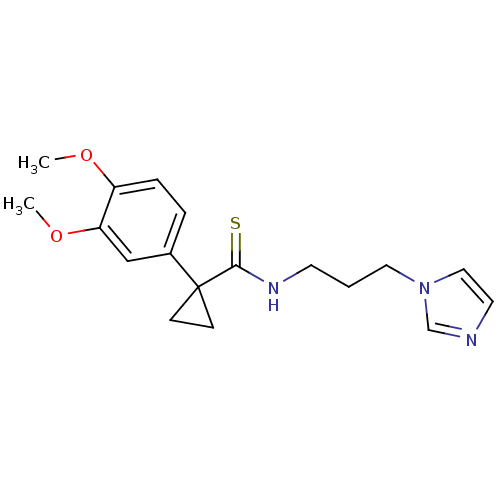
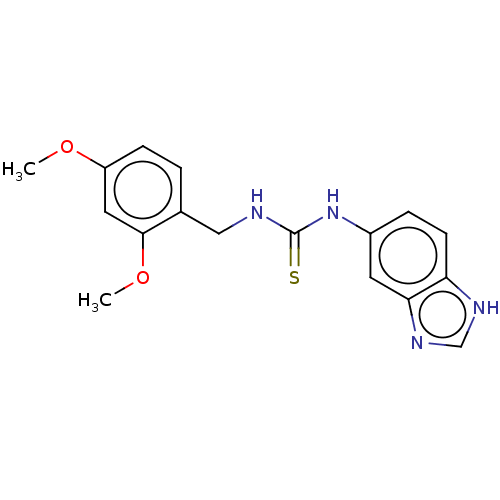
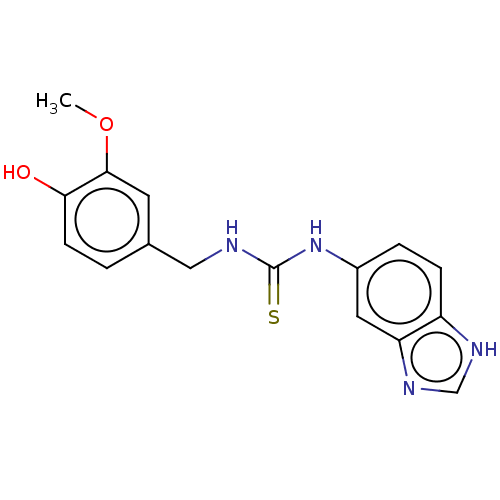
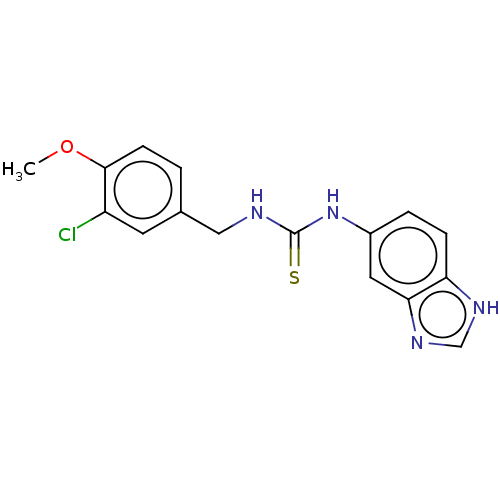
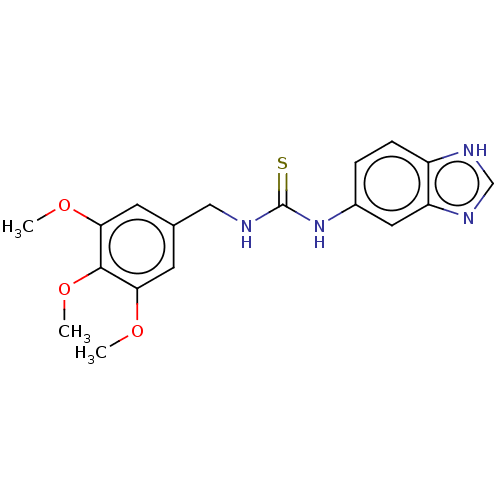
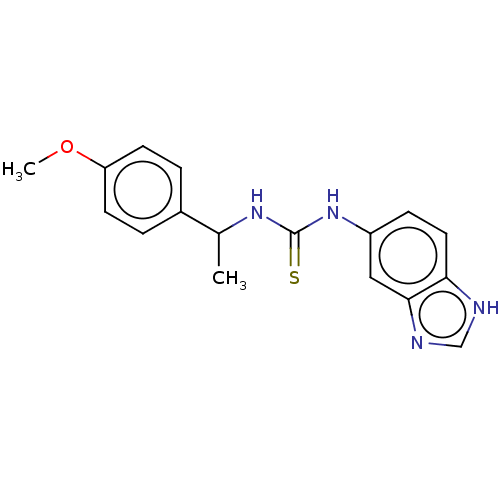
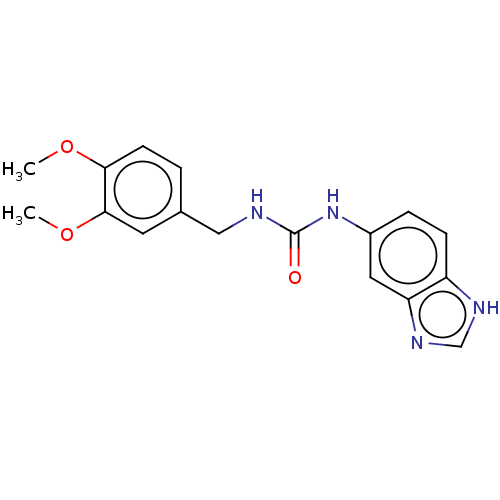
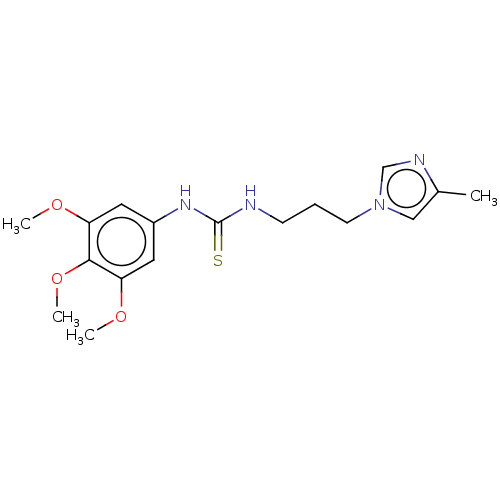
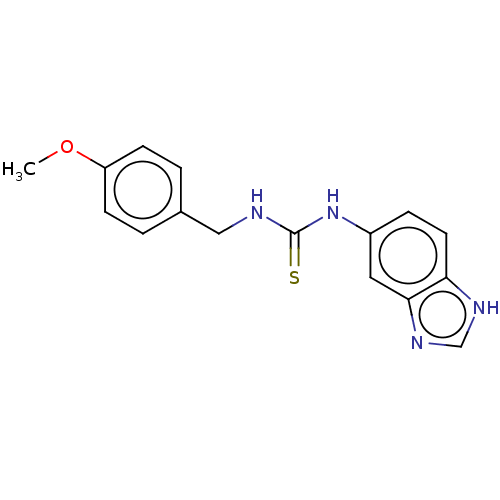
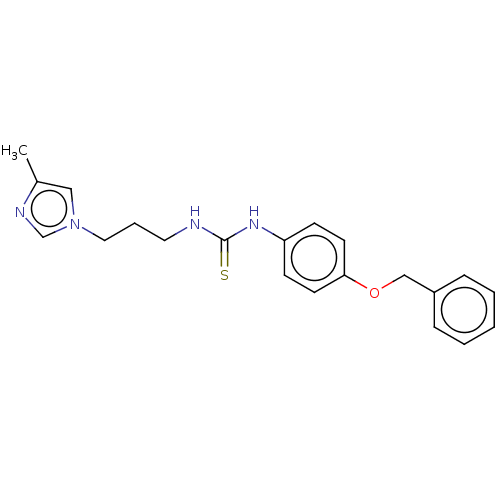
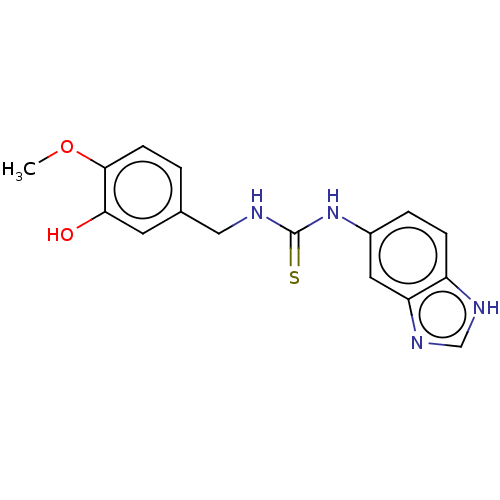
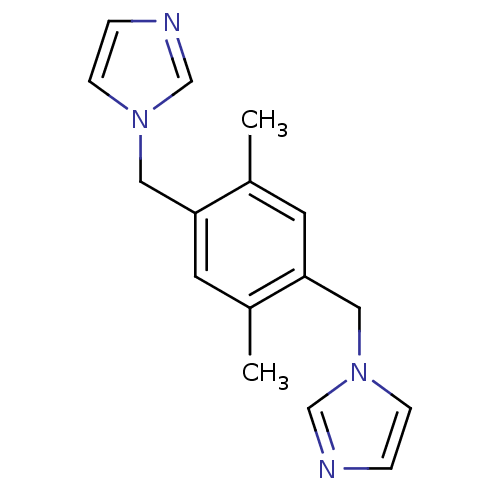
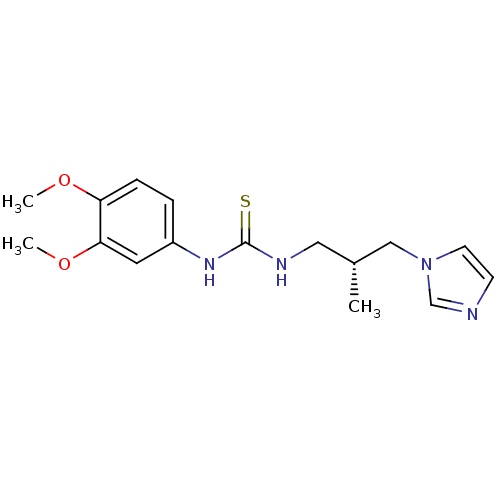
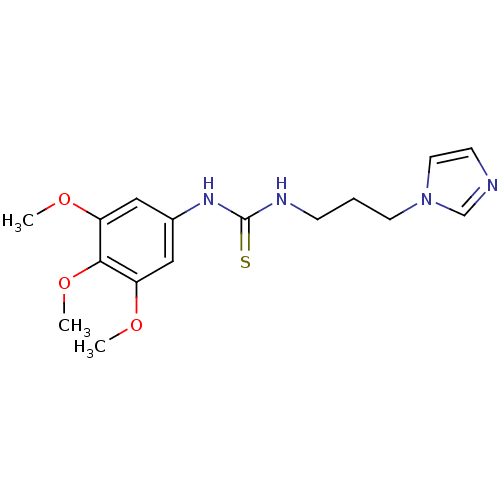
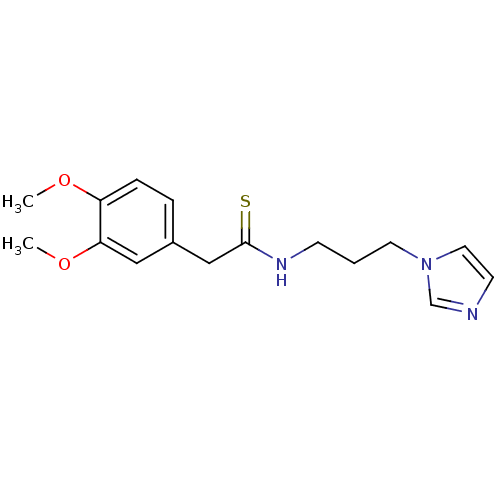
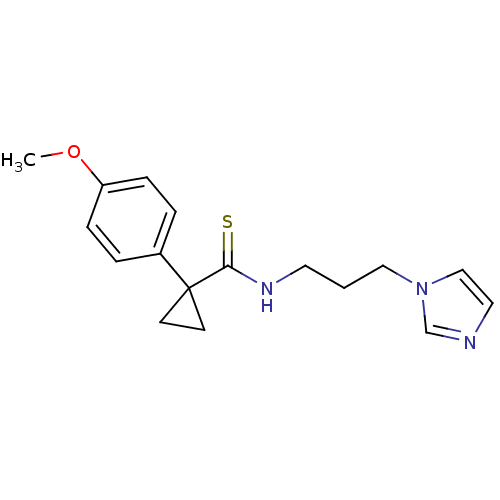
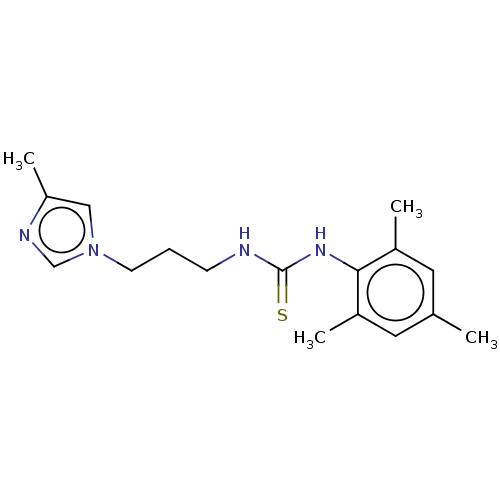
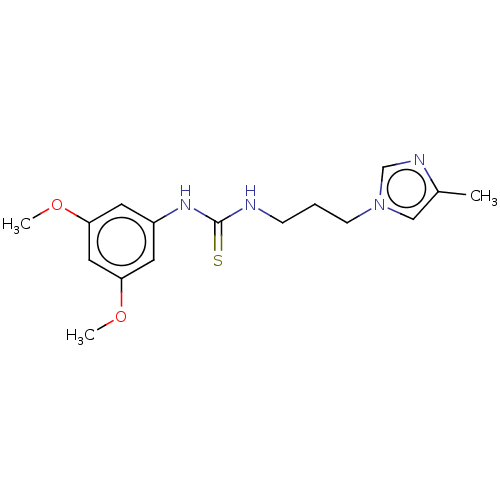
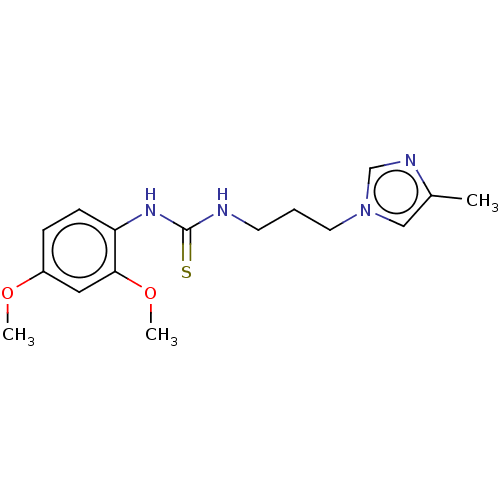
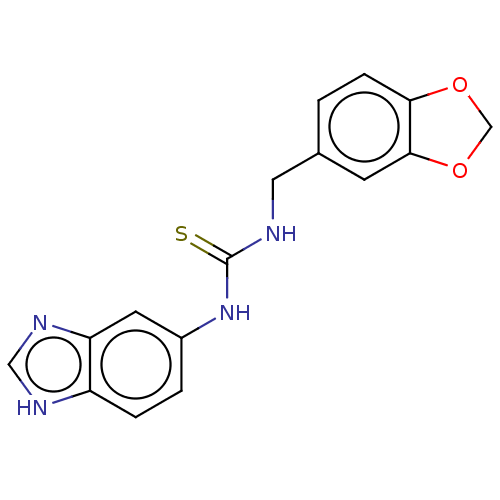
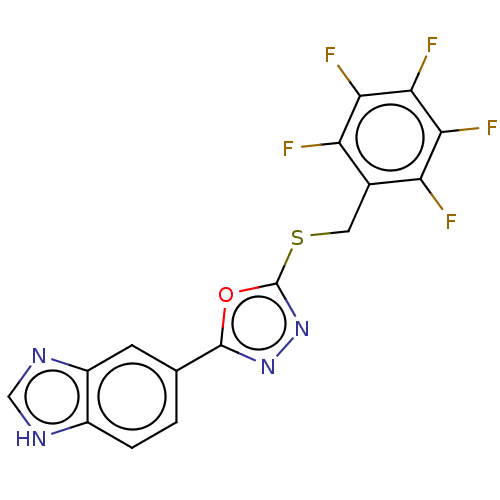
