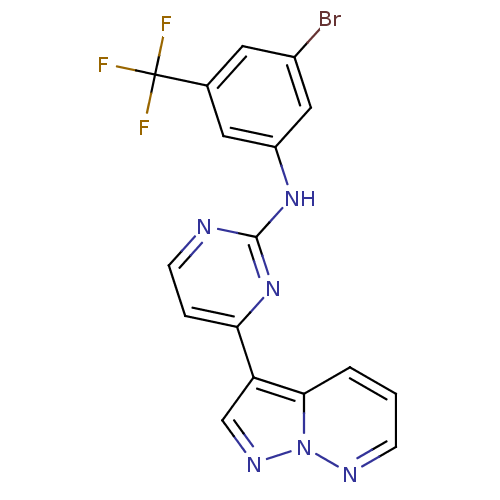
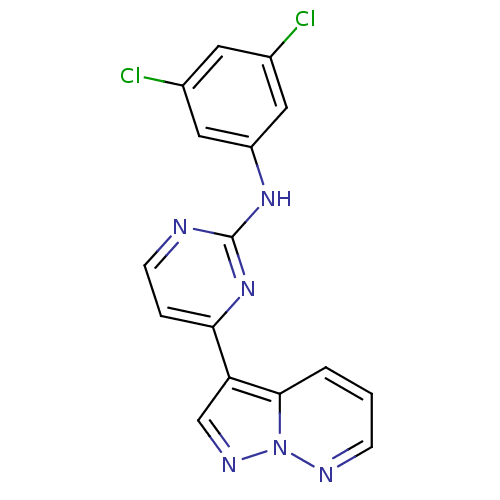
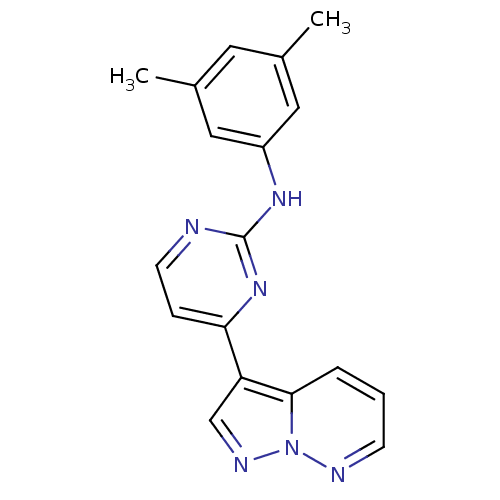
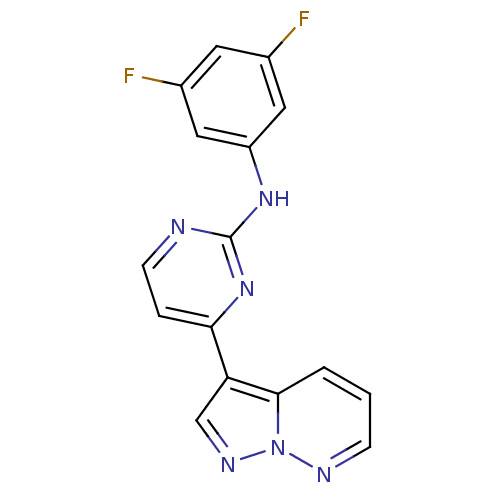
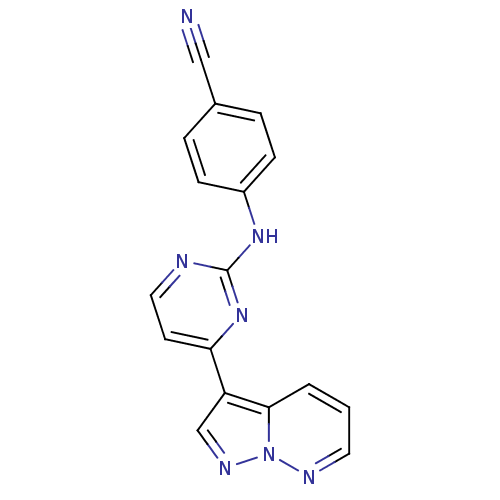
