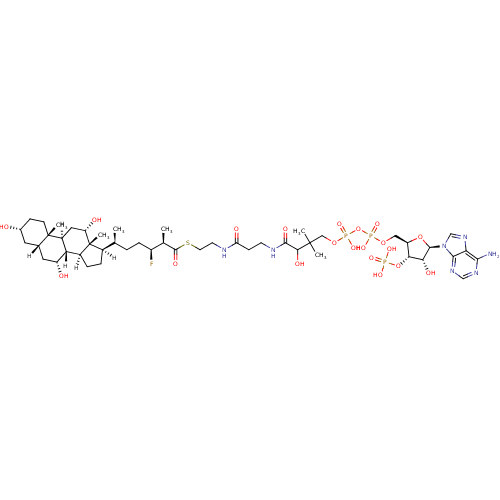
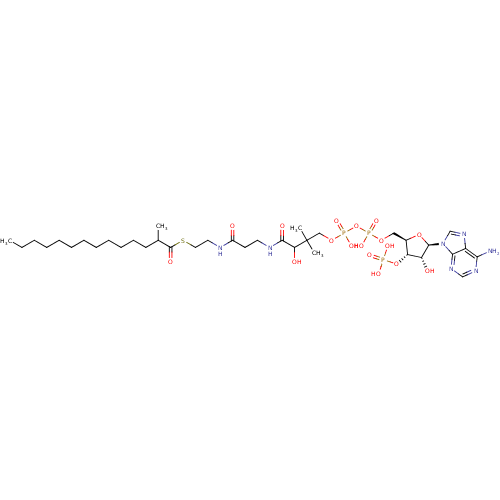
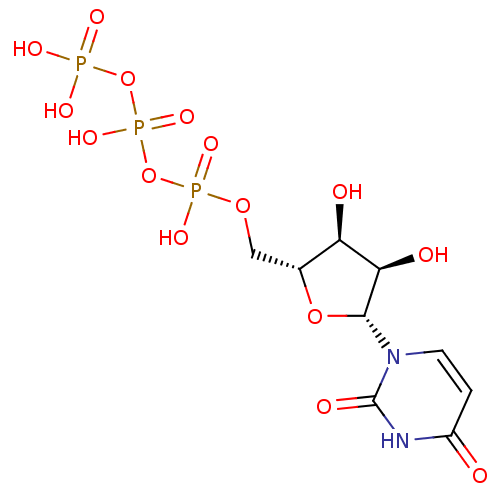
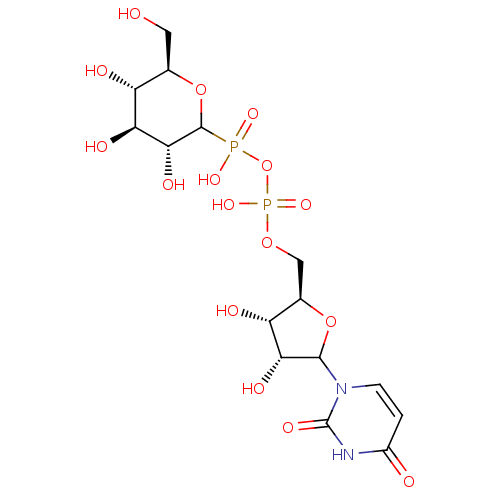
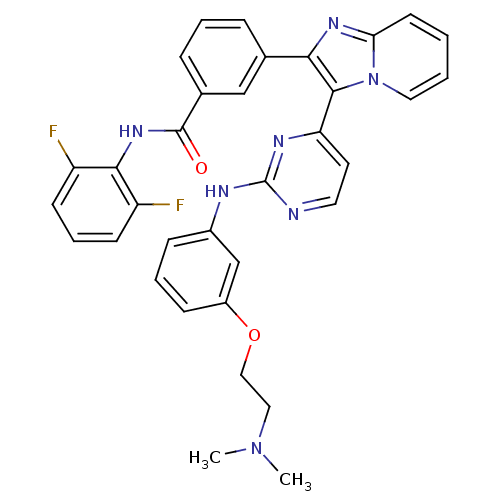
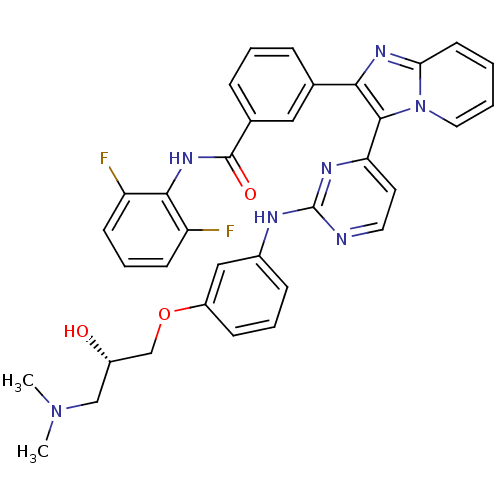
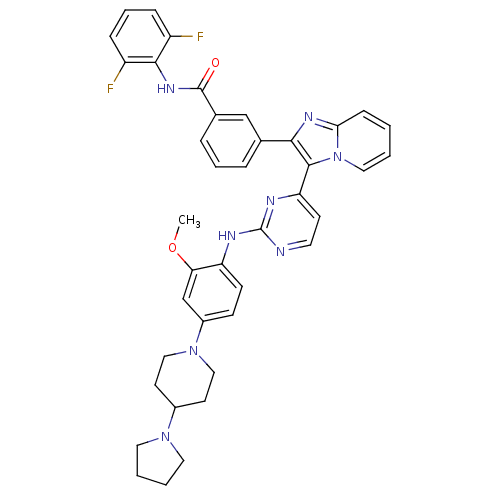
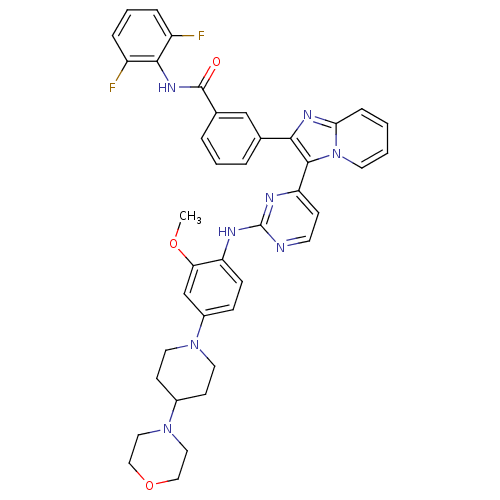
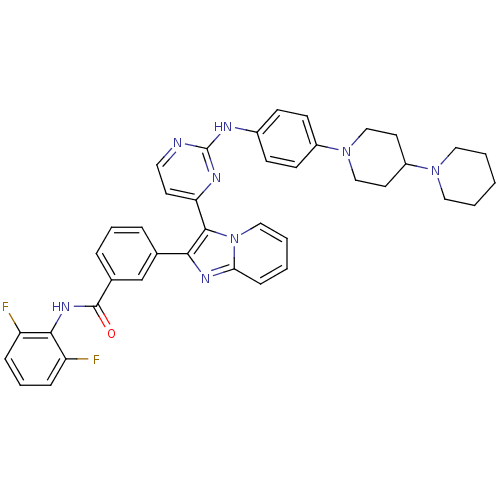
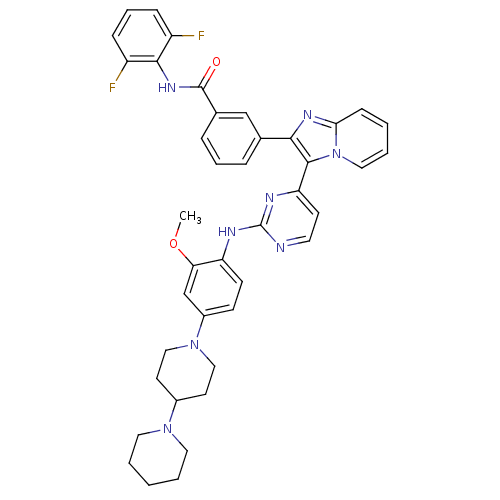
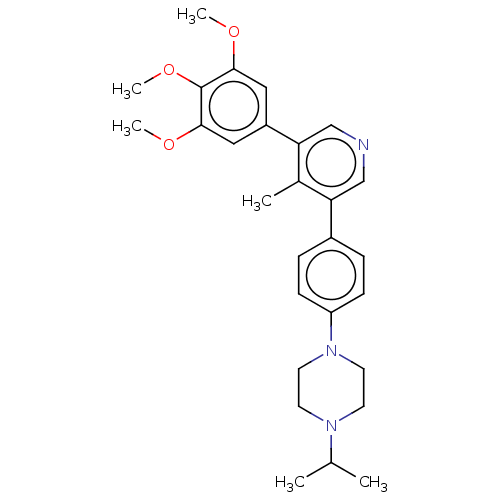
Affinity DataKi: 1.30nMAssay Description:Binding affinity to insulin receptor by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Binding affinity to IGF1R by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Binding affinity to insulin receptor by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 5.20nMAssay Description:Binding affinity to IGF1R by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 900nM ΔG°: -35.9kJ/molepH: 7.0 T: 2°CAssay Description:AMACR activity was determined by monitoring the interconversion of the (25R/S)-isomer of THC-CoA. Reaction product was analyzed by resolution of (25S...More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+3nM ΔG°: -34.9kJ/molepH: 7.0 T: 2°CAssay Description:AMACR activity was determined by monitoring the interconversion of the (25R/S)-isomer of THC-CoA. Reaction product was analyzed by resolution of (25S...More data for this Ligand-Target Pair
Affinity DataKi: 2.30E+3nM ΔG°: -33.5kJ/molepH: 7.0 T: 2°CAssay Description:AMACR activity was determined by monitoring the interconversion of the (25R/S)-isomer of THC-CoA. Reaction product was analyzed by resolution of (25S...