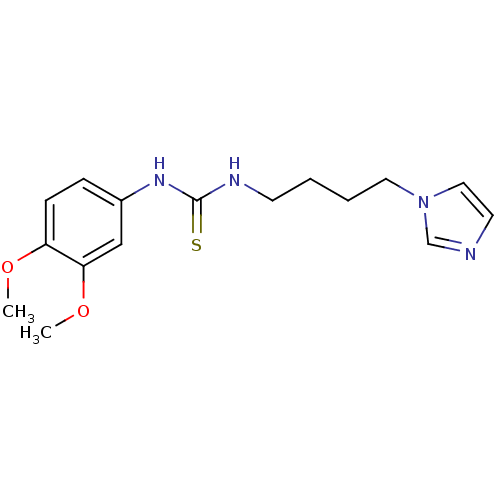
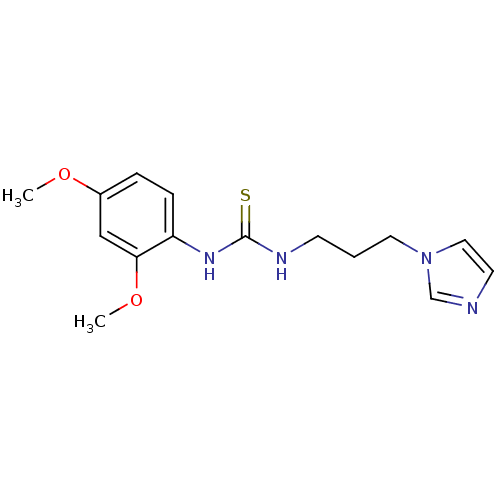
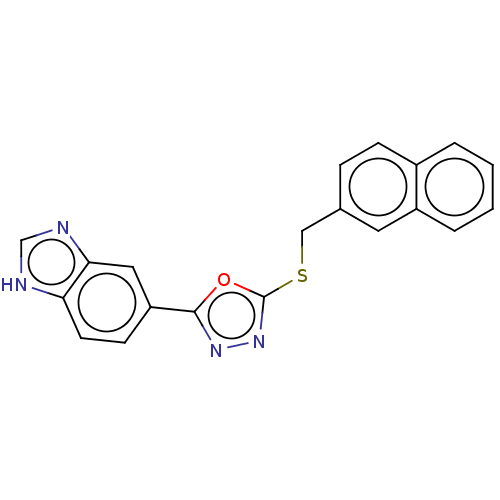
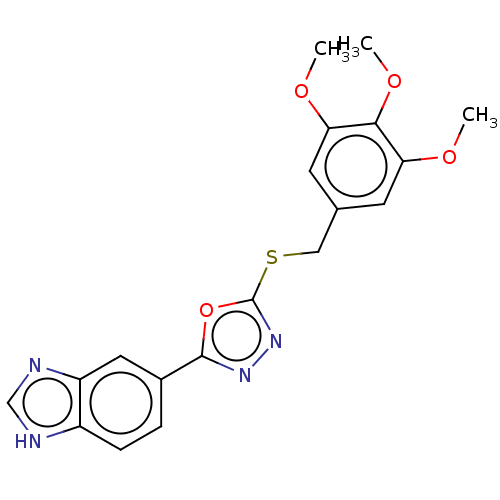
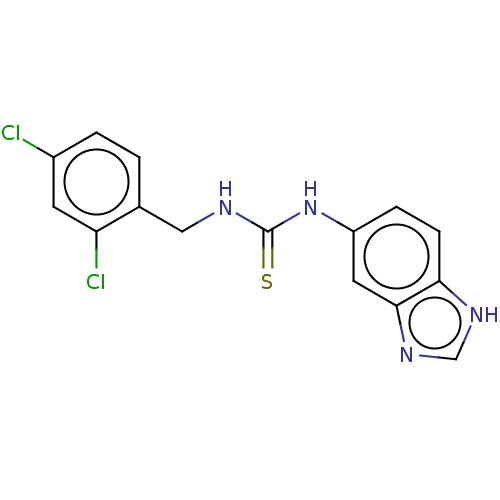
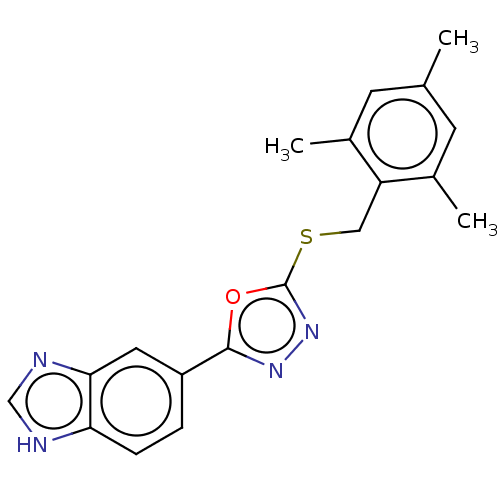
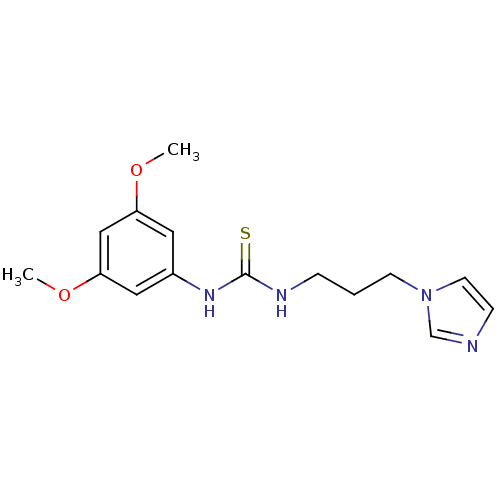
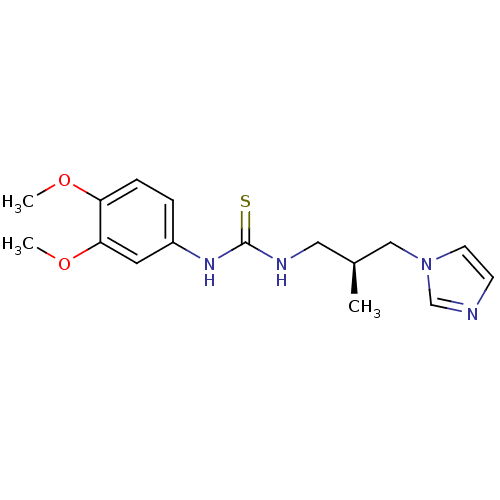
Affinity DataKi: 6nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
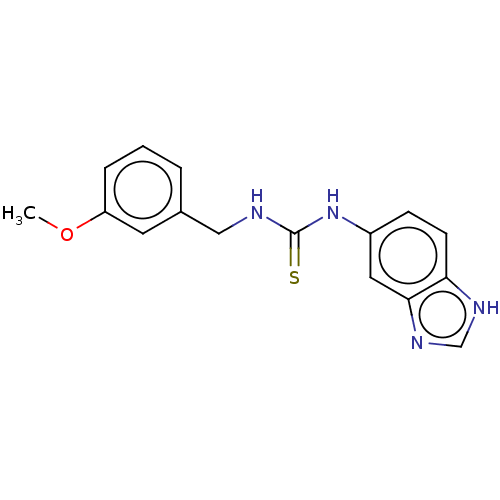
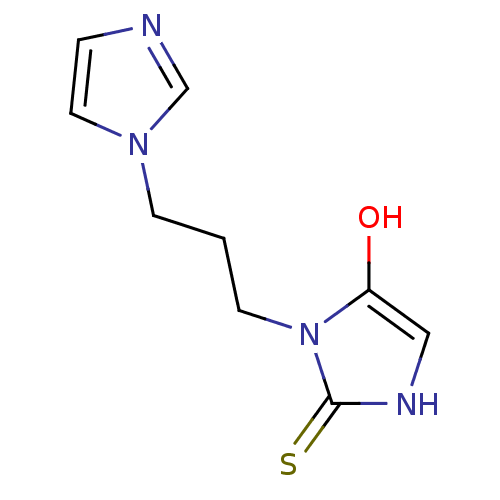
Affinity DataKi: 15nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
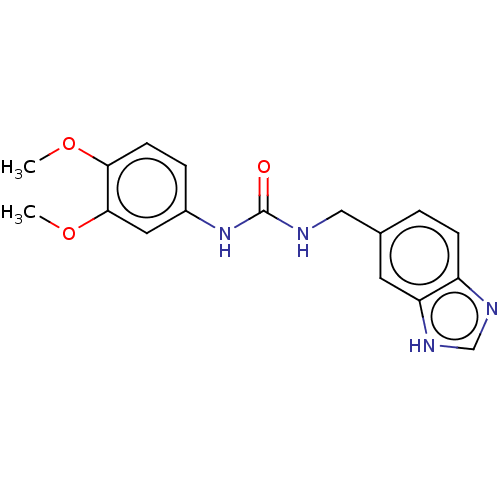
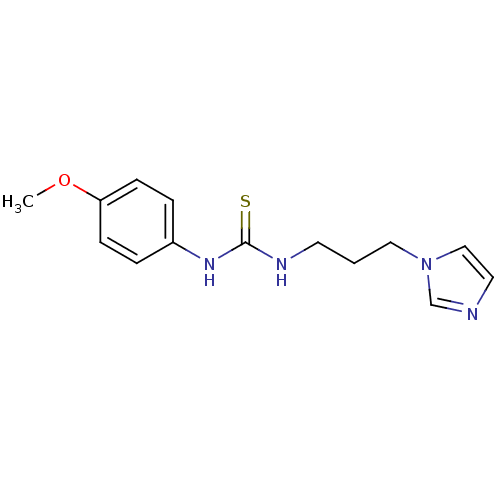
Affinity DataKi: 24nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
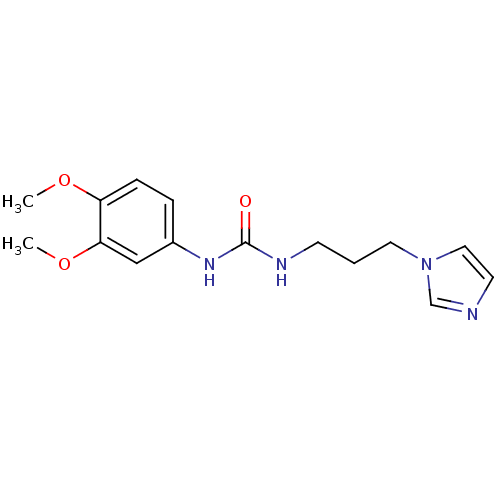
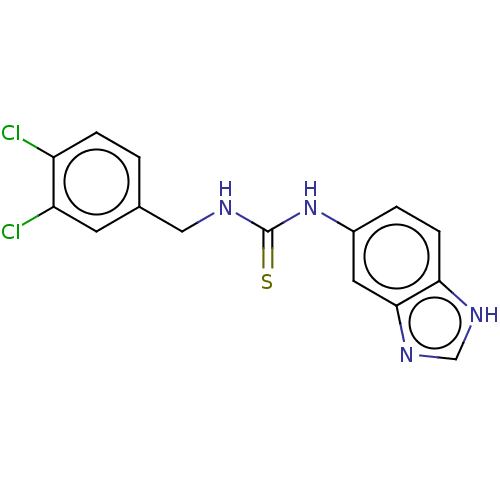
