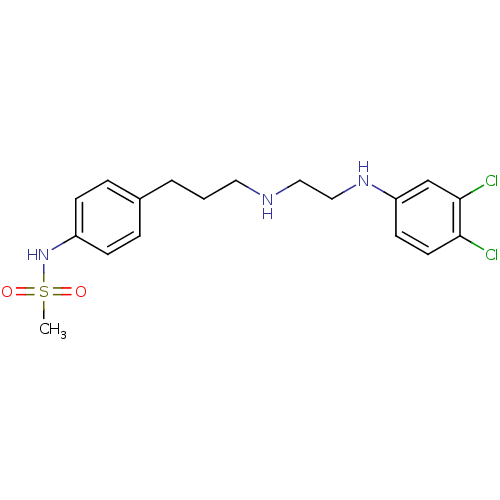
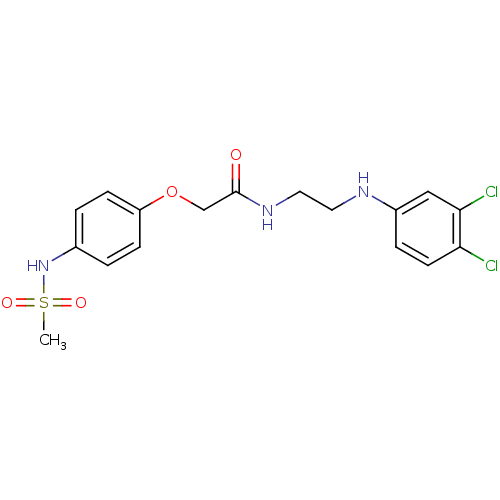
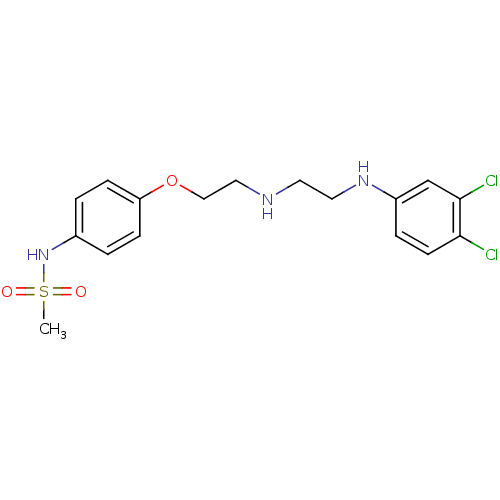
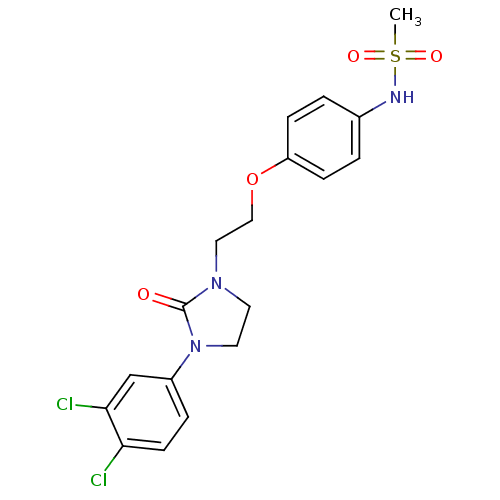
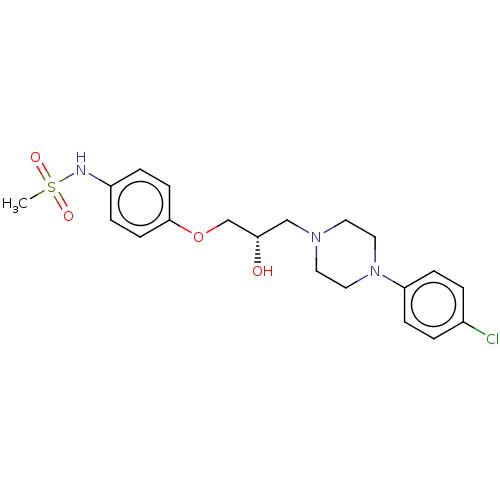
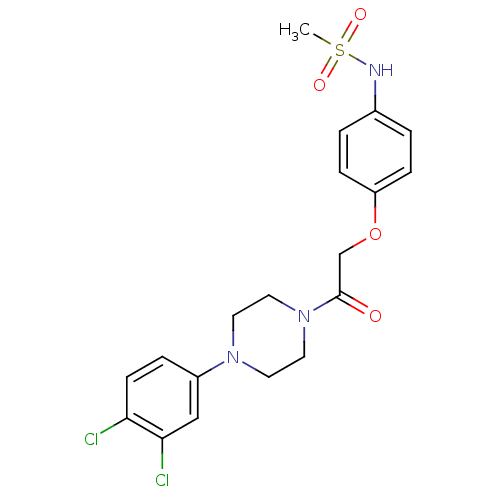
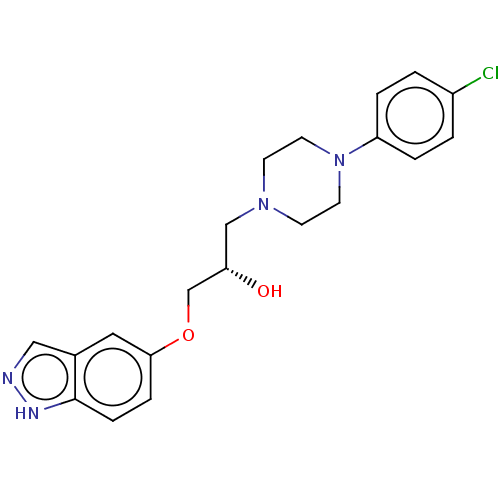
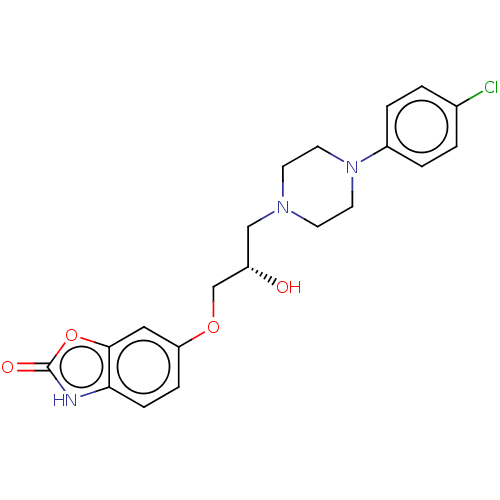
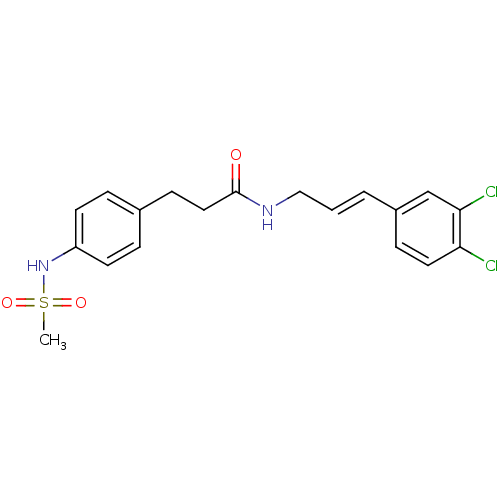
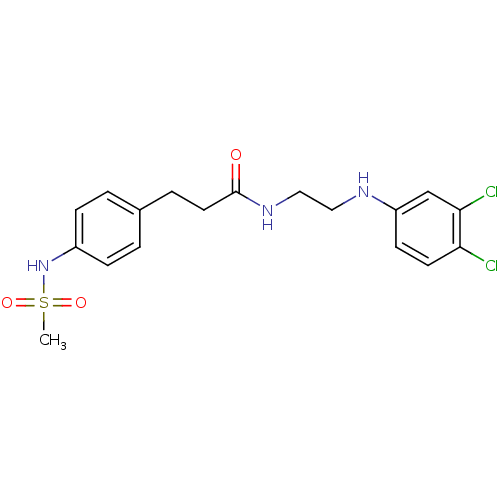
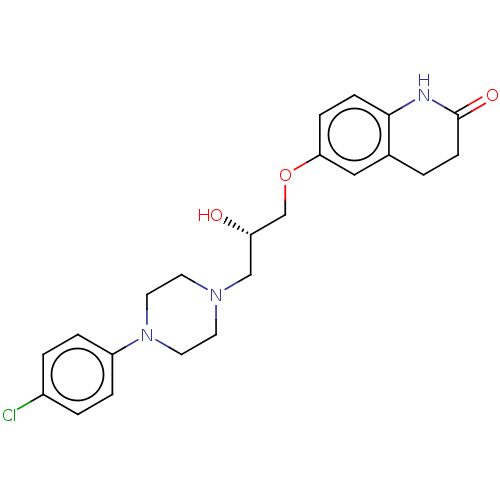
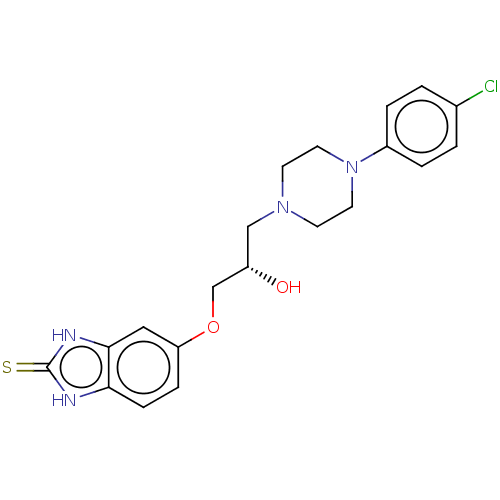
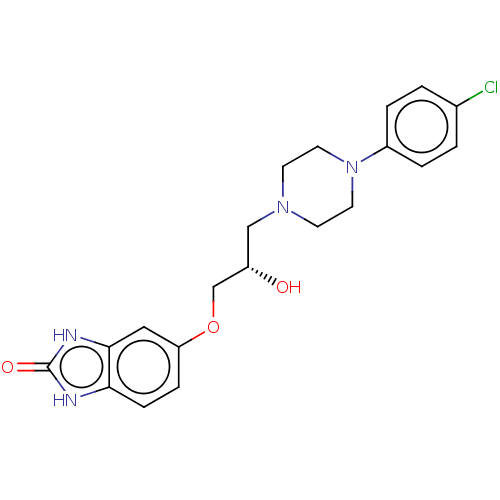
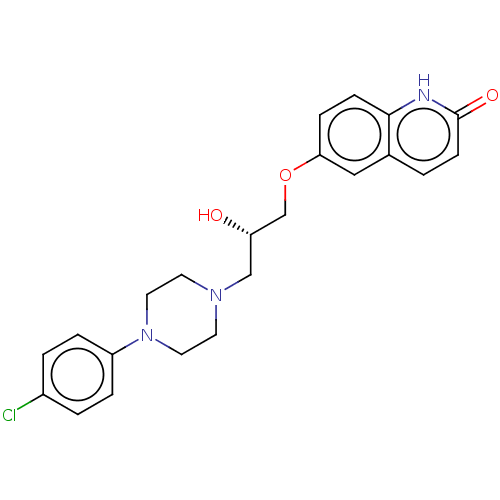
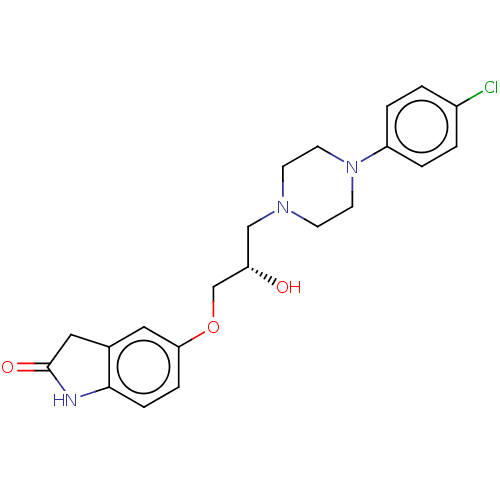
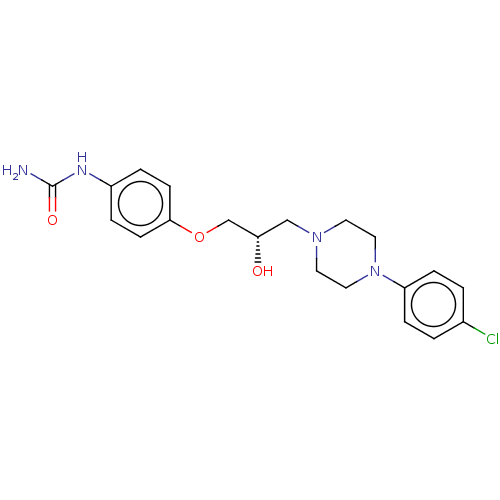
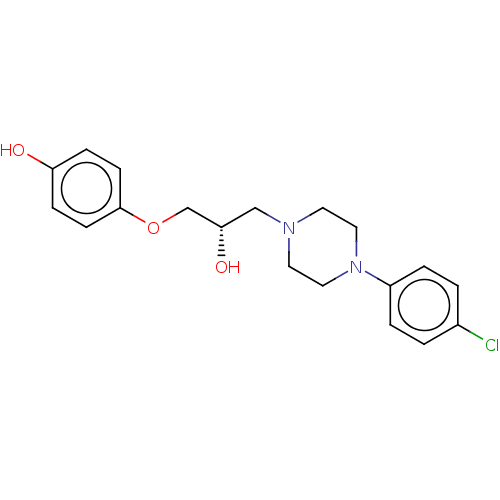
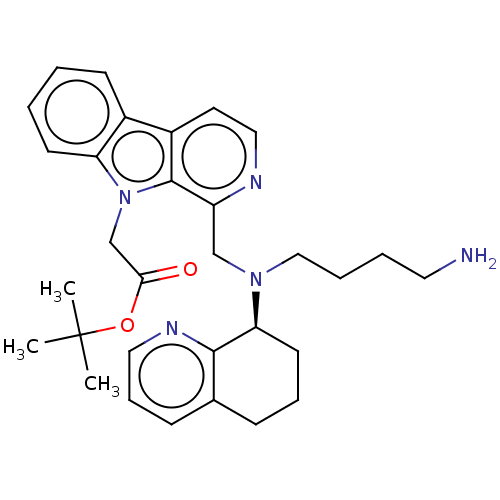
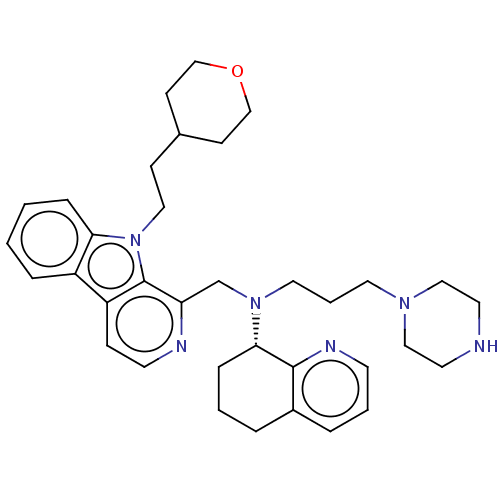
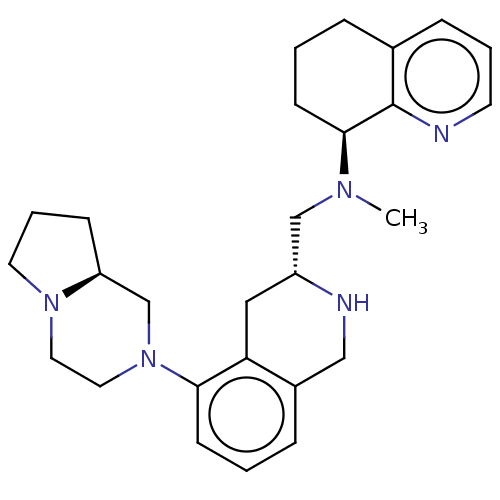
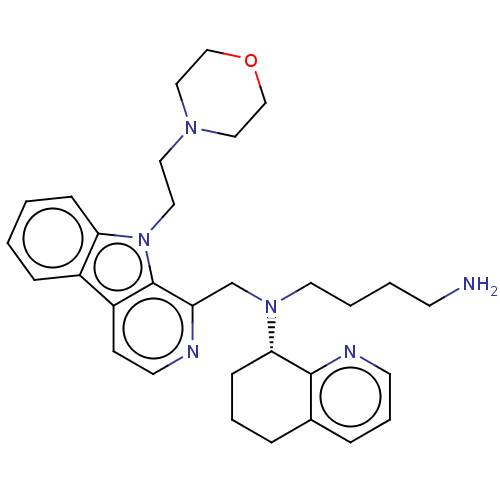
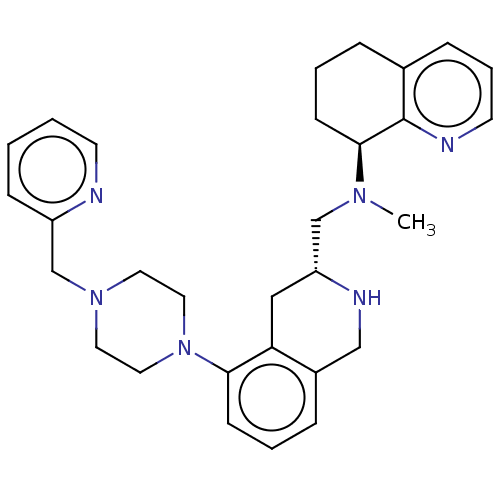
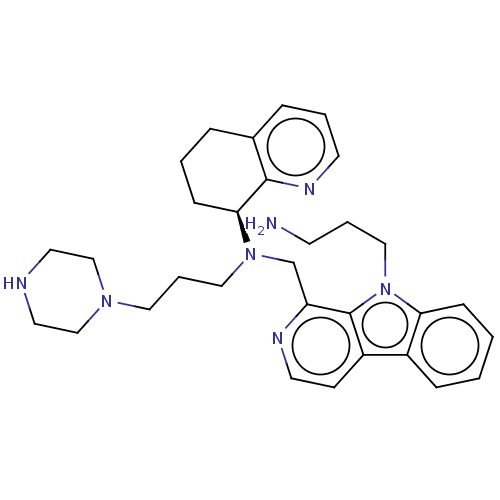
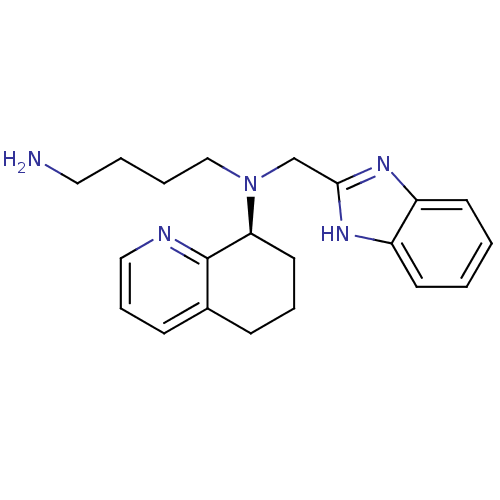
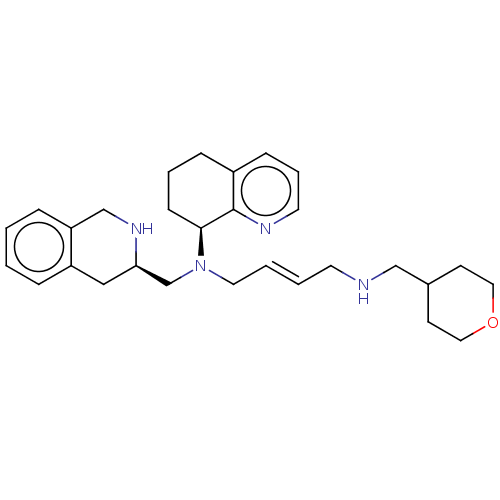
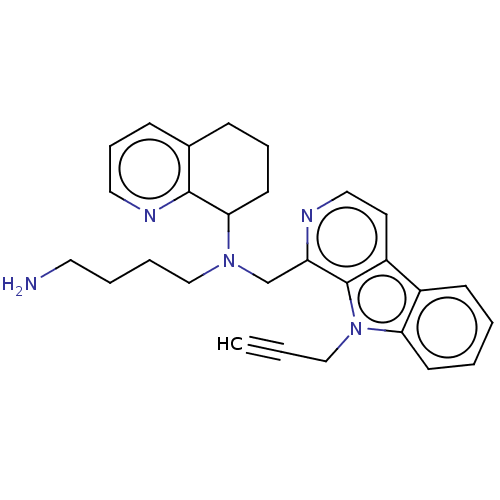
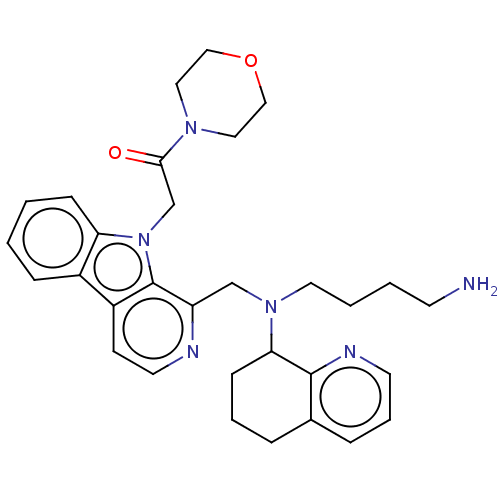
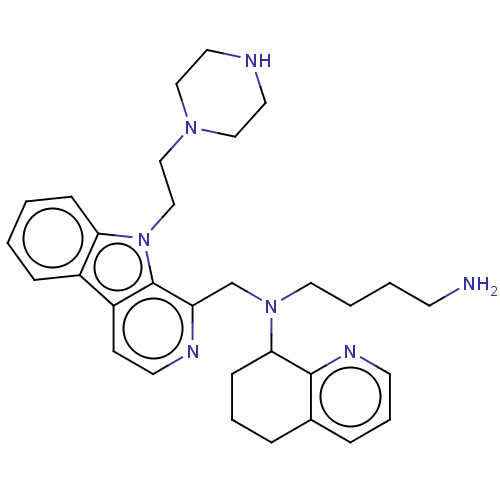
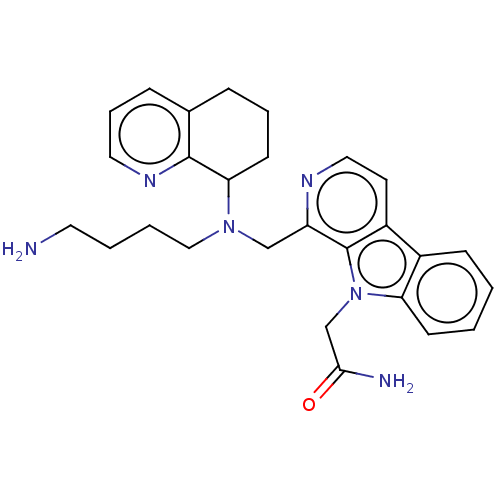
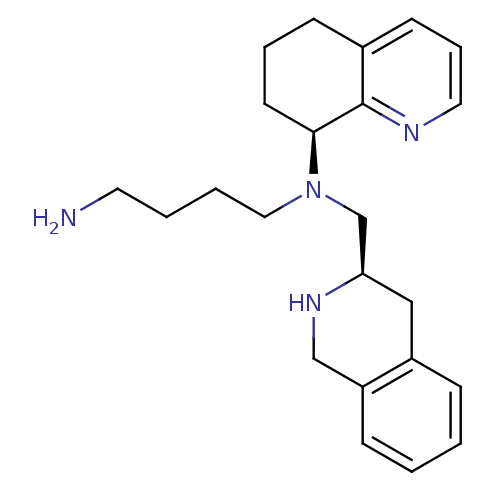
