Affinity DataKi: 0.0300nM ΔG°: -55.8kJ/moleT: 2°CAssay Description:For the membrane binding, NIH-3T3 cells were grown to 70% confluence in 15-cm2 dishes and transfected with 10 ug of receptor plasmid DNA using Polyfe...More data for this Ligand-Target Pair
Affinity DataKi: 0.0460nM ΔG°: -54.8kJ/moleT: 2°CAssay Description:For the membrane binding, NIH-3T3 cells were grown to 70% confluence in 15-cm2 dishes and transfected with 10 ug of receptor plasmid DNA using Polyfe...More data for this Ligand-Target Pair
Affinity DataKi: 0.0830nM ΔG°: -53.5kJ/moleT: 2°CAssay Description:For the membrane binding, NIH-3T3 cells were grown to 70% confluence in 15-cm2 dishes and transfected with 10 ug of receptor plasmid DNA using Polyfe...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nM ΔG°: -54.8kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nM ΔG°: -54.8kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 0.320nM ΔG°: -53.6kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 0.440nM ΔG°: -49.6kJ/moleT: 2°CAssay Description:For the membrane binding, NIH-3T3 cells were grown to 70% confluence in 15-cm2 dishes and transfected with 10 ug of receptor plasmid DNA using Polyfe...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nM ΔG°: -49.3kJ/moleT: 2°CAssay Description:For the membrane binding, NIH-3T3 cells were grown to 70% confluence in 15-cm2 dishes and transfected with 10 ug of receptor plasmid DNA using Polyfe...More data for this Ligand-Target Pair
Affinity DataKi: 0.550nM ΔG°: -49.1kJ/moleT: 2°CAssay Description:For the membrane binding, NIH-3T3 cells were grown to 70% confluence in 15-cm2 dishes and transfected with 10 ug of receptor plasmid DNA using Polyfe...More data for this Ligand-Target Pair