Affinity DataKi: 25nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 27nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 28nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 31.2nM ΔG°: -43.6kJ/mole IC50: 192nMpH: 8.0 T: 2°CAssay Description:For inhibitor testing, the sample composition was the same as described below, except of the putative inhibitory compound added. For a rapid test of ...More data for this Ligand-Target Pair
Affinity DataKi: 32nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 32nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 35nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 60nM ΔG°: -41.9kJ/mole IC50: 343nMpH: 8.0 T: 2°CAssay Description:For inhibitor testing, the sample composition was the same as described below, except of the putative inhibitory compound added. For a rapid test of ...More data for this Ligand-Target Pair
Affinity DataKi: 60nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 60nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 90nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 108nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 110nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 120nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 166nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 166nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 171nM ΔG°: -39.3kJ/mole IC50: 1.67E+3nMpH: 8.0 T: 2°CAssay Description:For inhibitor testing, the sample composition was the same as described below, except of the putative inhibitory compound added. For a rapid test of ...More data for this Ligand-Target Pair
Affinity DataKi: 258nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 264nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 272nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 290nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 295nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 340nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 340nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 390nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 400nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 412nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 426nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 458nM ΔG°: -36.8kJ/mole IC50: 4.25E+3nMpH: 8.0 T: 2°CAssay Description:For inhibitor testing, the sample composition was the same as described below, except of the putative inhibitory compound added. For a rapid test of ...More data for this Ligand-Target Pair
