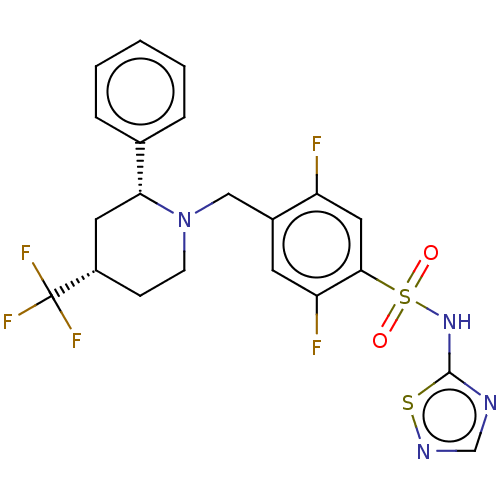
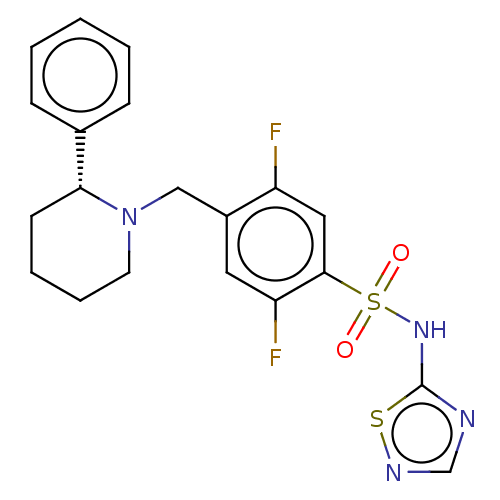
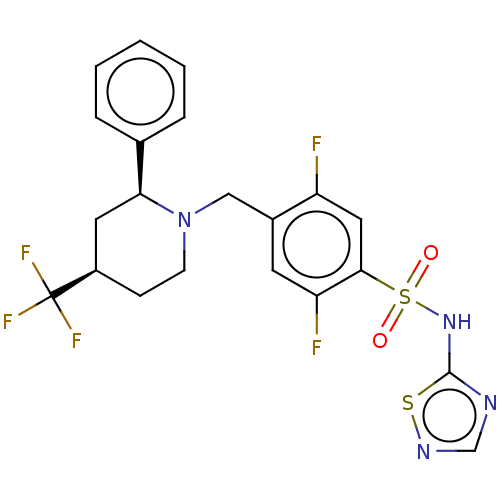
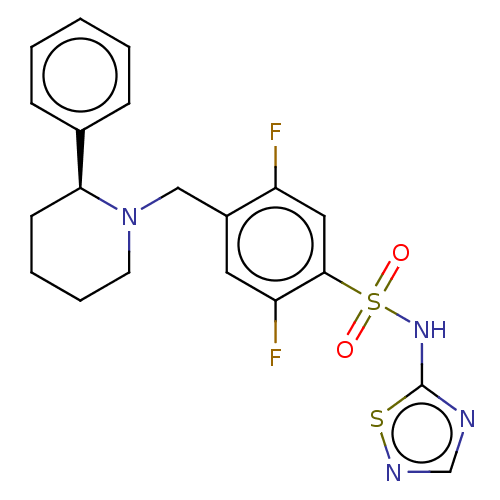
Affinity DataKi: 0.320nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 0.510nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 0.530nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 0.680nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 0.760nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 0.790nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 0.850nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 0.930nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 0.970nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 2.90nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 2.90nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 3.90nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: 29nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 72nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: 509nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: >700nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
Affinity DataKi: >700nMAssay Description:Displacement of [3H]GX-545 from full length human Nav1.7 VSD4 domain expressed in HEK cell membranes measured after 20 hrs by liquid scintillation co...More data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: 2.90E+3nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: 