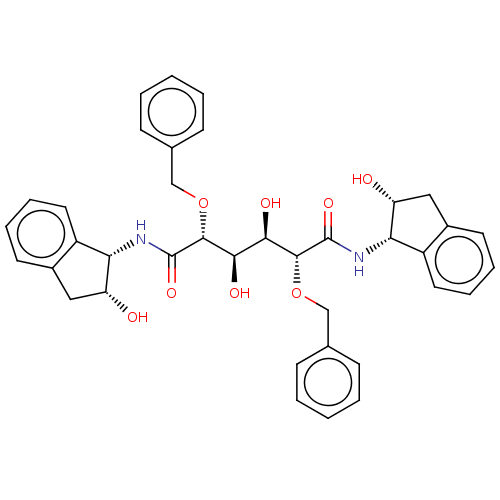
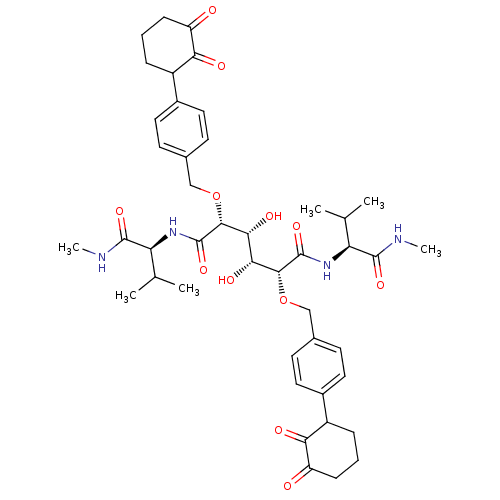
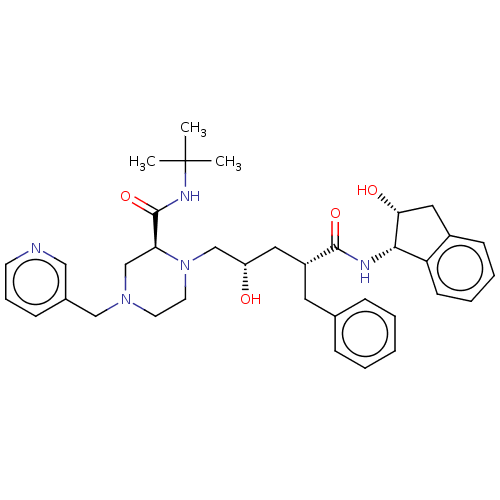
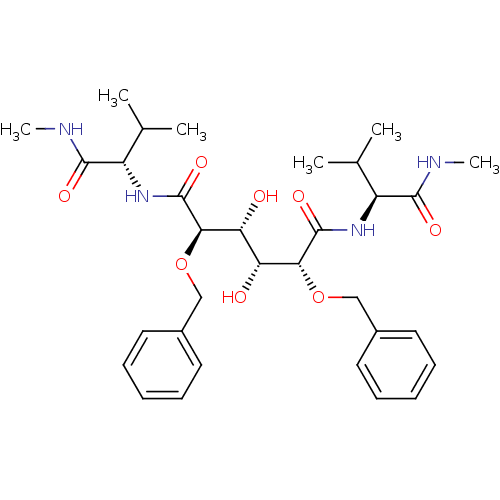
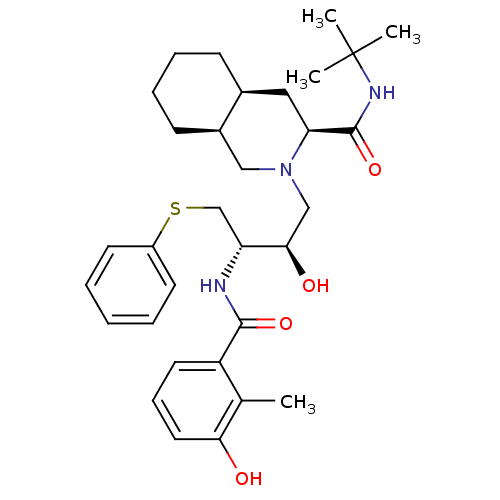
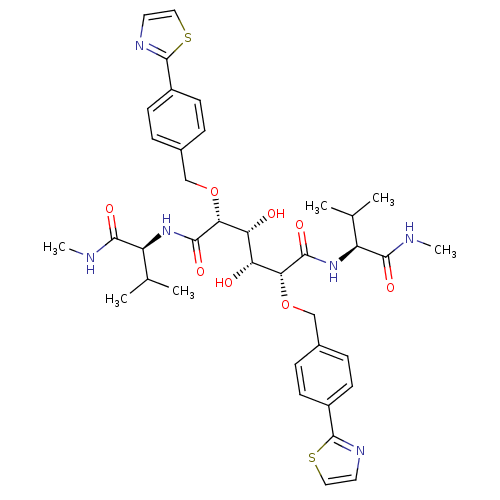
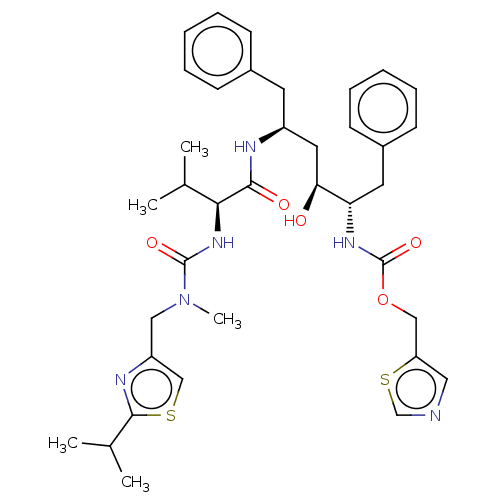
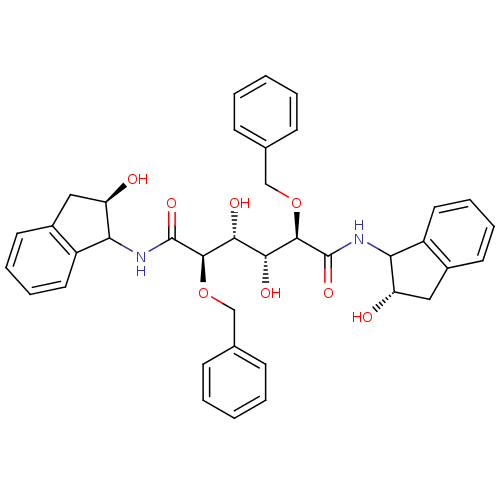
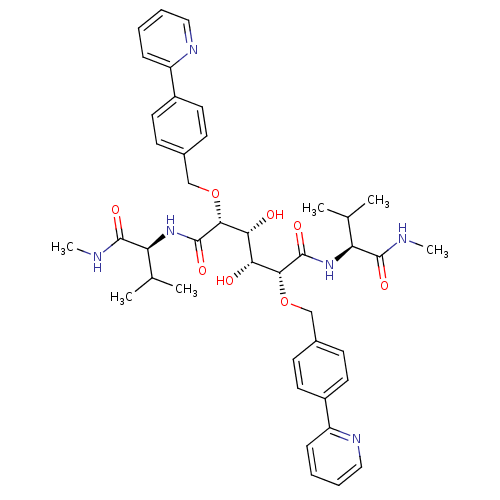
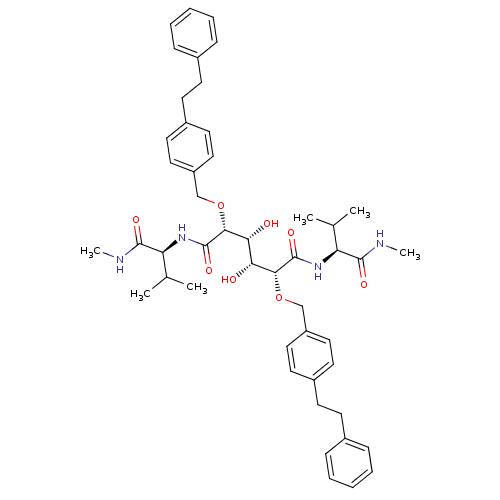
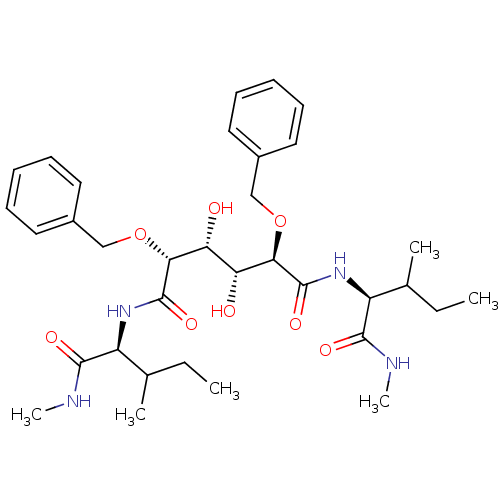
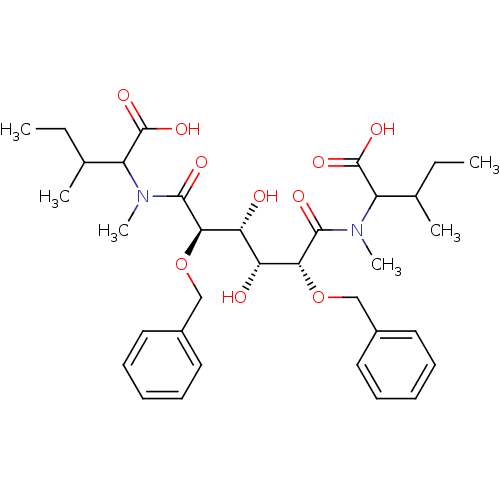
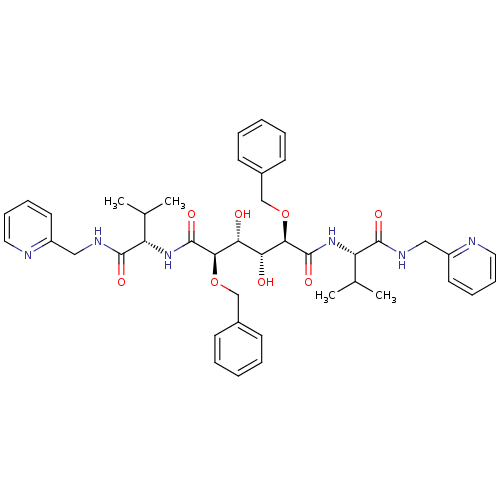
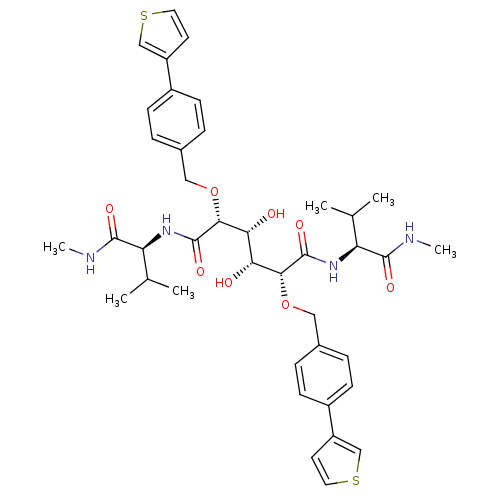
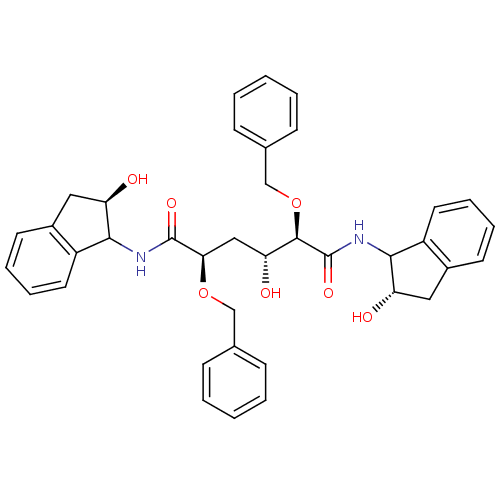
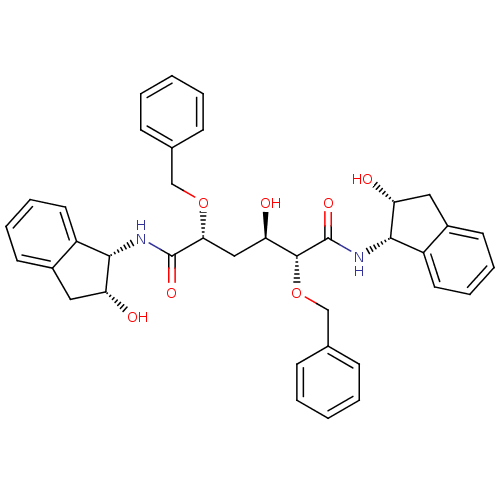
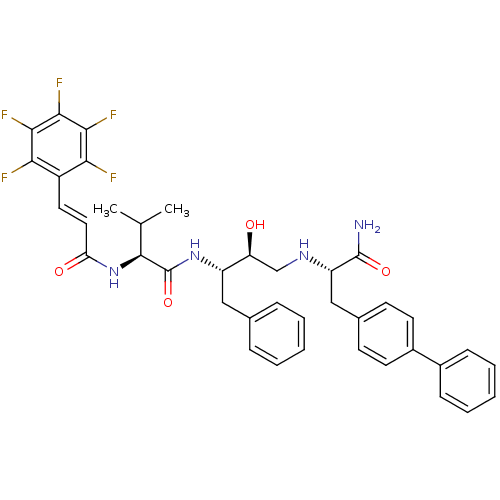
