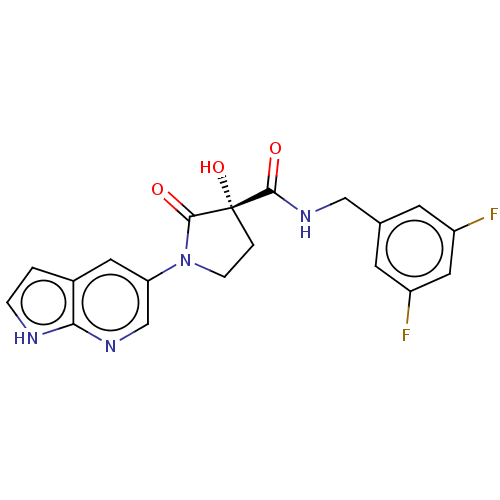
Affinity DataKi: 0.900nMAssay Description:Inhibition of 5-[(S)-3-(3-Chloro-5-fluoro-benzylcarbamoyl)-3-hydroxy-2-oxopyrrolidin-1-yl]-1H-indole-2-carboxylic Acid (3-Amino-propyl)-amide-Dy647 b...More data for this Ligand-Target Pair
Affinity DataKi: 0.900nMAssay Description:Inhibition of 5-[(S)-3-(3-Chloro-5-fluoro-benzylcarbamoyl)-3-hydroxy-2-oxopyrrolidin-1-yl]-1H-indole-2-carboxylic Acid (3-Amino-propyl)-amide-Dy647 b...More data for this Ligand-Target Pair
Affinity DataIC50: 55nMAssay Description:Inhibition of MetAP2 in HUVEC assessed as reduction in viability incubated for 3 days by CyQUANT Direct Cell proliferation assayMore data for this Ligand-Target Pair
Affinity DataIC50: 55nMAssay Description:Inhibition of MetAP2 in HUVEC assessed as reduction in viability incubated for 3 days by CyQUANT Direct Cell proliferation assayMore data for this Ligand-Target Pair
Affinity DataIC50: 87nMAssay Description:Inhibition of recombinant human N-terminal His-tagged MetAP2 (2 to 478 residues) using Met-Ala-Ser as substrate and MnCl2 as co-facor preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 87nMAssay Description:Inhibition of recombinant human N-terminal His-tagged MetAP2 (2 to 478 residues) using Met-Ala-Ser as substrate and MnCl2 as co-facor preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 91nMAssay Description:The MetAP-2 activity is determined by coupling enzymatic reactions. The tripeptide Met-Arg-Ser (MAS) is employed as substrate. The methionine liberat...More data for this Ligand-Target Pair
Affinity DataKd: 0.880nMAssay Description:Inhibition of 5-[(S)-3-(3-Chloro-5-fluoro-benzylcarbamoyl)-3-hydroxy-2-oxopyrrolidin-1-yl]-1H-indole-2-carboxylic Acid (3-Amino-propyl)-amide-Dy647 b...More data for this Ligand-Target Pair
Affinity DataKd: 0.880nMAssay Description:Inhibition of 5-[(S)-3-(3-Chloro-5-fluoro-benzylcarbamoyl)-3-hydroxy-2-oxopyrrolidin-1-yl]-1H-indole-2-carboxylic Acid (3-Amino-propyl)-amide-Dy647 b...More data for this Ligand-Target Pair