Affinity DataKi: 490nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 496nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 513nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 544nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 550nM ΔG°: -36.3kJ/mole IC50: 2.05E+3nMpH: 8.0 T: 2°CAssay Description:All measurements were performed with a BioAssay Reader HTS-7000Plus for microplates (Perkin Elmer) at 30 C. QC activity was evaluated fluorometricall...More data for this Ligand-Target Pair
Affinity DataKi: 550nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 560nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 690nM ΔG°: -35.8kJ/mole IC50: 8.67E+3nMpH: 8.0 T: 2°CAssay Description:All measurements were performed with a BioAssay Reader HTS-7000Plus for microplates (Perkin Elmer) at 30 C. QC activity was evaluated fluorometricall...More data for this Ligand-Target Pair
Affinity DataKi: 690nM ΔG°: -35.8kJ/mole IC50: 1.53E+4nMpH: 8.0 T: 2°CAssay Description:All measurements were performed with a BioAssay Reader HTS-7000Plus for microplates (Perkin Elmer) at 30 C. QC activity was evaluated fluorometricall...More data for this Ligand-Target Pair
Affinity DataKi: 699nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 700nM ΔG°: -35.7kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 700nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 722nMAssay Description:This novel assay was used to determine the kinetic parameters for most of the QC substrates. QC activity was analyzed spectrophotometrically using a ...More data for this Ligand-Target Pair
Affinity DataKi: 740nM ΔG°: -35.6kJ/mole IC50: 7.30E+3nMpH: 8.0 T: 2°CAssay Description:All measurements were performed with a BioAssay Reader HTS-7000Plus for microplates (Perkin Elmer) at 30 C. QC activity was evaluated fluorometricall...More data for this Ligand-Target Pair
