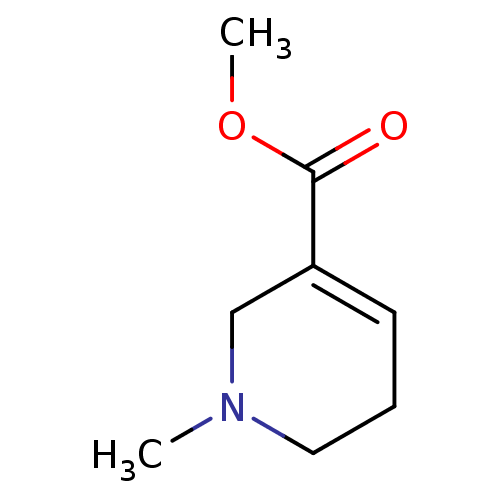
Found 32 Enz. Inhib. hit(s) with Target = 'Muscarinic acetylcholine receptor M1' and Ligand = 'BDBM46858'
Found 32 Enz. Inhib. hit(s) with Target = 'Muscarinic acetylcholine receptor M1' and Ligand = 'BDBM46858' Affinity DataKi: 3.30nMAssay Description:Binding affinity (low) of [3H](R)-QNB binding to Muscarinic acetylcholine receptor M1 expressed in A9 L cell line in the absence of 300 uM GTP-gamma-...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Homo sapiens (Human))
University of Arizona
Curated by PDSP Ki Database
University of Arizona
Curated by PDSP Ki Database
Affinity DataKi: 5.5nMAssay Description:Binding affinity of [3H]-(R)-QNB binding to Muscarinic acetylcholine receptor M1 expressed in A9 L cell line in the presence of 300 uM GTP-gamma-SMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Homo sapiens (Human))
University of Arizona
Curated by PDSP Ki Database
University of Arizona
Curated by PDSP Ki Database
Affinity DataKi: 14nMAssay Description:Inhibition of [3H]- oxotremorine-M binding to human Muscarinic acetylcholine receptor M1 expressed in CHO cell membranesMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Homo sapiens (Human))
University of Arizona
Curated by PDSP Ki Database
University of Arizona
Curated by PDSP Ki Database
Affinity DataKi: 2.90E+4nMAssay Description:Binding affinity of compound was determined towards human Muscarinic acetylcholine receptor M1 using [3H]-QNB radioligandMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Homo sapiens (Human))
University of Arizona
Curated by PDSP Ki Database
University of Arizona
Curated by PDSP Ki Database
Affinity DataKi: 2.95E+4nMAssay Description:Binding affinity to human cloned muscarinic M1 receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 8.60E+4nMAssay Description:Displacement of [3H]Quinuclidinyl benzillate from muscarinic M1 receptor in Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 8.60E+4nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor in Wistar rat brain cortex membrane after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 8.80E+4nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor in Wistar rat brain cortex after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 77nMAssay Description:In vitro binding affinity against rat hippocampus M1 receptor using [3H]-pirenzepine (Pz) as radioligandMore data for this Ligand-Target Pair
Affinity DataIC50: 77nMAssay Description:In vitro binding affinity against muscarinic acetylcholine receptor M1 from rat hippocampus, using [3H]-oxotremorine-M (Oxo-M) as radioligandMore data for this Ligand-Target Pair
Affinity DataIC50: 77nMAssay Description:Binding activity against muscarinic acetylcholine receptor M1 in rat brain, using [3H]OXO-M as the radioligand.More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Homo sapiens (Human))
University of Arizona
Curated by PDSP Ki Database
University of Arizona
Curated by PDSP Ki Database
Affinity DataIC50: 115nMAssay Description:Ability to displace [3H]-N-methylscopolamine from Muscarinic acetylcholine receptor M1 expressed in CHO cells.More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Homo sapiens (Human))
University of Arizona
Curated by PDSP Ki Database
University of Arizona
Curated by PDSP Ki Database
Affinity DataIC50: 545nMAssay Description:Efficacy at muscarinic acetylcholine receptor M1 measured by the ability to inhibit the electrically stimulated twitch of the rabbit vas deferensMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of [3H]pirenzepine binding to rat brain membrane Muscarinic acetylcholine receptor M1More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Displacement of [3H]-pirenzepine (Pz) from rat hippocampus muscarinic acetylcholine receptor M1More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:In vitro binding affinity against muscarinic acetylcholine receptor M1 from rat hippocampus, using [3H]-pirenzepine (Pz) as radioligandMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Binding activity against muscarinic acetylcholine receptor M1 in rat brain, using [3H]-Pz as the radioligand.More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Displacement of [3H]-Pz (pirenzepine) from the muscarinic receptor M1 of the rat hippocampusMore data for this Ligand-Target Pair
Affinity DataIC50: 1.74E+3nMAssay Description:Evaluated for inhibition of binding of [3H]pirenzepine to homogenized rat cerebral cortex membranesChecked by AuthorMore data for this Ligand-Target Pair
Affinity DataIC50: 1.74E+3nMAssay Description:Ability to displace [3H]-pirenzepine (pir) from muscarinic acetylcholine receptor M1 in rat cortical tissue.More data for this Ligand-Target Pair
Affinity DataIC50: 1.19E+4nMAssay Description:Compound was evaluated for the displacement of the muscarinic QNB in a genetically transformed rat cell line (m1c2) transfected with cloned m1 recept...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Homo sapiens (Human))
University of Arizona
Curated by PDSP Ki Database
University of Arizona
Curated by PDSP Ki Database
Affinity DataIC50: 6.50E+4nMAssay Description:In vitro affinity against human Muscarinic acetylcholine receptor M1 using quinuclidynyl benzylate (QNB)More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Homo sapiens (Human))
University of Arizona
Curated by PDSP Ki Database
University of Arizona
Curated by PDSP Ki Database
Affinity DataIC50: 1.16E+5nMAssay Description:Ability to displace [3H]-N-methylscopolamine from Muscarinic acetylcholine receptor M1 expressed in CHO cells.More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Homo sapiens (Human))
University of Arizona
Curated by PDSP Ki Database
University of Arizona
Curated by PDSP Ki Database
Affinity DataIC50: 1.45E+5nMAssay Description:Compound was evaluated for its ability to displace [3H]-N-methylscopolamine ([3H]-NMS) binding to cloned CHO cell lines expressing Muscarinic acetylc...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60E+5nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor in Wistar rat brain cortex after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 4.69E+5nMAssay Description:Displacement of [3H]Quinuclidinyl benzillate from muscarinic M1 receptor in Wistar rat cortex synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 4.69E+5nMAssay Description:Displacement of [3H]QNB from muscarinic M1 receptor in Wistar rat brain cortex membrane after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Homo sapiens (Human))
University of Arizona
Curated by PDSP Ki Database
University of Arizona
Curated by PDSP Ki Database
Affinity DataEC50: 5.60E+3nMAssay Description:Stimulation of phosphoinositide hydrolysis in A9L cells expressing human m1 receptorMore data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Homo sapiens (Human))
University of Arizona
Curated by PDSP Ki Database
University of Arizona
Curated by PDSP Ki Database
Affinity DataEC50: 3.20E+3nMAssay Description:In Vitro evaluation of the compound activity at the cloned Human Muscarinic acetylcholine receptor M1 determined by receptor selection and amplificat...More data for this Ligand-Target Pair
Affinity DataEC50: 2.70E+4nMAssay Description:Tested against Muscarinic acetylcholine receptor M1 expressed in A9 L cell lineMore data for this Ligand-Target Pair