Affinity DataKi: 0.790nM ΔG°: -51.4kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 1.20nM ΔG°: -50.4kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 1.30nM ΔG°: -47.1kJ/moleT: 2°CAssay Description:For the membrane binding, NIH-3T3 cells were grown to 70% confluence in 15-cm2 dishes and transfected with 10 ug of receptor plasmid DNA using Polyfe...More data for this Ligand-Target Pair
Affinity DataKi: 1.30nM ΔG°: -47.1kJ/moleT: 2°CAssay Description:For the membrane binding, NIH-3T3 cells were grown to 70% confluence in 15-cm2 dishes and transfected with 10 ug of receptor plasmid DNA using Polyfe...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nM ΔG°: -46.7kJ/moleT: 2°CAssay Description:For the membrane binding, NIH-3T3 cells were grown to 70% confluence in 15-cm2 dishes and transfected with 10 ug of receptor plasmid DNA using Polyfe...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nM ΔG°: -49.7kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -49.2kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -48.2kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -48.2kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 3.80nM ΔG°: -44.7kJ/moleT: 2°CAssay Description:For the membrane binding, NIH-3T3 cells were grown to 70% confluence in 15-cm2 dishes and transfected with 10 ug of receptor plasmid DNA using Polyfe...More data for this Ligand-Target Pair
Affinity DataKi: 25nM ΔG°: -43.0kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 40nM ΔG°: -41.8kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 398nM ΔG°: -36.2kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 1.00E+3nM ΔG°: -33.9kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 1.26E+3nM ΔG°: -33.3kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nM ΔG°: -32.2kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataKi: 5.01E+3nM ΔG°: -29.9kJ/molepH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0690nMAssay Description:R-SAT assays were performed as described previously (Weiner et al., 2001), with the following modifications. In brief, NIH-3T3 cells were grown to 80...