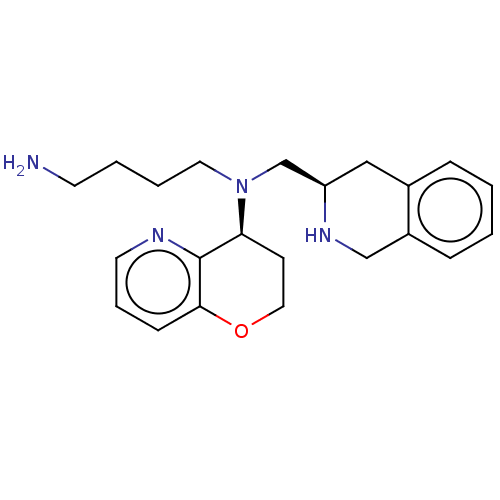
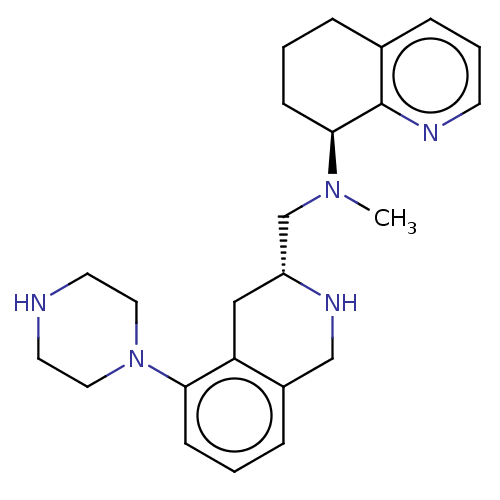
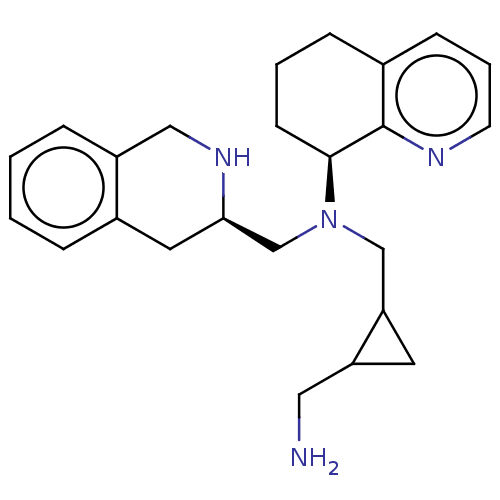
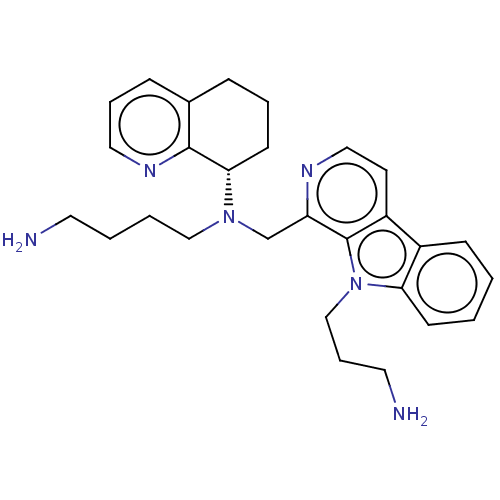
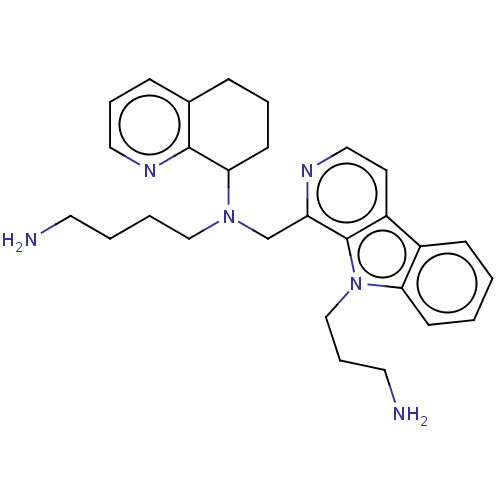
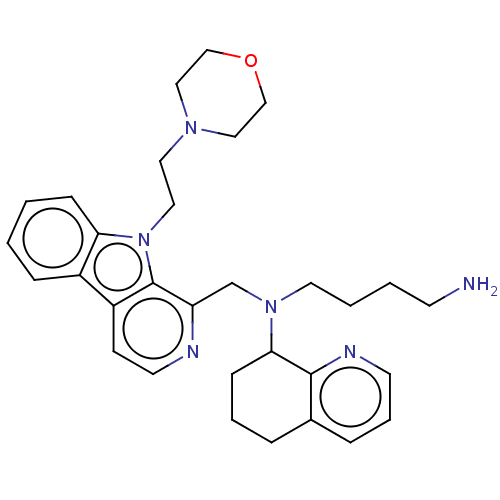
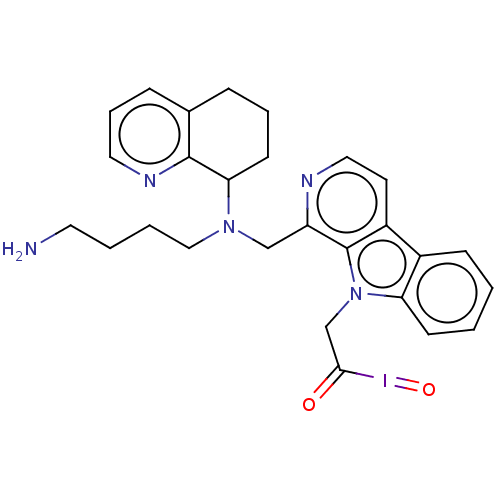
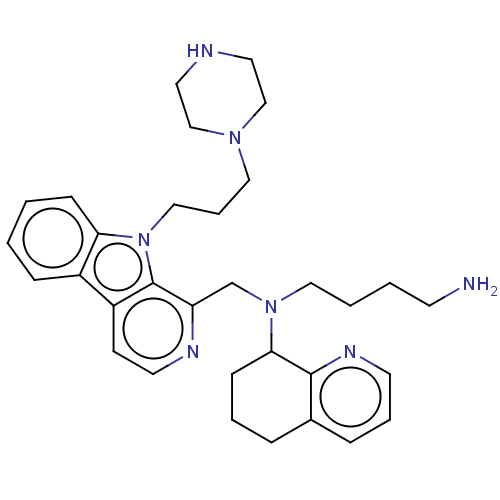
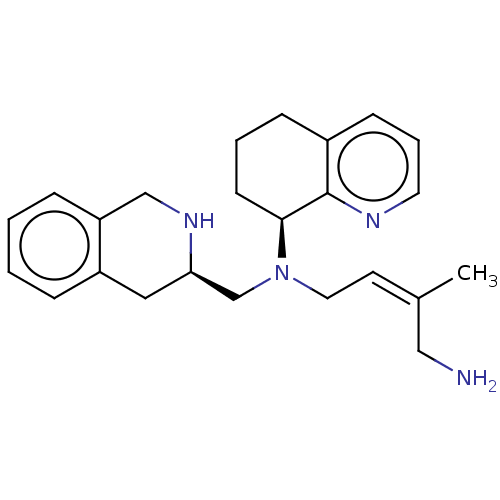
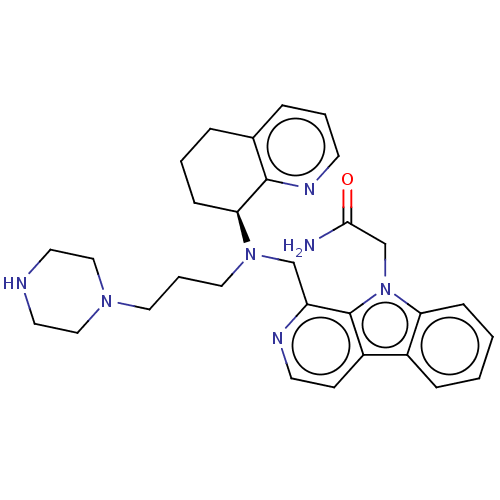
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 14nMAssay Description:Displacement of [3H]astemizole from human recombinant ERG expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 119nMAssay Description:Displacement of [3H]ifenprodil form NR2B receptor in Wistar rat cerebral cortex membraneMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 300nMAssay Description:Displacement of [3H]astemizole from human recombinant ERG expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 300nMAssay Description:Displacement of [3H]ifenprodil form NR2B receptor in Wistar rat cerebral cortex membraneMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 553nMAssay Description:Compounds were evaluated for binding to the human ether-a-go-go potassium channel (hERG) expressed in HEK293 cells by displacement of 3[H]-astemizole...