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase VIM-2(Pseudomonas aeruginosa (g-Proteobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 3.70E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase VIM-2(Pseudomonas aeruginosa (g-Proteobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 3.80E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 3.80E+3nM ΔG°: -32.2kJ/molepH: 7.0 T: 2°CAssay Description:AMACR activity was determined by monitoring the interconversion of the (25R/S)-isomer of THC-CoA. Reaction product was analyzed by resolution of (25S...More data for this Ligand-Target Pair
TargetVIM-24(Klebsiella pneumoniae (Enterobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 4.90E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 5.40E+3nM ΔG°: -31.3kJ/molepH: 7.0 T: 2°CAssay Description:AMACR activity was determined by monitoring the interconversion of the (25R/S)-isomer of THC-CoA. Reaction product was analyzed by resolution of (25S...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase VIM-2(Pseudomonas aeruginosa (g-Proteobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 5.40E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
TargetVIM-24(Klebsiella pneumoniae (Enterobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 6.00E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
TargetVIM-24(Klebsiella pneumoniae (Enterobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 1.10E+4nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
TargetVIM-24(Klebsiella pneumoniae (Enterobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 1.20E+4nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase VIM-2(Pseudomonas aeruginosa (g-Proteobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 1.40E+4nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 1.92E+4nM ΔG°: -28.0kJ/molepH: 7.0 T: 2°CAssay Description:AMACR activity was determined by monitoring the interconversion of the (25R/S)-isomer of THC-CoA. Reaction product was analyzed by resolution of (25S...