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:R-SAT assays were performed as described previously (Weiner et al., 2001), with the following modifications. In brief, NIH-3T3 cells were grown to 80...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:R-SAT assays were performed as described previously (Weiner et al., 2001), with the following modifications. In brief, NIH-3T3 cells were grown to 80...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90nMAssay Description:R-SAT assays were performed as described previously (Weiner et al., 2001), with the following modifications. In brief, NIH-3T3 cells were grown to 80...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:R-SAT assays were performed as described previously (Weiner et al., 2001), with the following modifications. In brief, NIH-3T3 cells were grown to 80...More data for this Ligand-Target Pair
Affinity DataIC50: 4.5nMAssay Description:R-SAT assays were performed as described previously (Weiner et al., 2001), with the following modifications. In brief, NIH-3T3 cells were grown to 80...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:R-SAT assays were performed as described previously (Weiner et al., 2001), with the following modifications. In brief, NIH-3T3 cells were grown to 80...More data for this Ligand-Target Pair
Affinity DataIC50: 91nMAssay Description:R-SAT assays were performed as described previously (Weiner et al., 2001), with the following modifications. In brief, NIH-3T3 cells were grown to 80...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:R-SAT assays were performed as described previously (Weiner et al., 2001), with the following modifications. In brief, NIH-3T3 cells were grown to 80...