Affinity DataKi: 750nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 760nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair

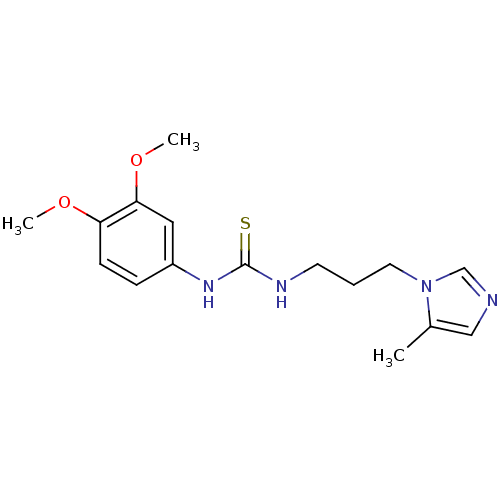
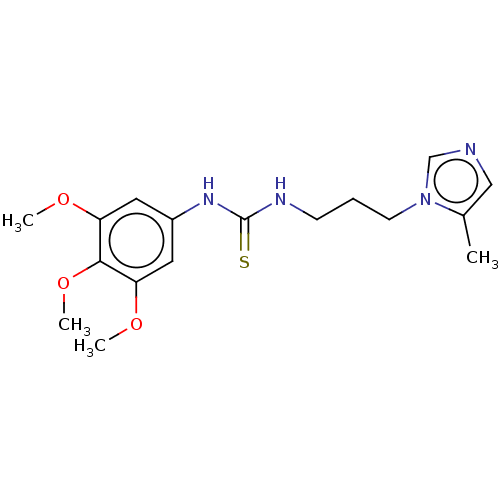
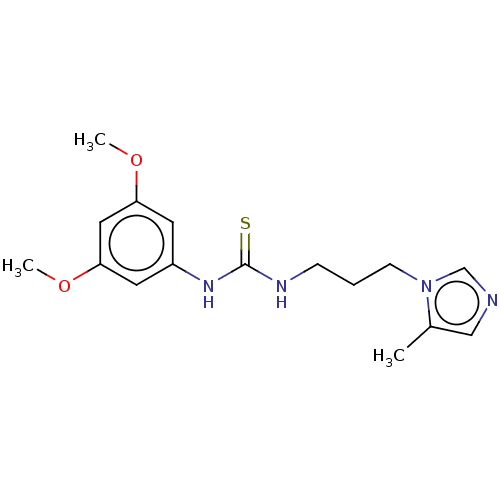
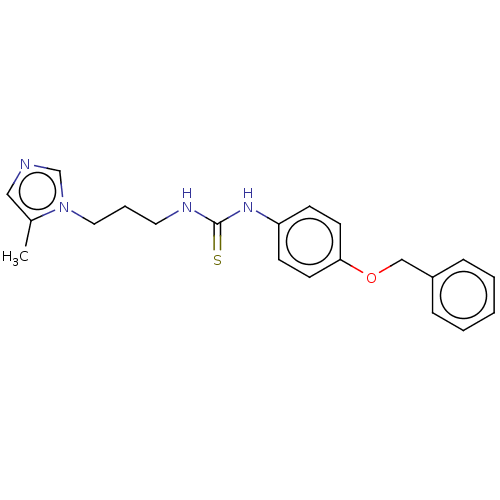
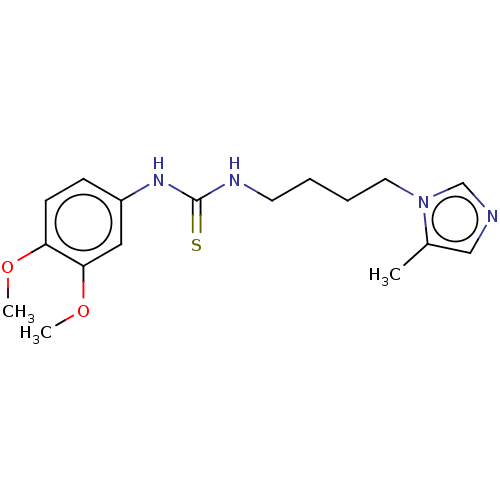
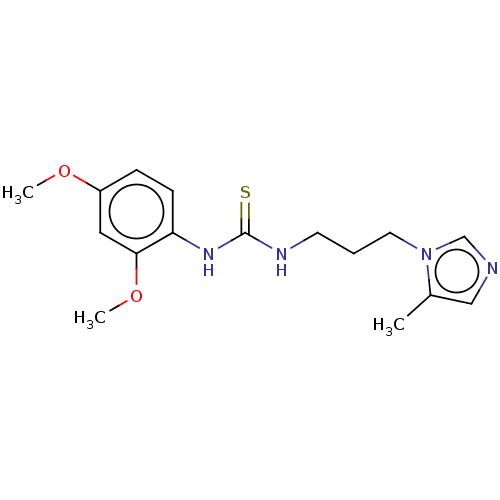
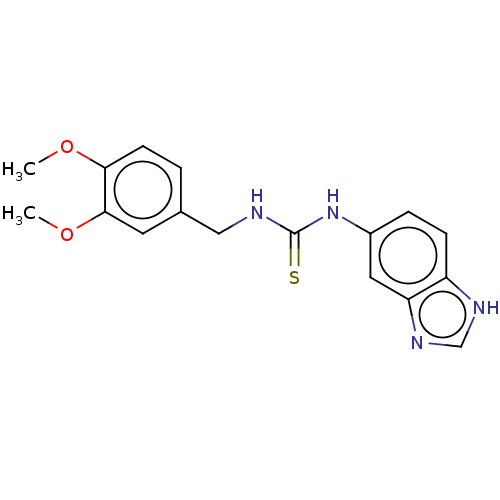
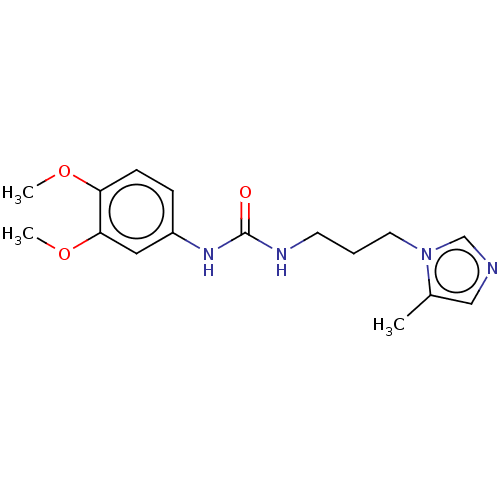
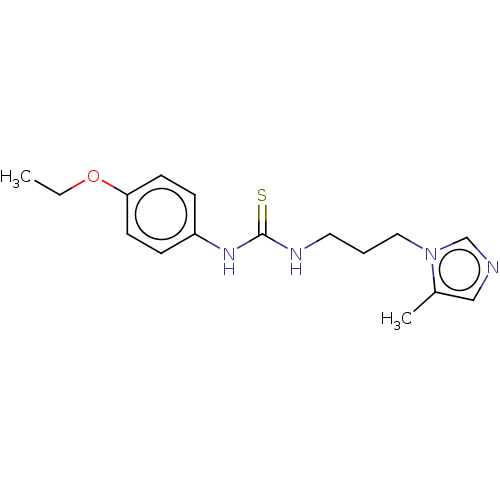
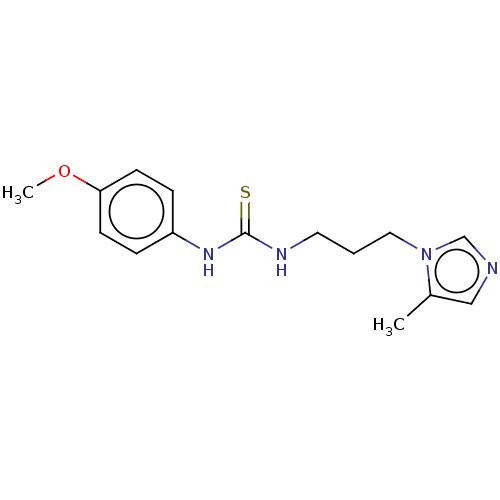
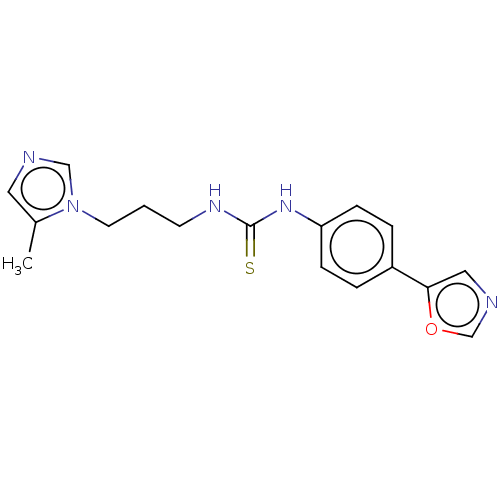
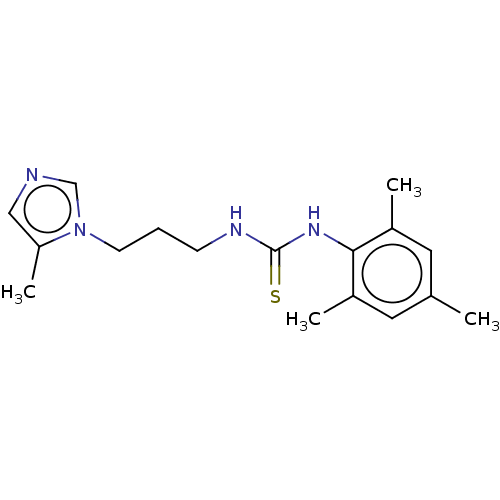
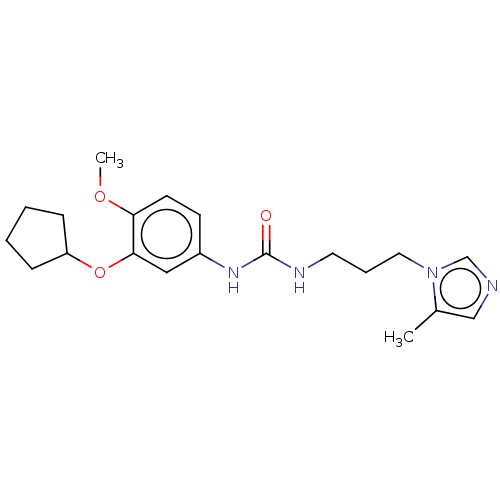
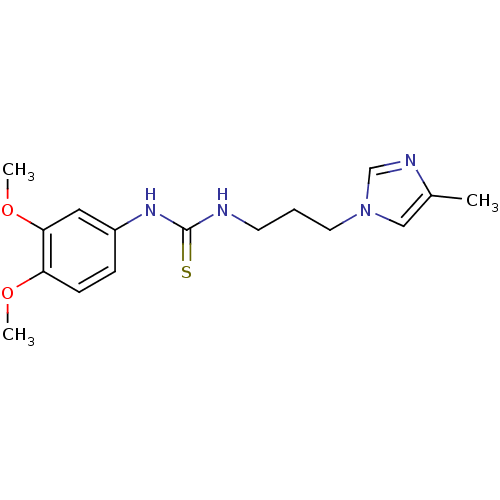
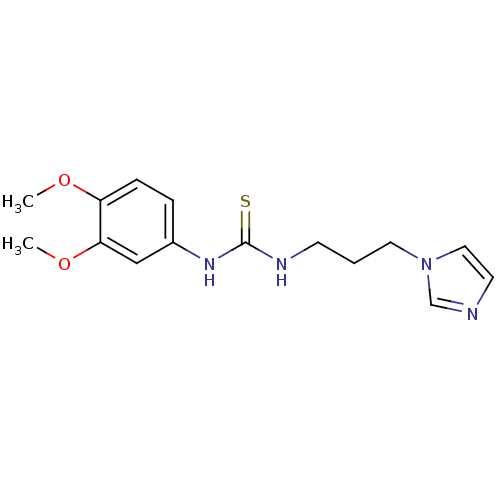
 3D Structure (crystal)
3D Structure (crystal)