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 817nMAssay Description:Displacement of [3H]ifenprodil form NR2B receptor in Wistar rat cerebral cortex membraneMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 1.00E+3nMAssay Description:Compounds were evaluated for binding to the human ether-a-go-go potassium channel (hERG) expressed in HEK293 cells by displacement of 3[H]-astemizole...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 1.60E+3nMAssay Description:Compounds were evaluated for binding to the human ether-a-go-go potassium channel (hERG) expressed in HEK293 cells by displacement of 3[H]-astemizole...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 3.00E+3nMAssay Description:Displacement of [3H]astemizole from human recombinant ERG expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 4.62E+3nMAssay Description:Displacement of [3H]astemizole from human recombinant ERG expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 5.00E+3nMAssay Description:Compounds were evaluated for binding to the human ether-a-go-go potassium channel (hERG) expressed in HEK293 cells by displacement of 3[H]-astemizole...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 7.50E+3nMAssay Description:Compounds were evaluated for binding to the human ether-a-go-go potassium channel (hERG) expressed in HEK293 cells by displacement of 3[H]-astemizole...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 7.50E+3nMAssay Description:Compounds were evaluated for binding to the human ether-a-go-go potassium channel (hERG) expressed in HEK293 cells by displacement of 3[H]-astemizole...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 7.50E+3nMAssay Description:Compounds were evaluated for binding to the human ether-a-go-go potassium channel (hERG) expressed in HEK293 cells by displacement of 3[H]-astemizole...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 1.00E+4nMAssay Description:Compounds were evaluated for binding to the human ether-a-go-go potassium channel (hERG) expressed in HEK293 cells by displacement of 3[H]-astemizole...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 1.30E+4nMAssay Description:Compounds were evaluated for binding to the human ether-a-go-go potassium channel (hERG) expressed in HEK293 cells by displacement of 3[H]-astemizole...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Emory University
Curated by ChEMBL
