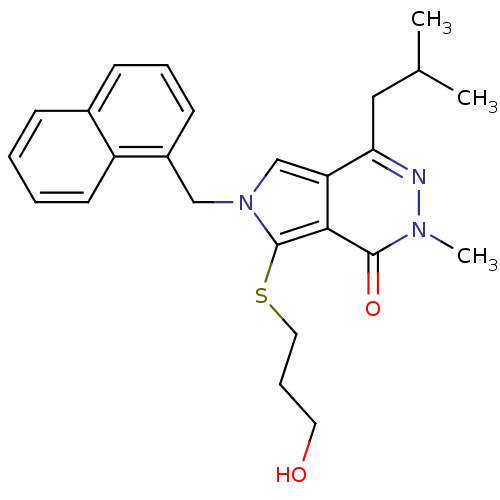
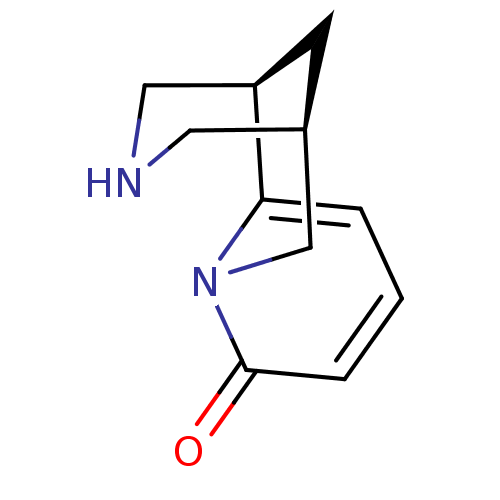
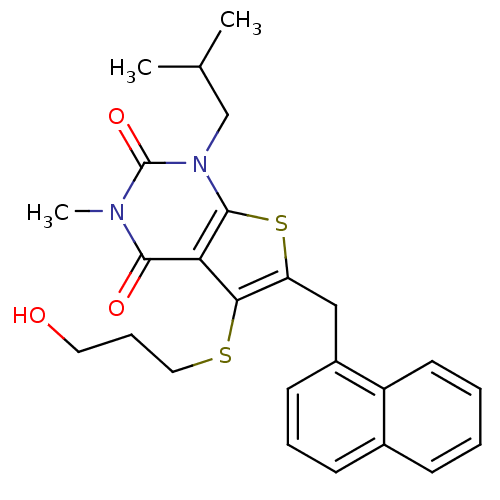
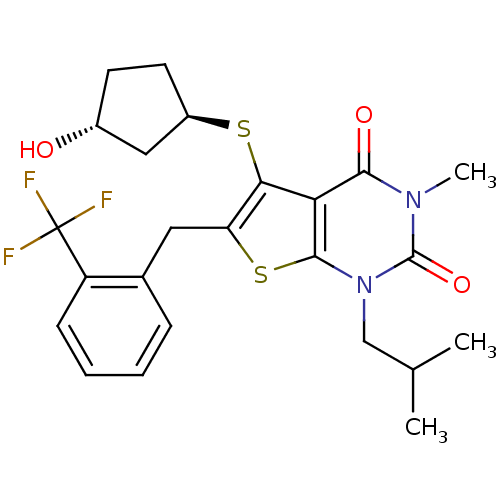
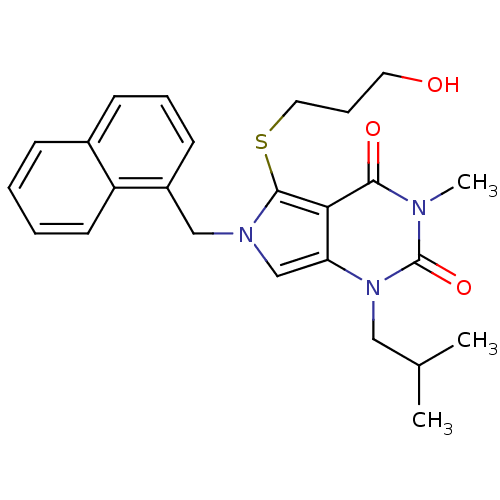
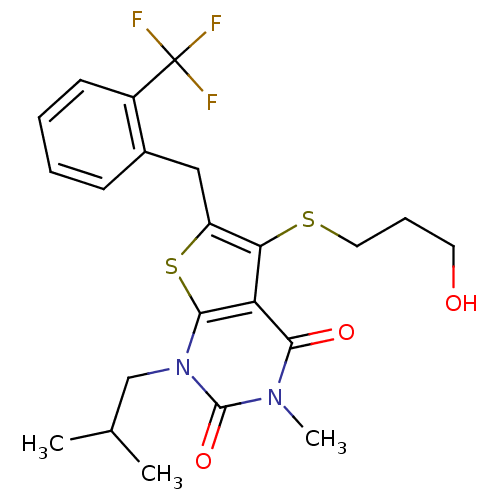
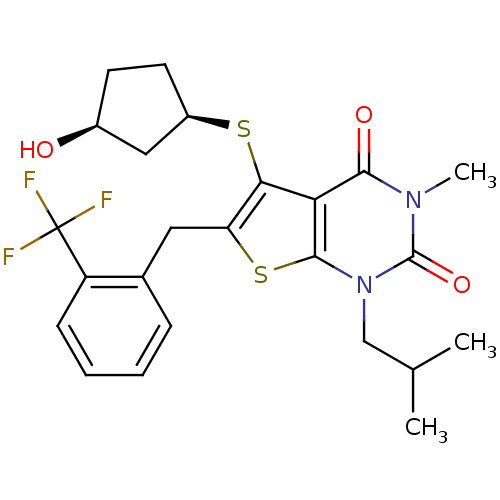
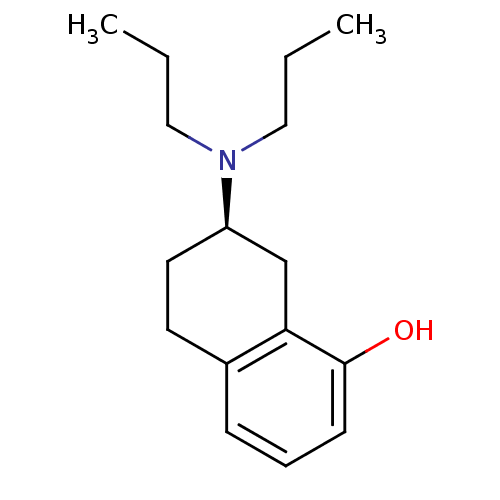