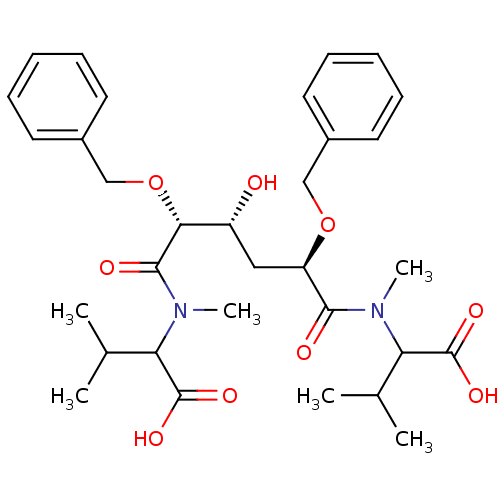
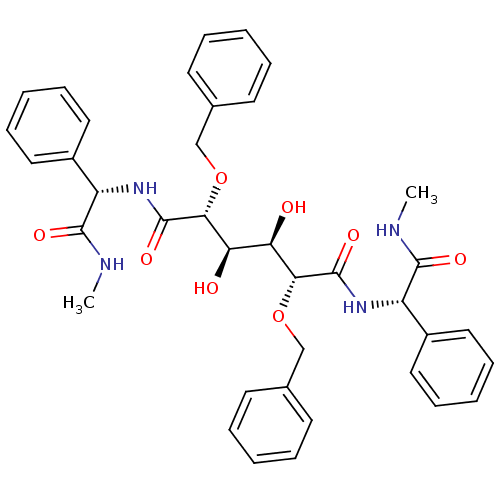
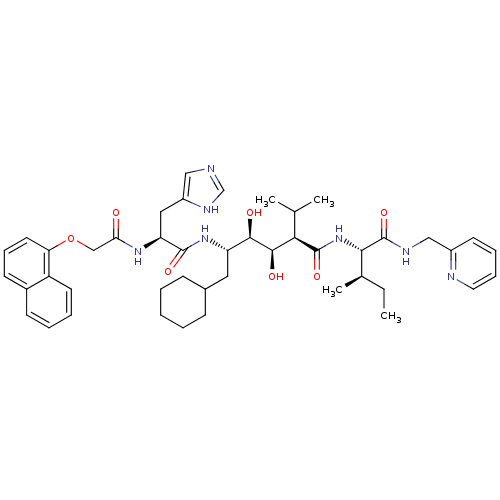
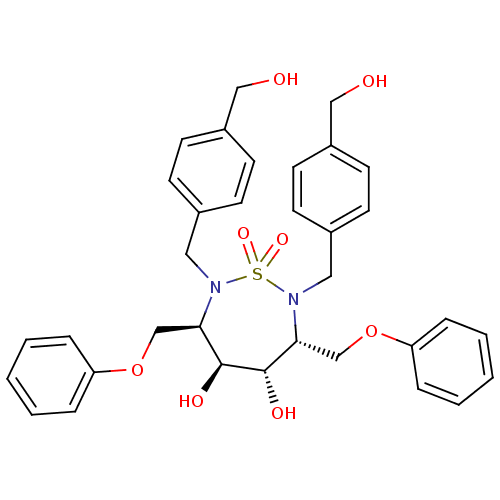
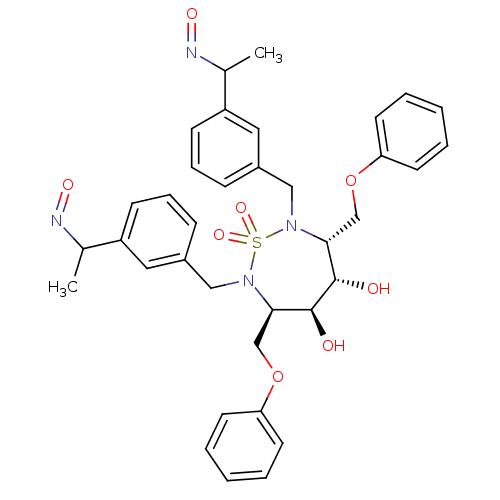
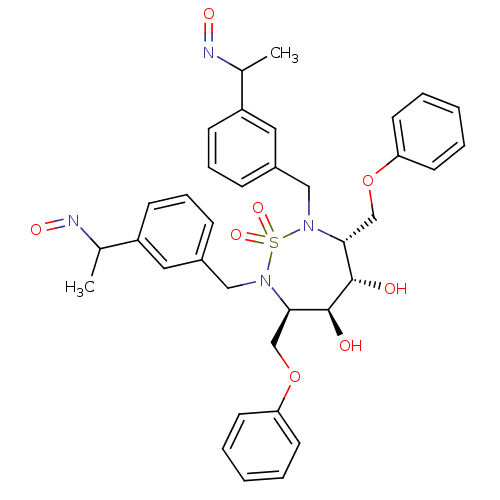
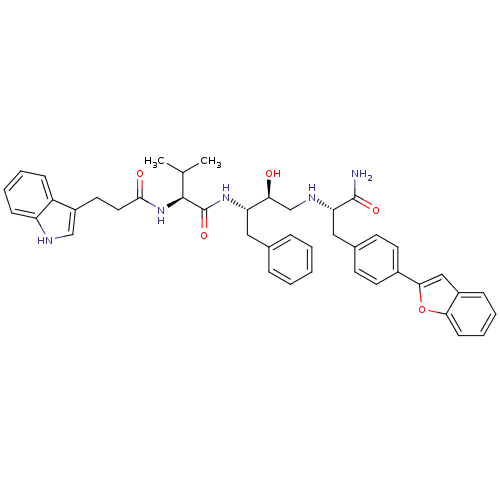
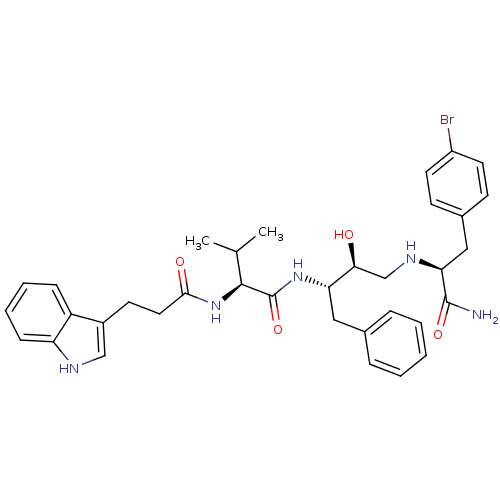
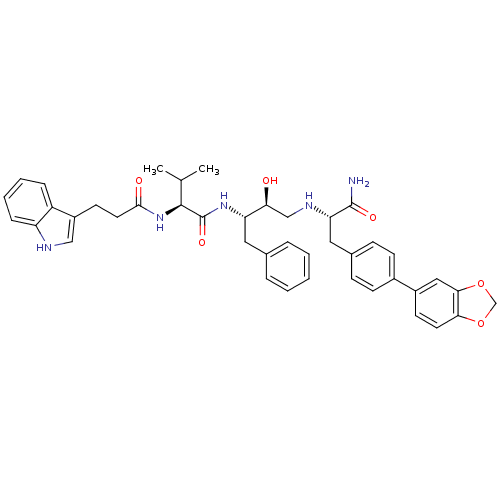
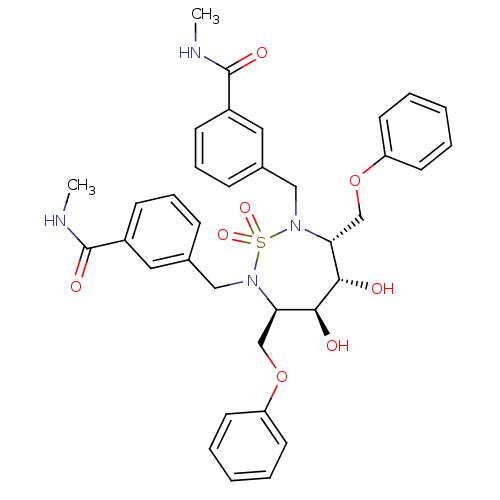
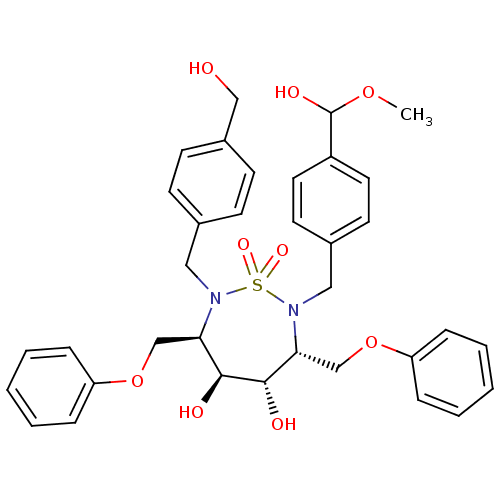
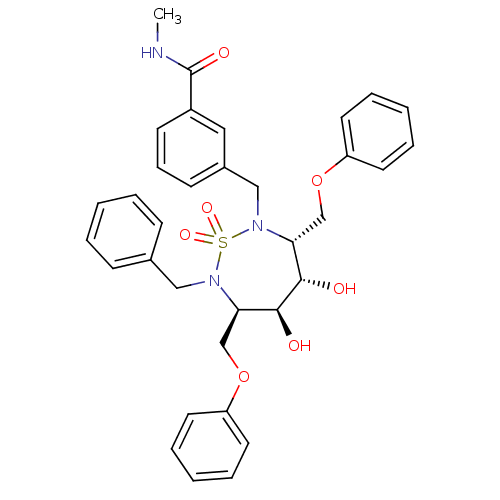
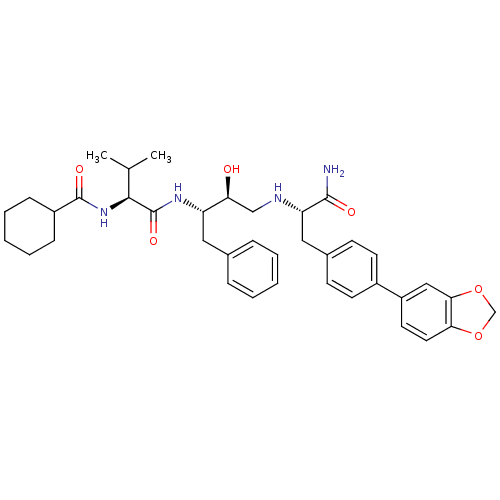
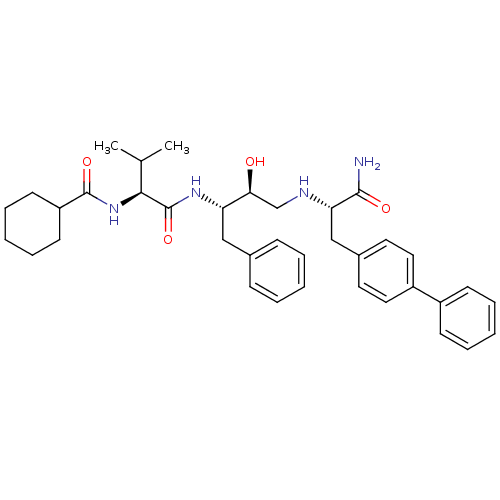
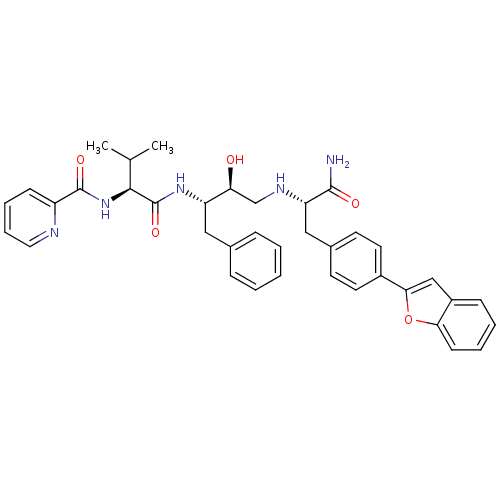
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.0540nM Kd: 4.42nM Koff: 5.97E+6s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