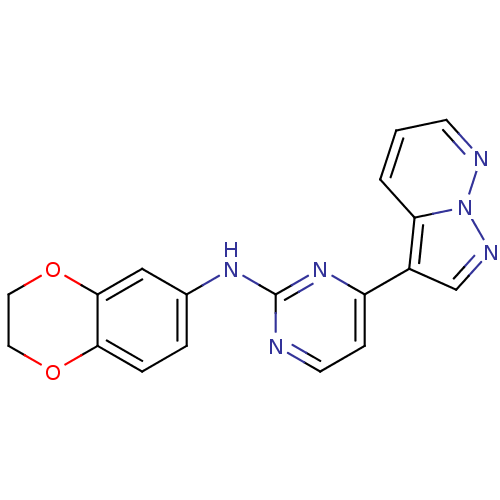
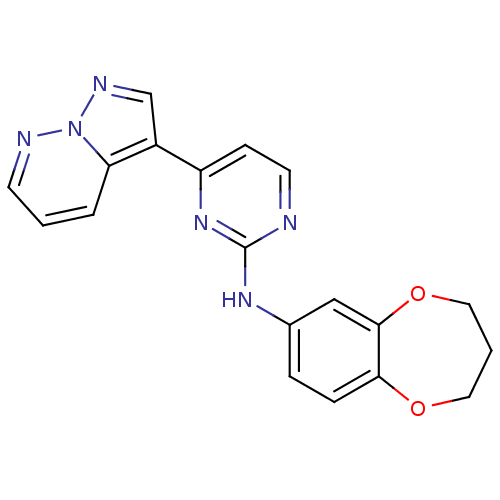
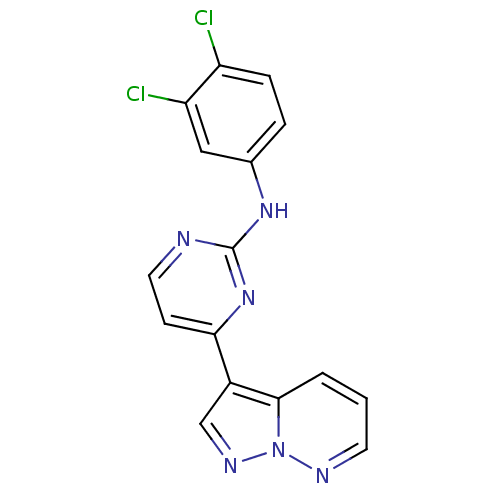
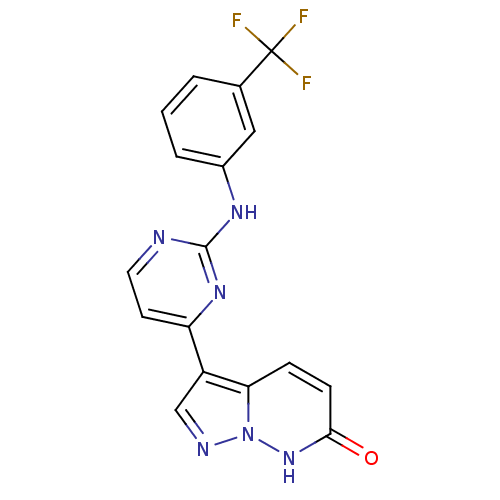
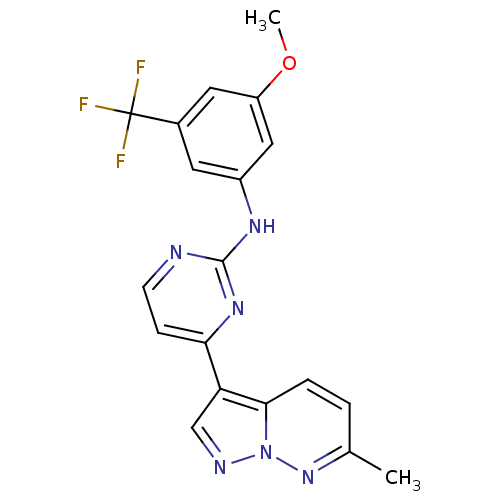
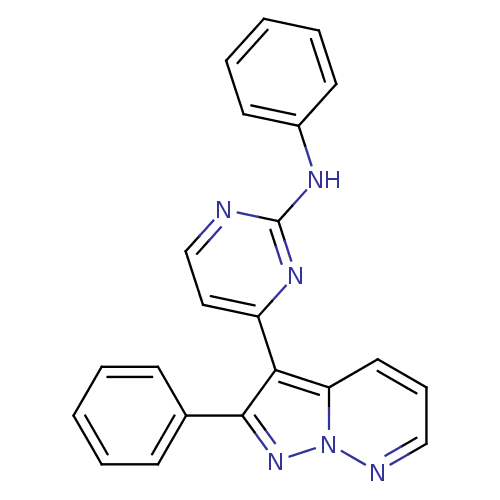
