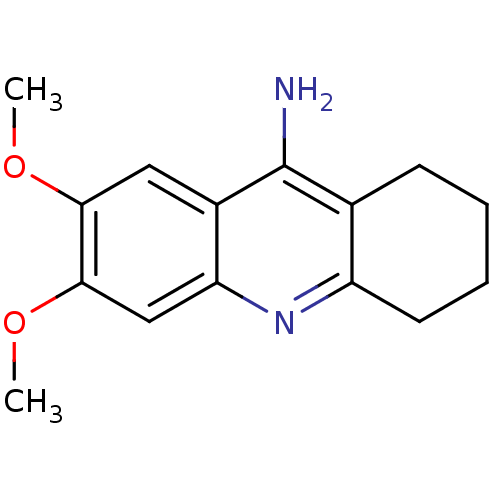
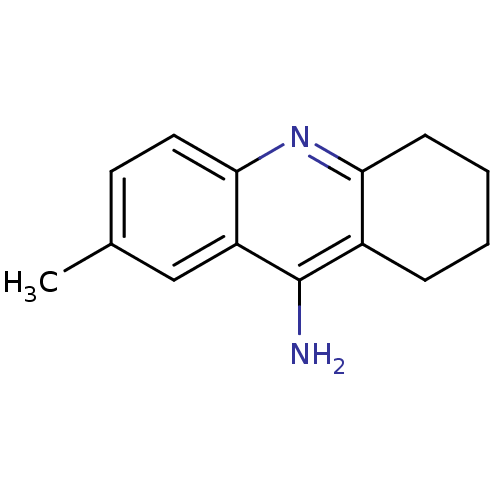
Report error Found 23 Enz. Inhib. hit(s) with all data for entry = 1116
Affinity DataIC50: 9.90nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 28nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 45nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 87nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 250nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 290nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 350nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 390nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 460nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 470nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 550nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 750nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 