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:R-SAT assays were performed as described previously (Weiner et al., 2001), with the following modifications. In brief, NIH-3T3 cells were grown to 80...More data for this Ligand-Target Pair
Affinity DataIC50: 540nMAssay Description:R-SAT assays were performed as described previously (Weiner et al., 2001), with the following modifications. In brief, NIH-3T3 cells were grown to 80...More data for this Ligand-Target Pair
Affinity DatapH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DatapH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DatapH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DatapH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DatapH: 7.2 T: 2°CAssay Description:IC50 values for test compounds were determined from nonlinear regression analysis of data collected from ligand displacement experiments. The inhibit...More data for this Ligand-Target Pair
Affinity DataEC50: 0.251nMAssay Description:Agonist activity at androgen receptor in human MDA-KB2 cells transfected with MMTV linked luciferase assessed as transcriptional activation by lucife...More data for this Ligand-Target Pair
Affinity DataEC50: 3.16nMAssay Description:Agonist activity at androgen receptor in human MDA-KB2 cells transfected with MMTV linked luciferase assessed as transcriptional activation by lucife...More data for this Ligand-Target Pair
Affinity DataEC50: 1nMAssay Description:Agonist activity at androgen receptor in mouse NIH3T3 cells transiently transfected with beta-galactosidase reporter gene assessed as cellular transf...More data for this Ligand-Target Pair