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+4nM ΔG°: -27.9kJ/molepH: 7.0 T: 2°CAssay Description:AMACR activity was determined by monitoring the interconversion of the (25R/S)-isomer of THC-CoA. Reaction product was analyzed by resolution of (25S...More data for this Ligand-Target Pair
Affinity DataKi: 4.50E+4nM ΔG°: -25.8kJ/molepH: 7.0 T: 2°CAssay Description:AMACR activity was determined by monitoring the interconversion of the (25R/S)-isomer of THC-CoA. Reaction product was analyzed by resolution of (25S...More data for this Ligand-Target Pair
Affinity DataKi: 5.60E+4nM ΔG°: -25.2kJ/molepH: 7.0 T: 2°CAssay Description:AMACR activity was determined by monitoring the interconversion of the (25R/S)-isomer of THC-CoA. Reaction product was analyzed by resolution of (25S...More data for this Ligand-Target Pair
Affinity DataKi: 1.28E+5nMAssay Description:Kinetic constant for galactosyltransferase inhibition was determinedMore data for this Ligand-Target Pair
Affinity DataKi: 1.37E+5nM ΔG°: -22.9kJ/molepH: 7.0 T: 2°CAssay Description:AMACR activity was determined by monitoring the interconversion of the (25R/S)-isomer of THC-CoA. Reaction product was analyzed by resolution of (25S...More data for this Ligand-Target Pair
Affinity DataKi: 1.65E+5nMAssay Description:Kinetic constant for galactosyltransferase inhibition was determinedMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
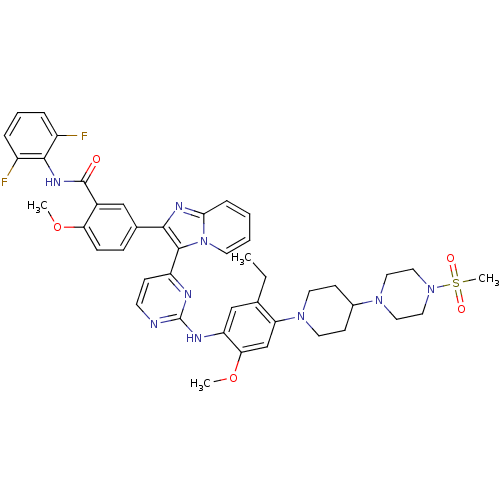
TargetActivin receptor type-1(Homo sapiens (Human))
Ontario Institute For Cancer Research
Curated by ChEMBL
Ontario Institute For Cancer Research