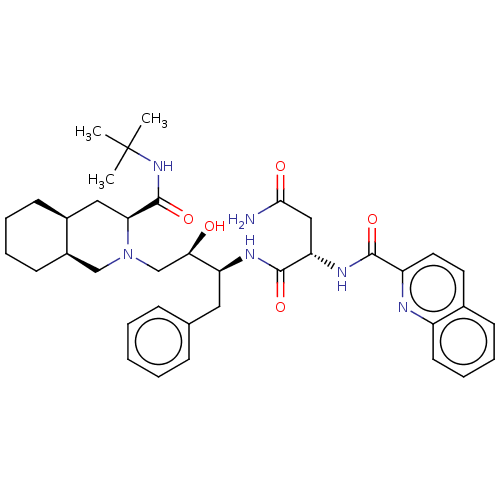
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.0900nM Kd: 1.95nM Koff: 2.04E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
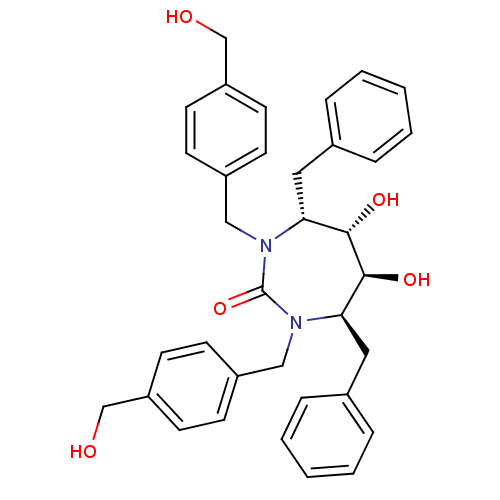
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.100nM Kd: 70nM Koff: 1.01E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -58.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -58.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nM ΔG°: -56.3kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
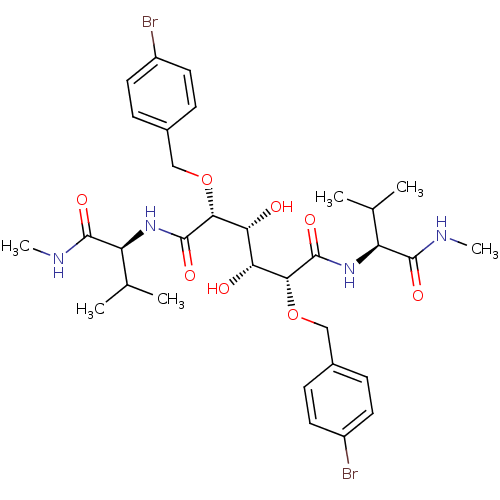