Affinity DataEC50: 3.98nMAssay Description:Agonist activity at androgen receptor in mouse NIH3T3 cells transiently transfected with beta-galactosidase reporter gene assessed as cellular transf...More data for this Ligand-Target Pair
Affinity DataEC50: 12.6nMAssay Description:Agonist activity at androgen receptor in mouse NIH3T3 cells transiently transfected with beta-galactosidase reporter gene assessed as cellular transf...More data for this Ligand-Target Pair
Affinity DataEC50: 19.9nMAssay Description:Agonist activity at androgen receptor in mouse NIH3T3 cells transiently transfected with beta-galactosidase reporter gene assessed as cellular transf...More data for this Ligand-Target Pair
Affinity DataEC50: 100nMAssay Description:Agonist activity at androgen receptor in mouse NIH3T3 cells transiently transfected with beta-galactosidase reporter gene assessed as cellular transf...More data for this Ligand-Target Pair
Affinity DataEC50: 2nMAssay Description:Agonist activity at androgen receptor in mouse NIH3T3 cells transiently transfected with beta-galactosidase reporter gene assessed as cellular transf...More data for this Ligand-Target Pair

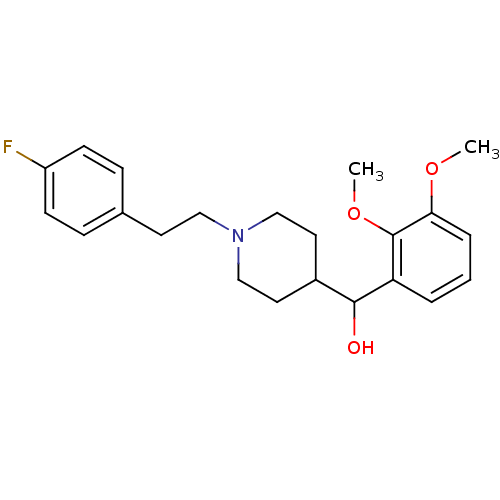
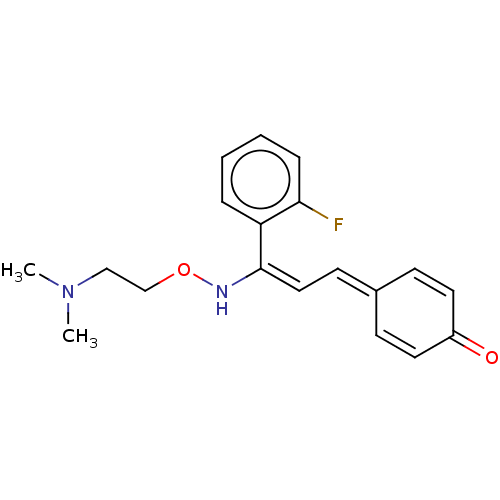
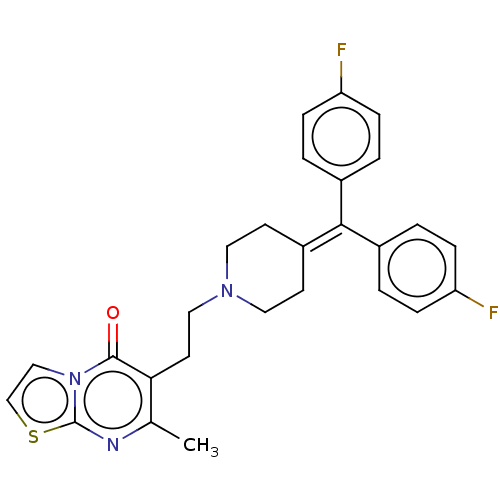
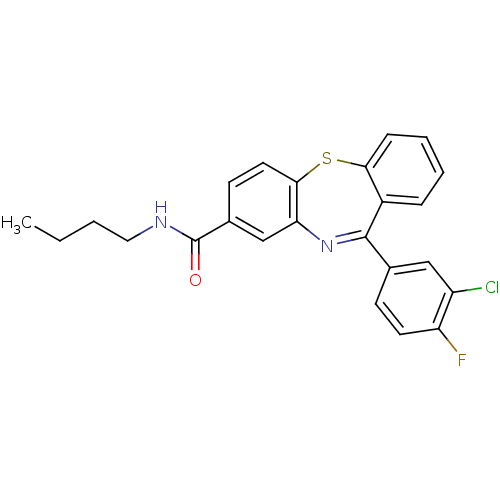
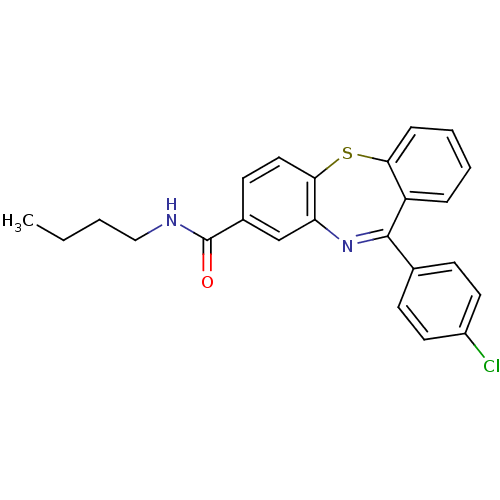
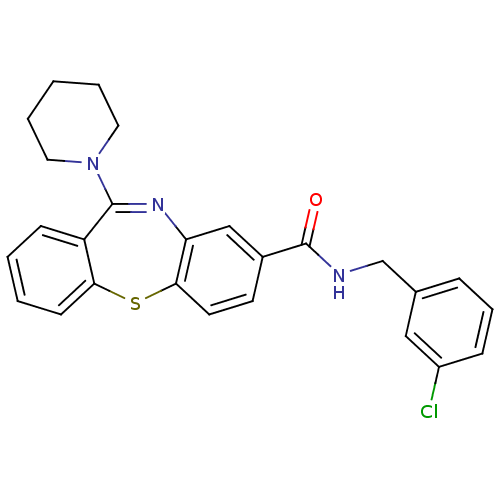
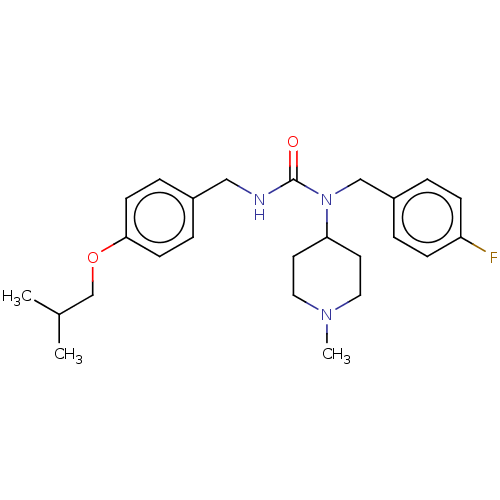
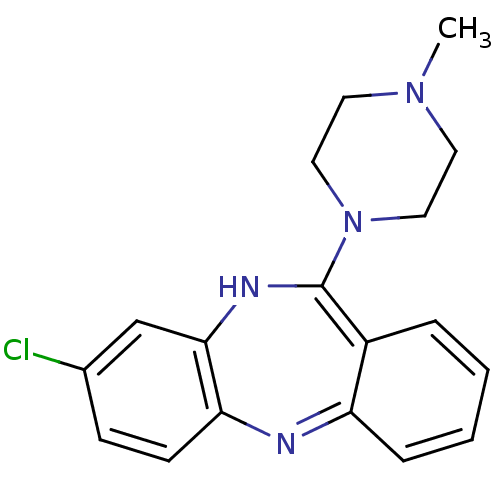
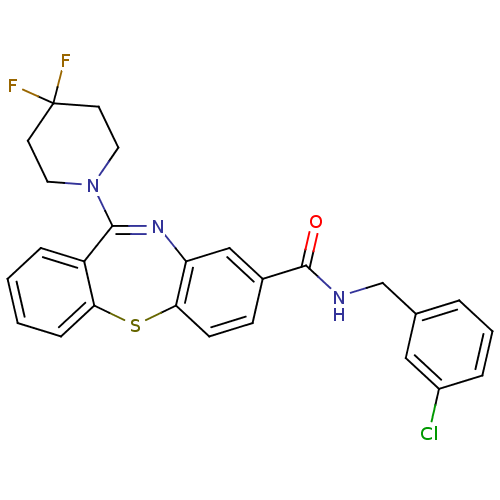
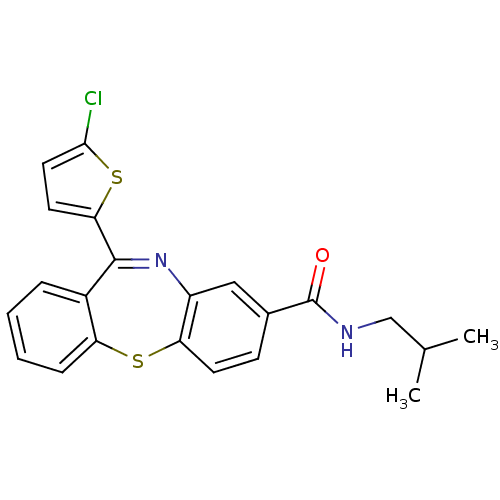
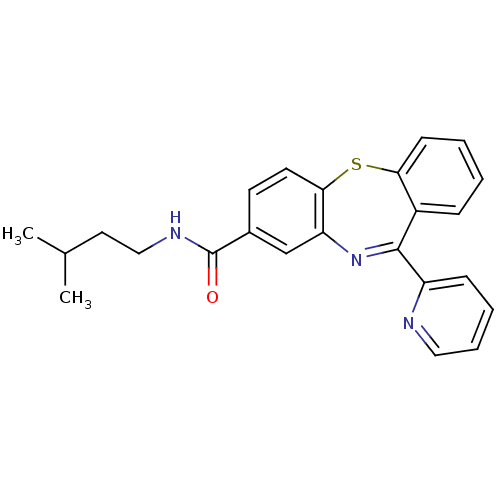
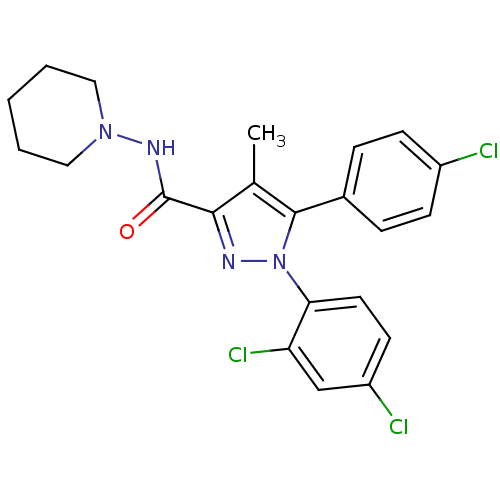
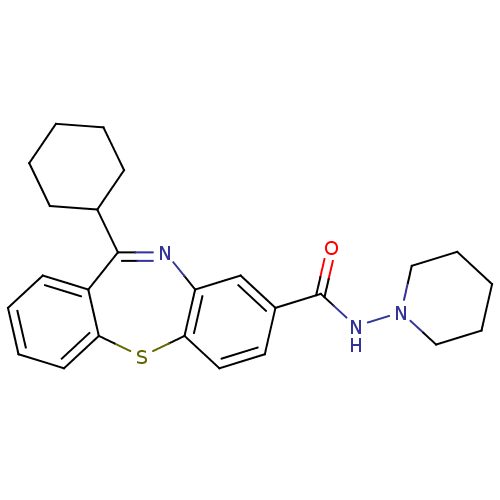
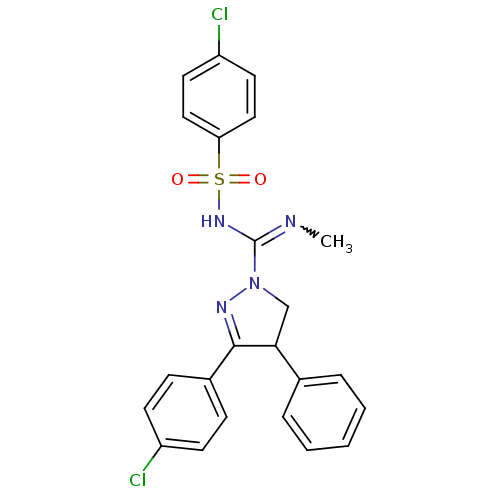
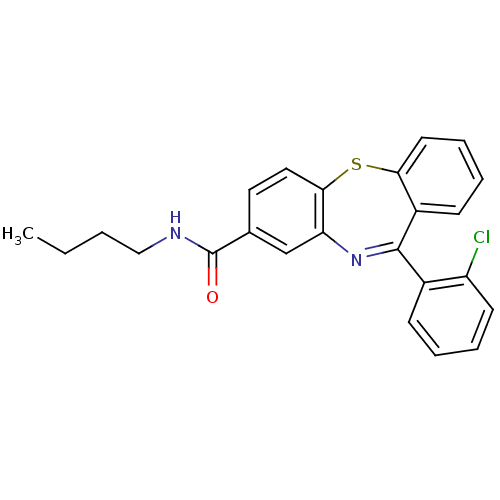
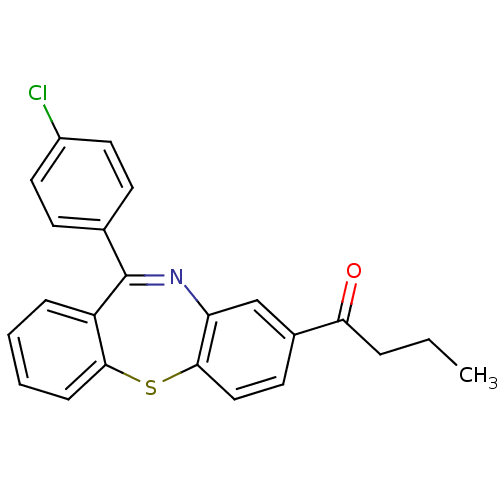
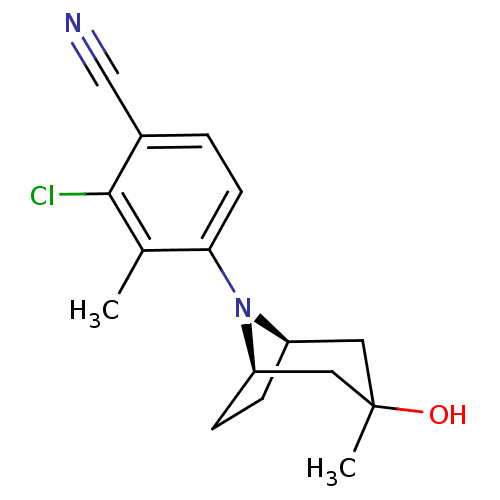
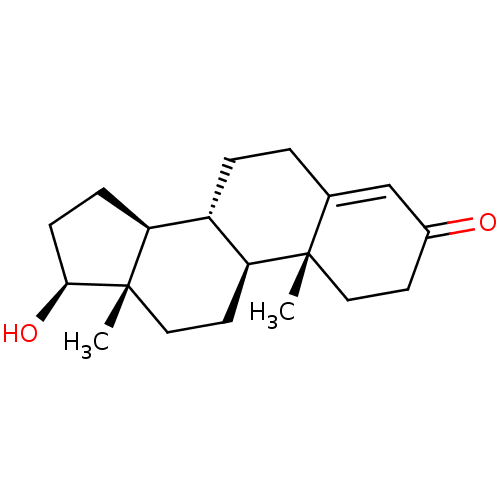
 3D Structure (crystal)
3D Structure (crystal)