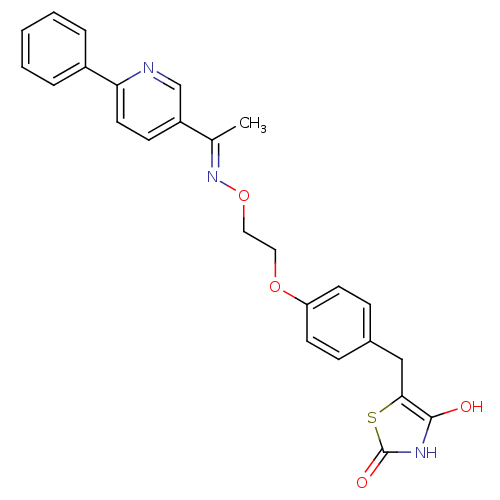
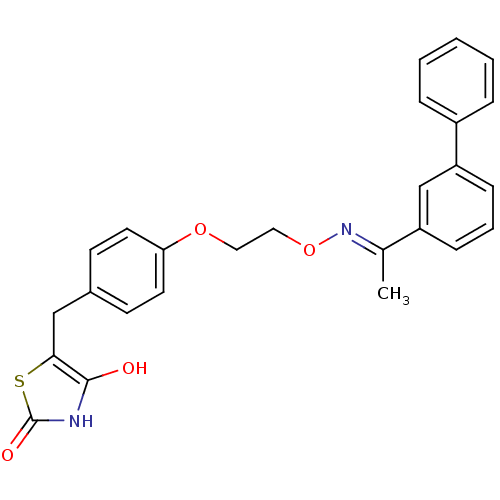
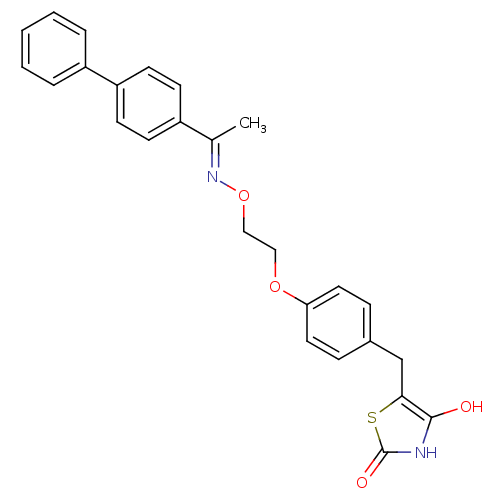
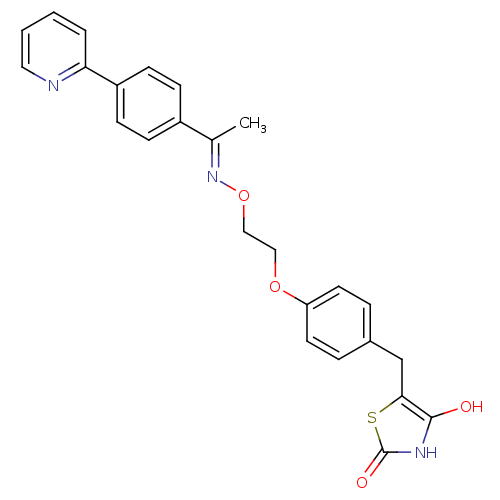
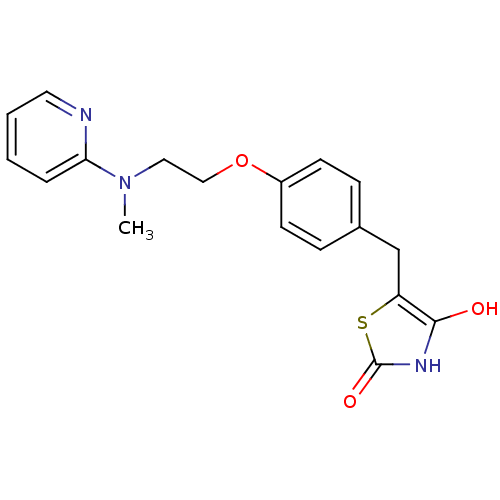
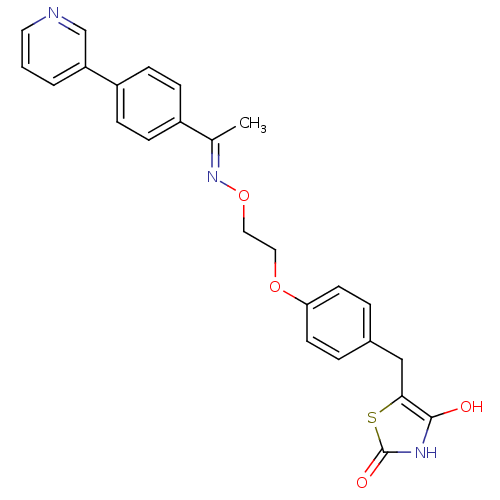
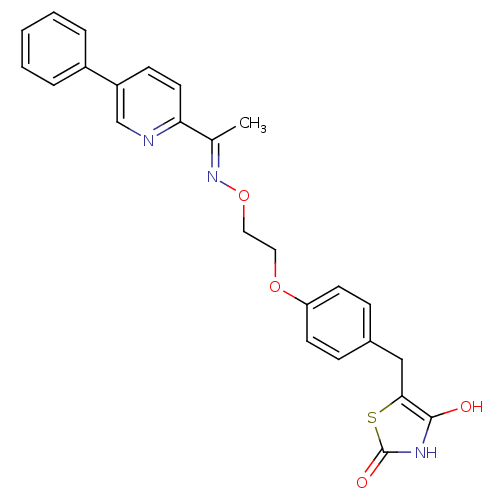
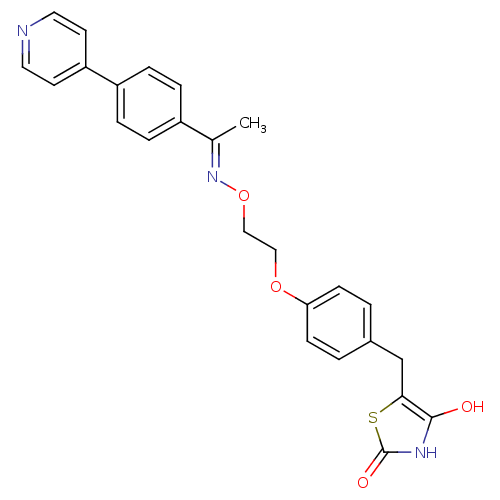
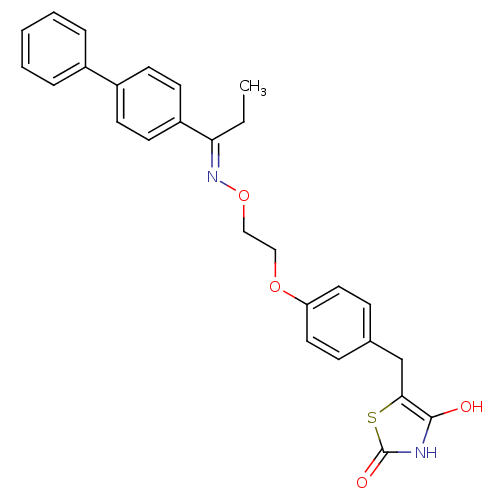
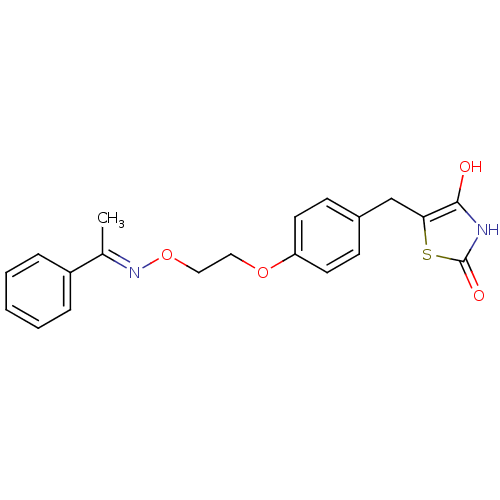
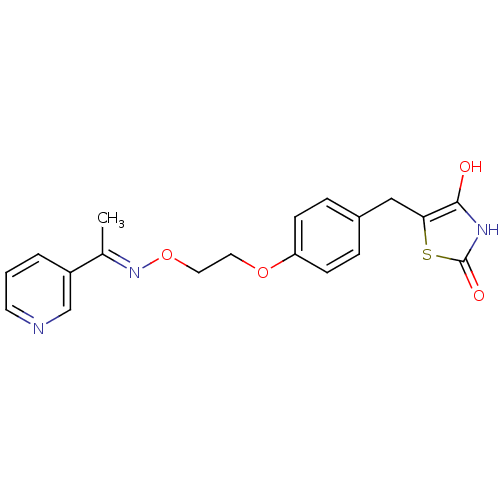
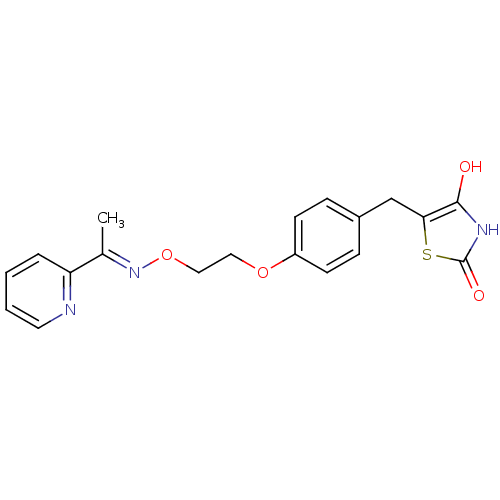
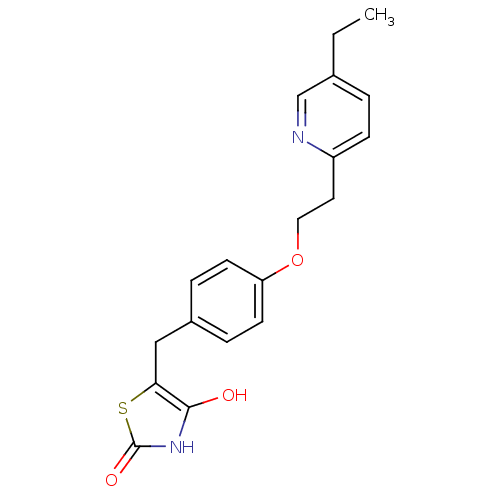
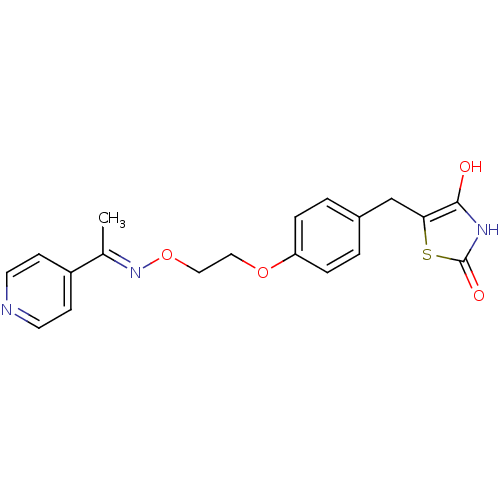
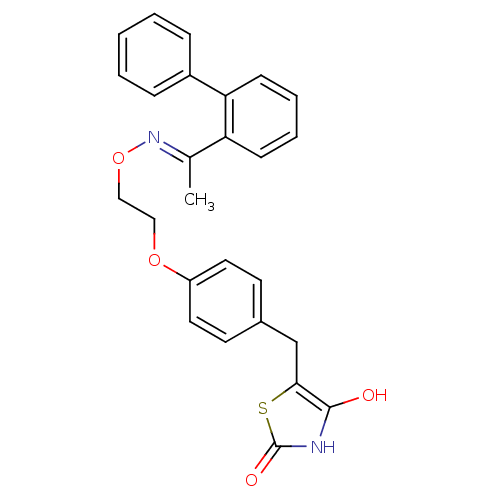