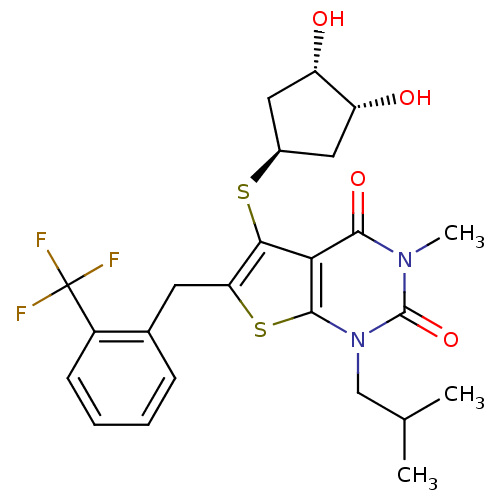
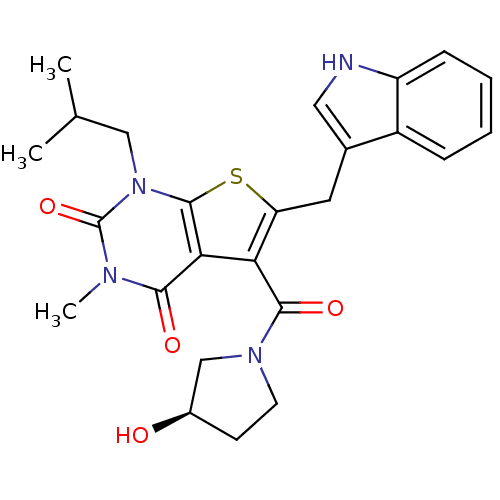
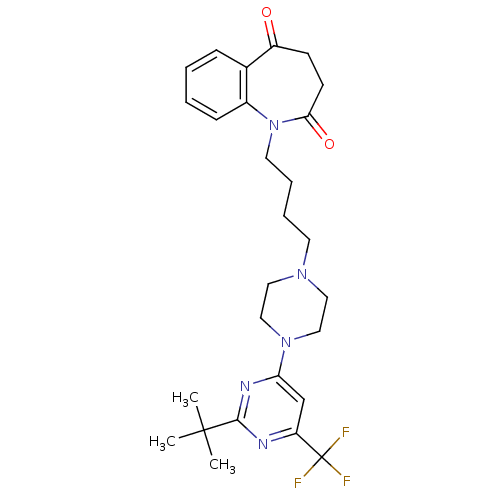
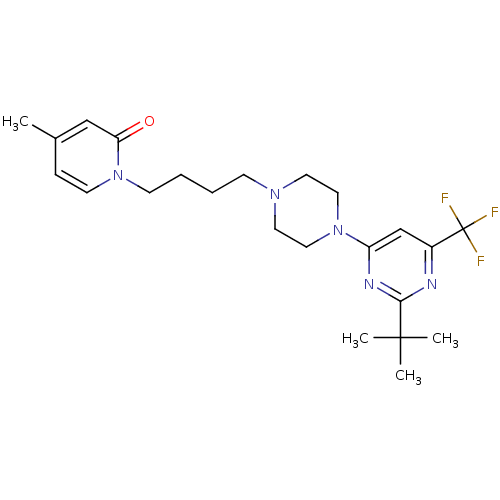
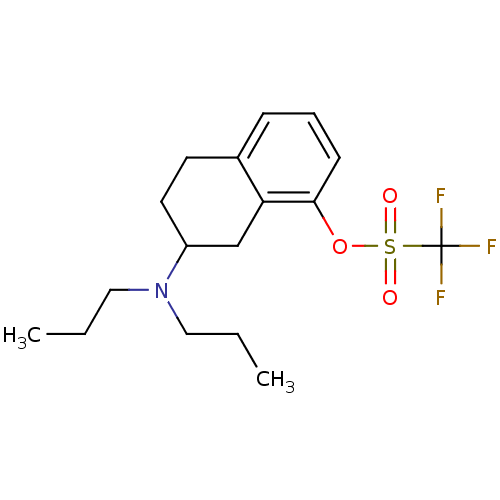
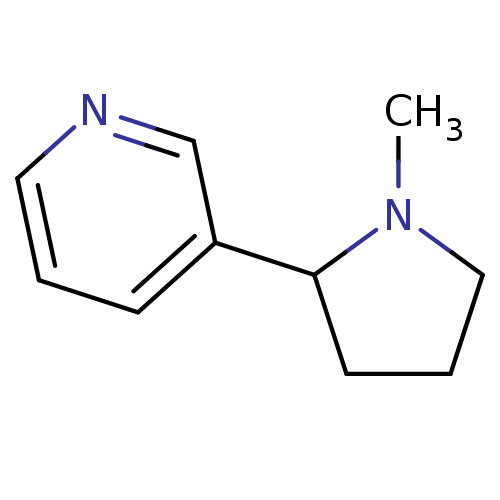
 3D Structure (crystal)
3D Structure (crystal)