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of human ALK2 using casein as substrate in presence of [gamma-33P]-ATP by radiometric hotspot assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
TargetActivin receptor type-1(Homo sapiens (Human))
Ontario Institute For Cancer Research
Curated by ChEMBL
Ontario Institute For Cancer Research
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of human ALK2 using casein as substrate in presence of [gamma-33P]-ATP by radiometric hotspot assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of GST-tagged insulin receptor expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
TargetActivin receptor type-1(Homo sapiens (Human))
Ontario Institute For Cancer Research
Curated by ChEMBL
Ontario Institute For Cancer Research
Curated by ChEMBL
Affinity DataIC50: 5nMAssay Description:Inhibition of human ALK2 using casein as substrate in presence of [gamma-33P]-ATP by radiometric hotspot assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
TargetActivin receptor type-1(Homo sapiens (Human))
Ontario Institute For Cancer Research
Curated by ChEMBL
Ontario Institute For Cancer Research
Curated by ChEMBL
Affinity DataIC50: 6nMAssay Description:Inhibition of human ALK2 using casein as substrate in presence of [gamma-33P]-ATP by radiometric hotspot assayMore data for this Ligand-Target Pair
TargetActivin receptor type-1(Homo sapiens (Human))
Ontario Institute For Cancer Research
Curated by ChEMBL
Ontario Institute For Cancer Research
Curated by ChEMBL
Affinity DataIC50: 6nMAssay Description:Competitive displacement of PBI-6908 from nanoluciferase-fused ALK2 G328V mutant (unknown origin) expressed in HEK293 cells incubated for 2 hrs by Na...More data for this Ligand-Target Pair
TargetActivin receptor type-1(Homo sapiens (Human))
Ontario Institute For Cancer Research
Curated by ChEMBL