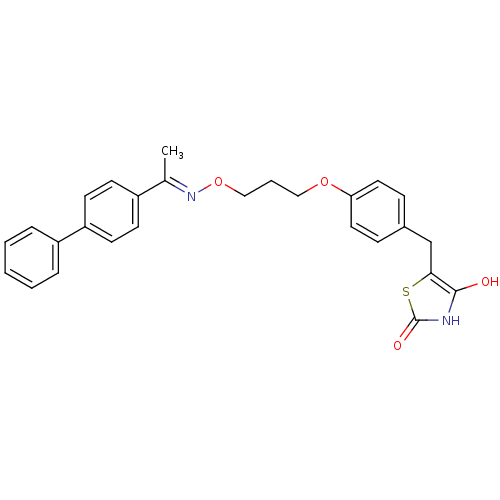
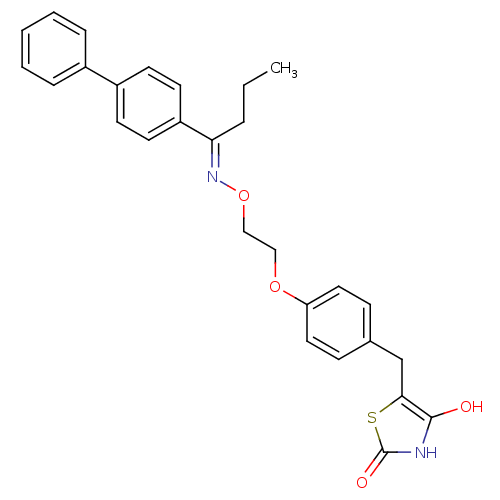
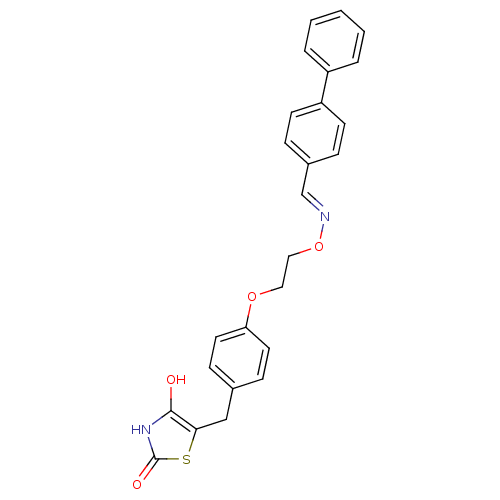
Report error Found 18 Enz. Inhib. hit(s) with all data for entry = 50035108
Affinity DataEC50: 190nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: 330nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: 400nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: 640nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: 730nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: 800nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: 1.10E+3nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: 1.60E+3nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: 1.60E+3nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: 1.70E+3nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: 2.80E+3nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: 2.80E+3nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: 7.60E+3nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: 7.90E+3nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:Peroxisome proliferator activated receptor gamma agonistic activity in HepG2 cells.More data for this Ligand-Target Pair