Emory University
Curated by ChEMBL
Affinity DataKi: 3.90E+4nMAssay Description:Compounds were evaluated for binding to the human ether-a-go-go potassium channel (hERG) expressed in HEK293 cells by displacement of 3[H]-astemizole...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 2.90nMAssay Description:Antagonist activity at CXCR4 in human CCRF-CEM cells assessed as inhibition of SDF-1alpha-induced calcium release preincubated for 25 mins followed b...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60nMAssay Description:Antagonist activity at CXCR4 in human CCRF-CEM cells assessed as inhibition of SDF-1alpha-induced calcium release preincubated for 25 mins followed b...More data for this Ligand-Target Pair
Affinity DataIC50: 3.70nMAssay Description:Antagonist activity at human CXCR4 in CCRF-CEM cells assessed as decrease in SDF-1alpha stimulated Ca2+ flux preincubated for 25 mins followed by SDF...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 4.5nMAssay Description:Antagonist activity at CXCR4 in human CCRF-CEM cells assessed as inhibition of SDF-1alpha-induced calcium release preincubated for 25 mins followed b...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Antagonist activity at CXCR4 in human CCRF-CEM cells assessed as inhibition of SDS1alpha-induced calcium flux pretreated for 25 mins followed by SDS1...More data for this Ligand-Target Pair