3.32E+3nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: 4.60E+3nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: 5.25E+3nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: 9.76E+3nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: 1.10E+4nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: 4.09E+4nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]Ifenprodil from NMDAR-2B in Sprague-Dawley rat frontal cortex homogenates after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 0.520nMAssay Description:Inhibition of AChE in Wistar rat brain homogenates using acetylthiocholine iodide and DTNB as substrate after 10 mins by Ellman methodMore data for this Ligand-Target Pair
Affinity DataIC50: 0.570nMAssay Description:Inhibition of full length human Nav1.7 expressed in HEK293 cells assessed as reduction of current amplitude at -60 mV holding potential measured afte...More data for this Ligand-Target Pair
Affinity DataIC50: 1.03nMAssay Description:Inhibition of AChE in Wistar rat brain homogenates using acetylthiocholine iodide and DTNB as substrate after 10 mins by Ellman methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.16nMAssay Description:Inhibition of AChE in Wistar rat brain homogenates using acetylthiocholine iodide and DTNB as substrate after 10 mins by Ellman methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.33nMAssay Description:Inhibition of AChE in Wistar rat brain homogenates using acetylthiocholine iodide and DTNB as substrate after 10 mins by Ellman methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.79nMAssay Description:Inhibition of AChE in Wistar rat brain homogenates using acetylthiocholine iodide and DTNB as substrate after 10 mins by Ellman methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.32nMAssay Description:Inhibition of AChE in Wistar rat brain homogenates using acetylthiocholine iodide and DTNB as substrate after 10 mins by Ellman methodMore data for this Ligand-Target Pair
Affinity DataIC50: 4.31nMAssay Description:Inhibition of AChE in Wistar rat brain homogenates using acetylthiocholine iodide and DTNB as substrate after 10 mins by Ellman methodMore data for this Ligand-Target Pair
TargetN-acylethanolamine-hydrolyzing acid amidase(Homo sapiens (Human))
Istituto Italiano Di Tecnologia
Curated by ChEMBL
Istituto Italiano Di Tecnologia
Curated by ChEMBL
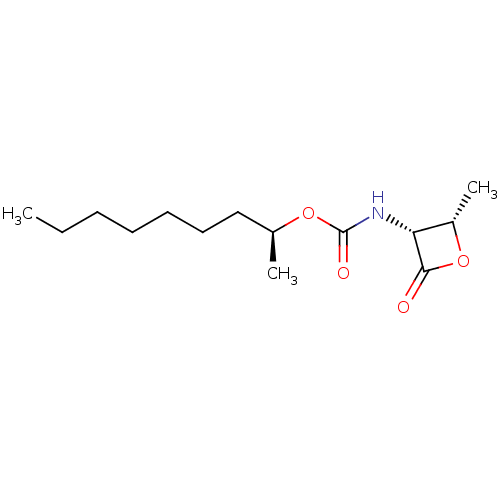
Affinity DataIC50: 5nMAssay Description:Inhibition of C-terminal His-6-tagged recombinant human spleen NAAA enzyme expressed in HEK293 cellsMore data for this Ligand-Target Pair

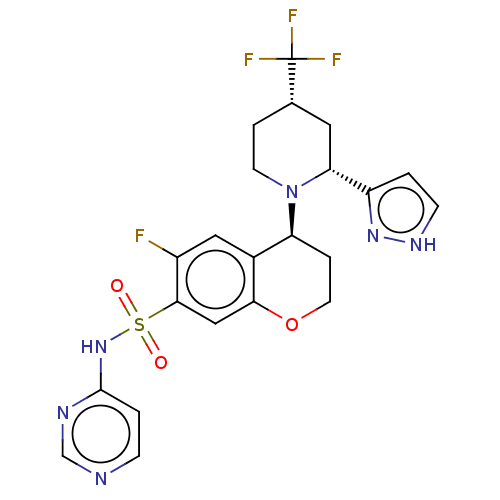
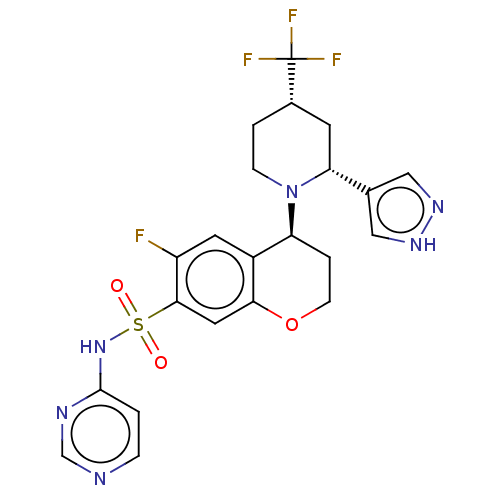
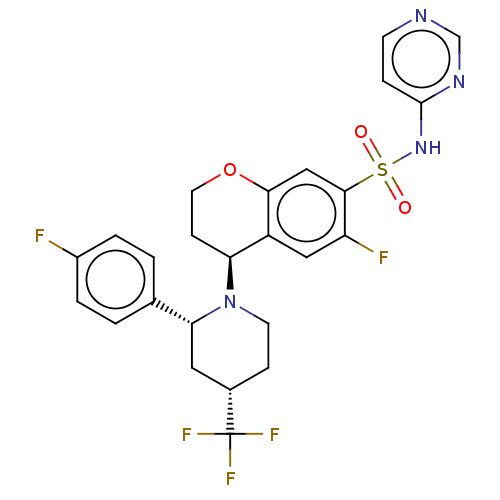
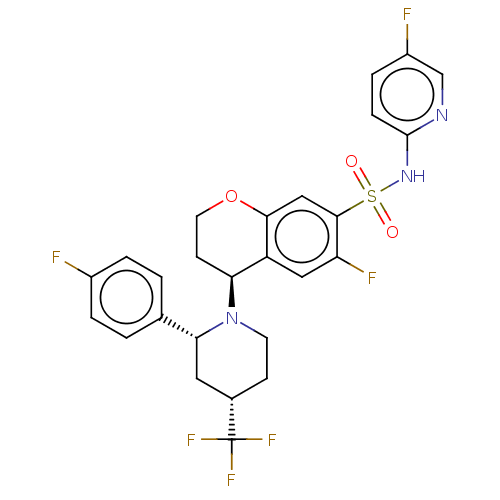
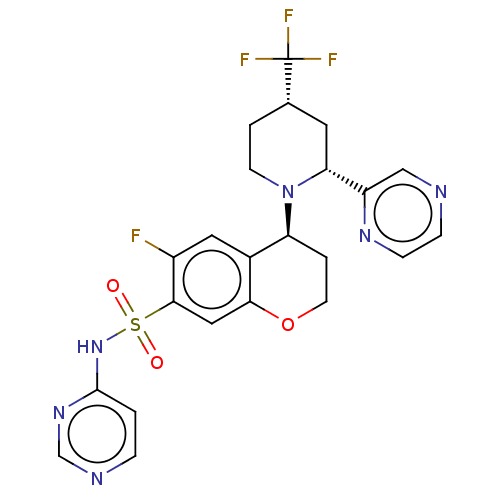
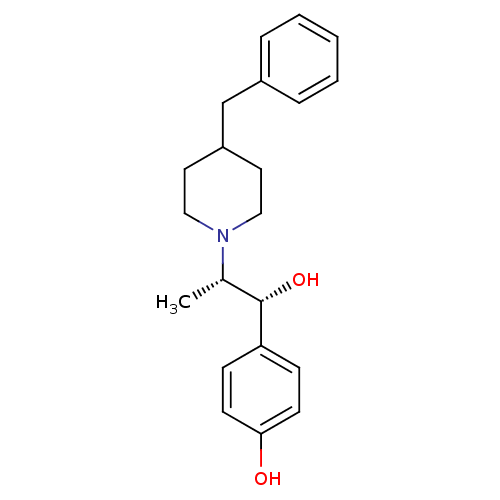
 3D Structure (crystal)
3D Structure (crystal)