Ontario Institute For Cancer Research
Curated by ChEMBL
Affinity DataIC50: 6nMAssay Description:Inhibition of human ALK2 using casein as substrate in presence of [gamma-33P]-ATP by radiometric hotspot assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Inhibition of GST-tagged insulin receptor expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Inhibition of GST-tagged IGF1R (957-1367) (unknown origin) expressed in baculovirus by time-resolved fluorescence assayMore data for this Ligand-Target Pair
TargetActivin receptor type-1(Homo sapiens (Human))
Ontario Institute For Cancer Research
Curated by ChEMBL
Ontario Institute For Cancer Research
Curated by ChEMBL
Affinity DataIC50: 7nMAssay Description:Inhibition of human ALK2 using casein as substrate in presence of [gamma-33P]-ATP by radiometric hotspot assayMore data for this Ligand-Target Pair
TargetActivin receptor type-1(Homo sapiens (Human))
Ontario Institute For Cancer Research
Curated by ChEMBL
Ontario Institute For Cancer Research
Curated by ChEMBL
Affinity DataIC50: 7nMAssay Description:Inhibition of human ALK2 using casein as substrate in presence of [gamma-33P]-ATP by radiometric hotspot assayMore data for this Ligand-Target Pair
TargetActivin receptor type-1(Homo sapiens (Human))
Ontario Institute For Cancer Research
Curated by ChEMBL
Ontario Institute For Cancer Research
Curated by ChEMBL
Affinity DataIC50: 7nMAssay Description:Competitive displacement of PBI-6908 from nanoluciferase-fused ALK2 Q207D mutant (unknown origin) expressed in HEK293 cells incubated for 2 hrs by Na...More data for this Ligand-Target Pair
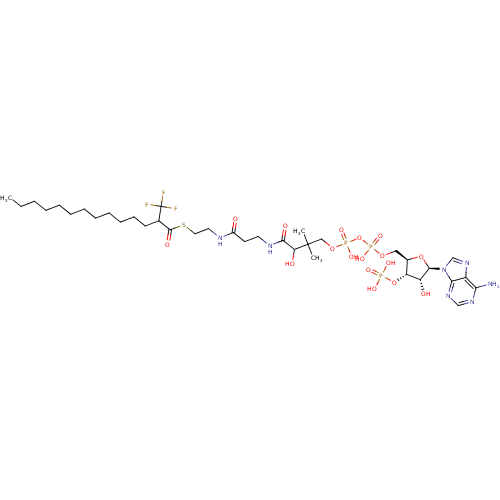



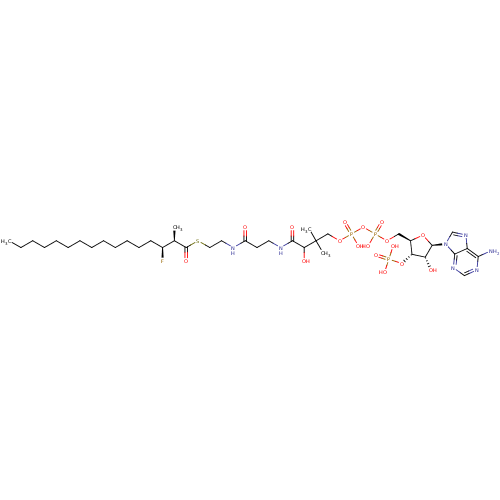
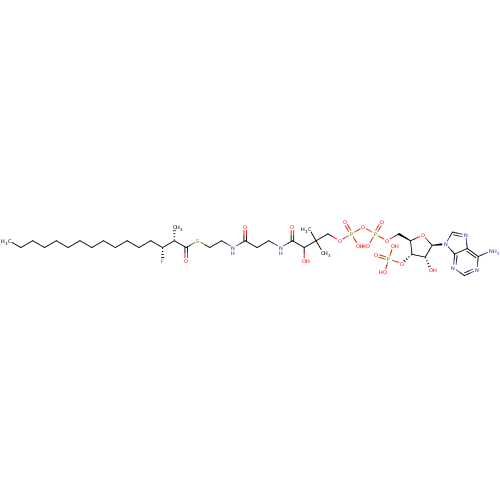

 3D Structure (crystal)
3D Structure (crystal)