Affinity DataIC50: 5.20nMAssay Description:Antagonist activity at CXCR4 receptor in human CCRF-CEM cells assessed as inhibition of SDF-1alpha-induced calcium release preincubated for 25 mins f...More data for this Ligand-Target Pair
Affinity DataIC50: 5.5nMAssay Description:Antagonist activity at mouse CXCR4More data for this Ligand-Target Pair
Affinity DataIC50: 5.5nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 5.60nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 5.70nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 5.80nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Antagonist activity at CXCR4 in human CCRF-CEM cells assessed as inhibition of SDS1alpha-induced calcium flux pretreated for 25 mins followed by SDS1...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Antagonist activity at CXCR4 in human CCRF-CEM cells assessed as inhibition of SDF1alpha-induced calcium flux pretreated for 25 mins followed by SDF1...More data for this Ligand-Target Pair
Affinity DataIC50: 6.10nMAssay Description:Antagonist activity at CXCR4 in human CCRF-CEM cells assessed as inhibition of SDF-1alpha-induced calcium release preincubated for 25 mins followed b...More data for this Ligand-Target Pair
Affinity DataIC50: 6.30nMAssay Description:Antagonist activity at CXCR4 in human CCRF-CEM cells assessed as inhibition of SDF-1alpha-induced calcium release preincubated for 25 mins followed b...More data for this Ligand-Target Pair
Affinity DataIC50: 6.30nMAssay Description:Antagonist activity at human CXCR4 in CCRF-CEM cells assessed as decrease in SDF-1alpha stimulated Ca2+ flux preincubated for 25 mins followed by SDF...More data for this Ligand-Target Pair
Affinity DataIC50: 6.30nMAssay Description:Antagonist activity at CXCR4 receptor in human CCRF-CEM cells assessed as inhibition of SDF-1alpha-induced calcium release preincubated for 25 mins f...More data for this Ligand-Target Pair
Affinity DataIC50: 6.40nMAssay Description:Antagonist activity at CXCR4 in human CCRF-CEM cells assessed as inhibition of CXCL12-mediated calcium flux incubated for 25 mins followed by CXCL12 ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.40nMAssay Description:Antagonist activity at CXCR4 in human CCRF-CEM cells assessed as inhibition of CXCL12-mediated calcium flux incubated for 25 mins followed by CXCL12 ...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Antagonist activity at CXCR4 in human CCRF-CEM cells assessed as inhibition of SDF1alpha-induced calcium flux pretreated for 25 mins followed by SDF1...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair
Affinity DataIC50: 7.30nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 7.70nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 8.10nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 8.30nMAssay Description:This assay measures the change in impedance that occurs when cells are stimulated with SDF-1a. Changes in shape and cytoskeleton result in a change o...More data for this Ligand-Target Pair
Affinity DataIC50: 8.60nMAssay Description:Antagonist activity at CXCR4 in human CCRF-CEM cells assessed as inhibition of CXCL12-mediated calcium flux incubated for 25 mins followed by CXCL12 ...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMT: 2°CAssay Description:Functional modulation of CXCR4 was determined by calcium mobilization assay using leukemic lymphoid CEM cells, which naturally express high levels of...More data for this Ligand-Target Pair