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.230nM Kd: 7nM Koff: 7.06E+9s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.230nM Kd: 1.13nM Koff: 4.43E+6s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.230nM Kd: 0.315nM Koff: 8.17E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.270nM Kd: 3.83nM Koff: 2.52E+10s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.300nM Kd: 21nM Koff: 8.89E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.300nM Kd: 18nM Koff: 2.93E+4s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.310nM Kd: 1.07nM Koff: 1.53E+6s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.400nM Kd: 11nM Koff: 3.55E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.400nM ΔG°: -54.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.540nM Kd: 1.64nM Koff: 6.63E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.550nM Kd: 0.643nM Koff: 4.77E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.590nM Kd: 0.608nM Koff: 3.92E+6s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.600nM Kd: 14nM Koff: 6.39E+6s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.610nM Kd: 1.18nM Koff: 3.23E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.800nM Kd: 34nM Koff: 8.11E+4s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.900nM ΔG°: -52.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 0.910nM Kd: 48nM Koff: 1.85E+6s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 0.920nM ΔG°: -52.4kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 1.20nM ΔG°: -51.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 1.20nM Kd: 3nM Koff: 3.48E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 1.40nM Kd: 36nM Koff: 6.66E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 1.40nM Kd: 18nM Koff: 1.81E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 1.40nM ΔG°: -51.4kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 2.10nM Kd: 102nM Koff: 3.04E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 2.30nM Kd: 27.4nM Koff: 2.05E+8s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 2.80nM Kd: 0.484nM Koff: 6.76E+6s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 3.10nM Kd: 39.8nM Koff: 