3.70E+3nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 3.80E+3nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 4.80E+3nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 5.20E+3nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
Affinity DataIC50: 8.10E+3nMpH: 8.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. The appearance of product was monitored at 412 nm for 5 min us...More data for this Ligand-Target Pair
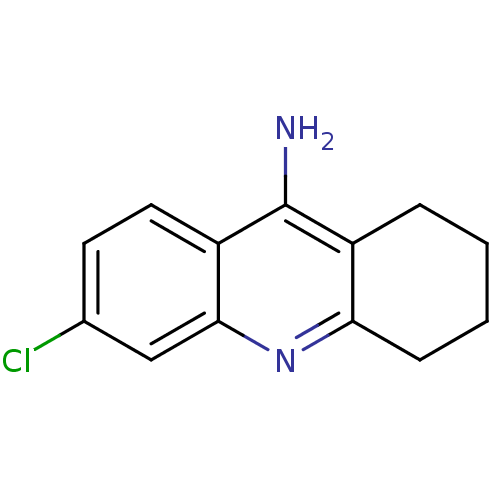
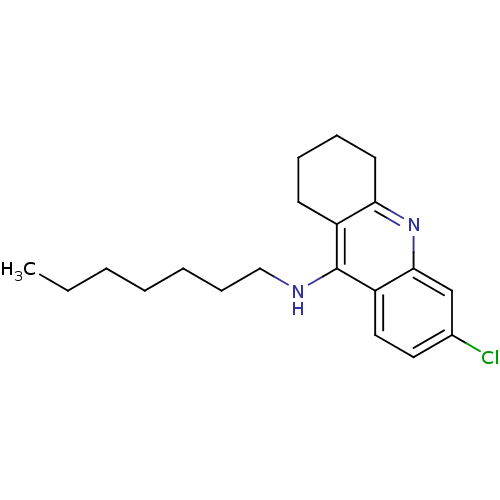
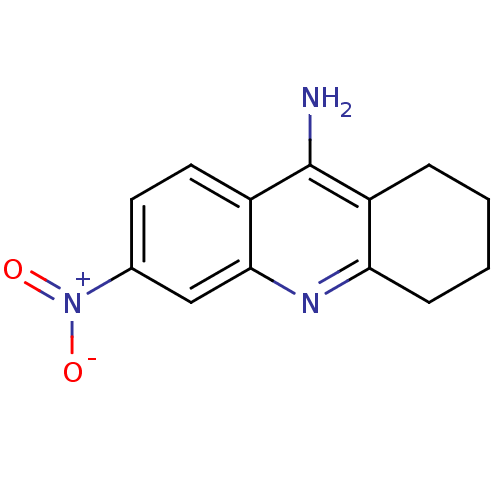
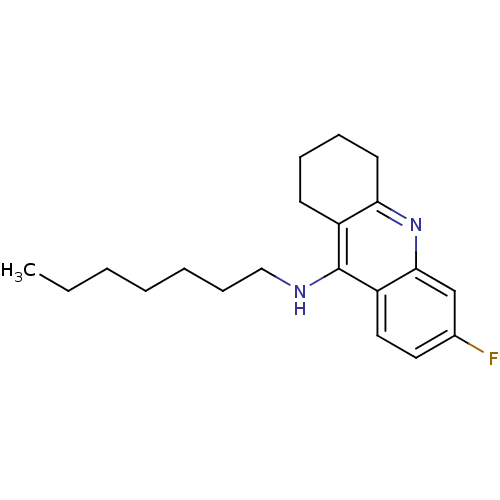
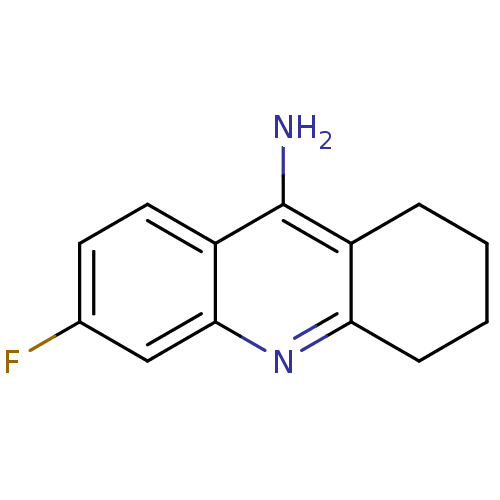
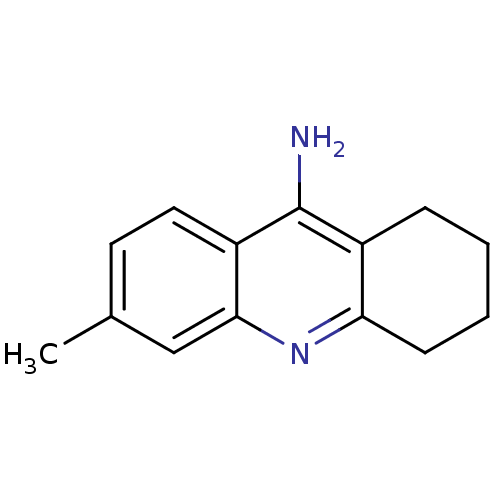
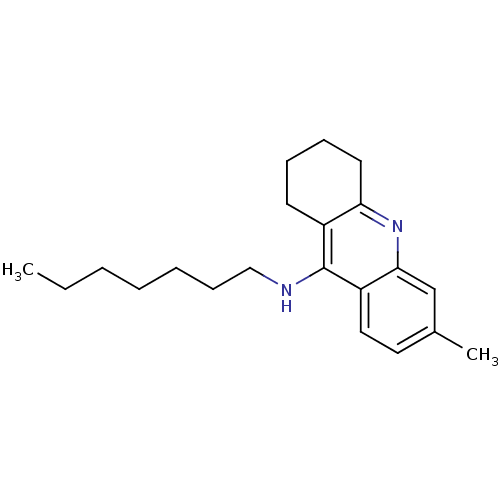
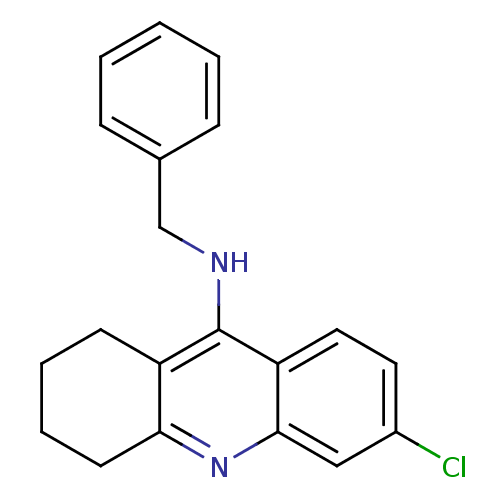
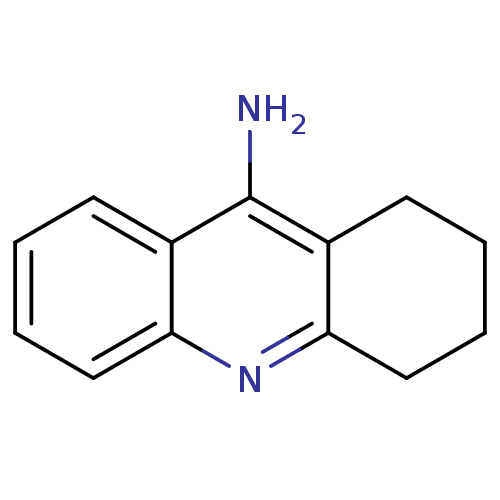
 3D Structure (crystal)
3D Structure (crystal)