6.87E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 3.10nMAssay Description:Inhibitory activity against HIV-1 Protease expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 3.10nM ΔG°: -49.4kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 3.40nM ΔG°: -49.1kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 3.40nMAssay Description:Inhibitory activity against HIV-1 Protease expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 4.10nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 4.40nM ΔG°: -48.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 4.70nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 4.80nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 5.60nM Kd: 169nM Koff: 5.12E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 6.20nM Kd: 675nM Koff: 4.99E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 7.30nM ΔG°: -47.2kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 10.1nM Kd: 577nM Koff: 1.88E+5s-1Assay Description:Association rate constant for the interaction between inhibitor and HIV-1 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 11nM ΔG°: -46.2kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair

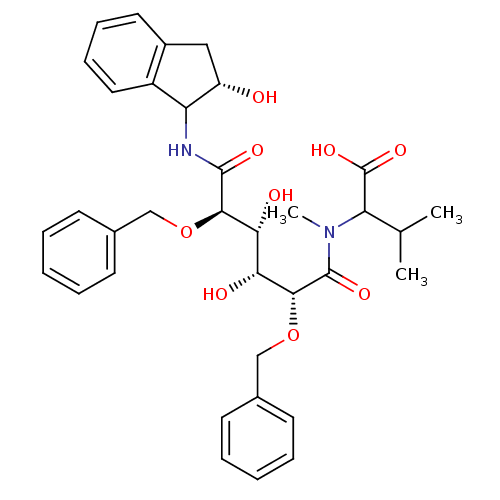
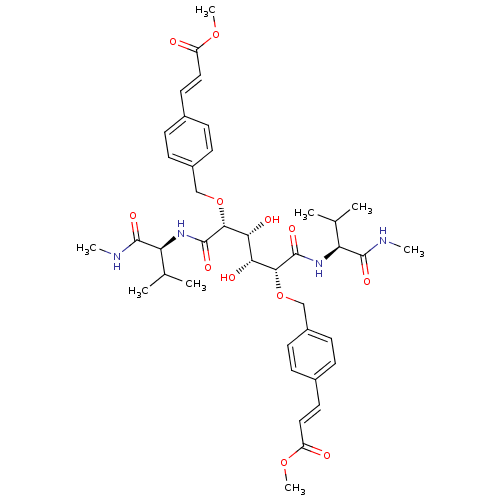
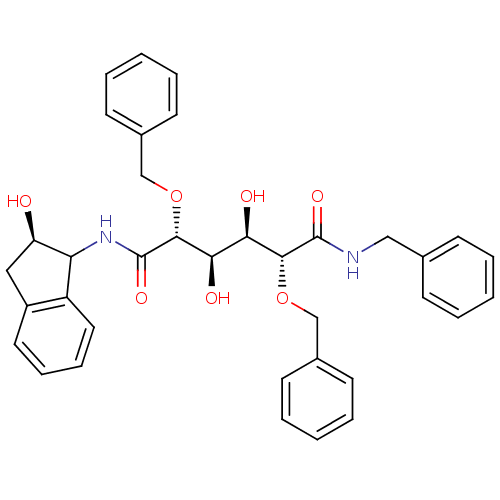
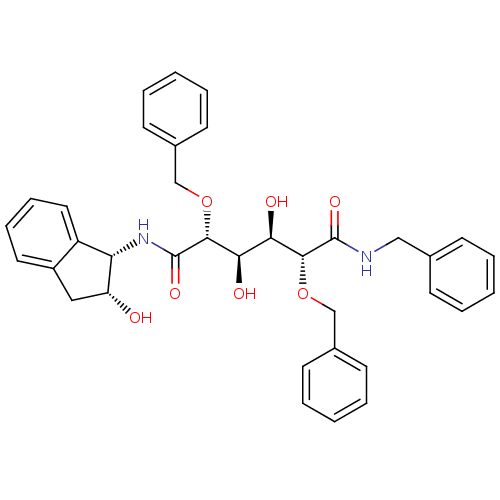
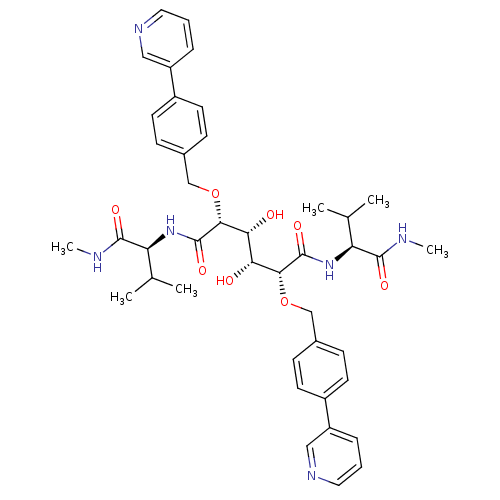
 3D Structure (crystal)
3D Structure (crystal)