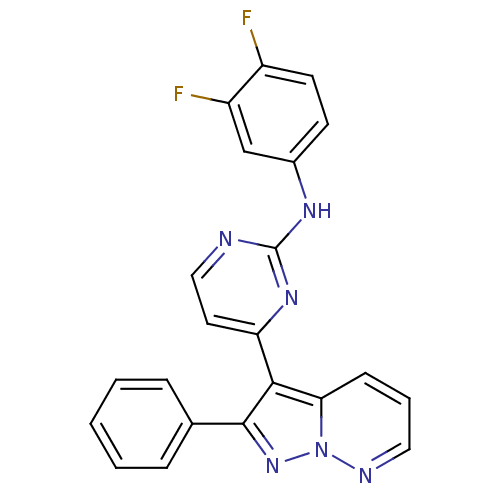
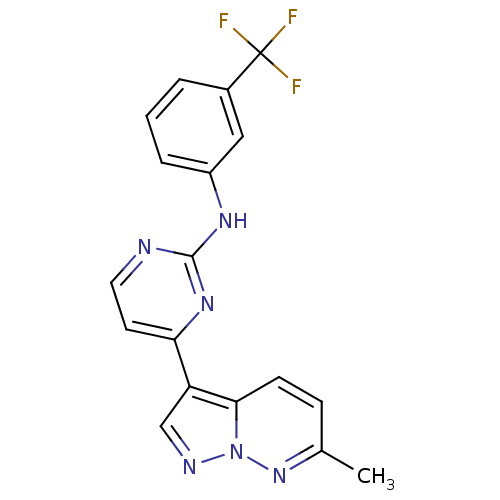
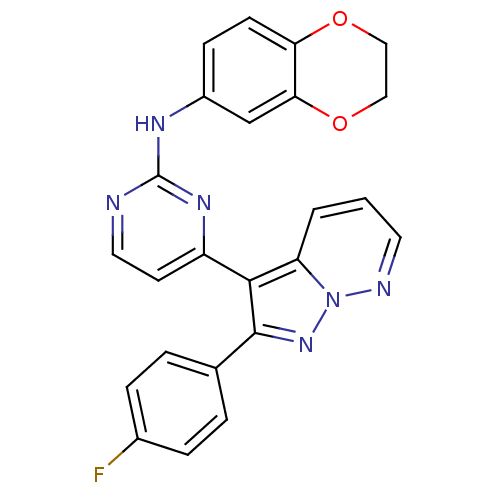
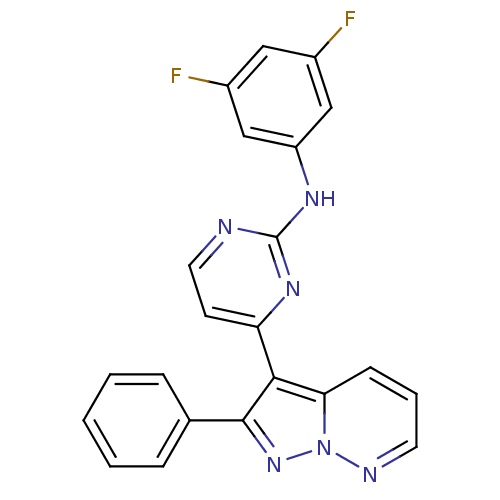
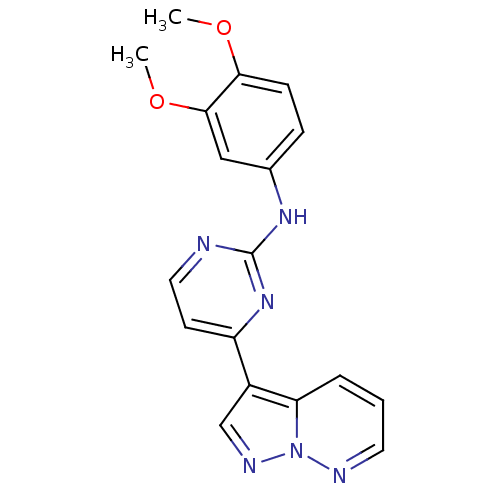
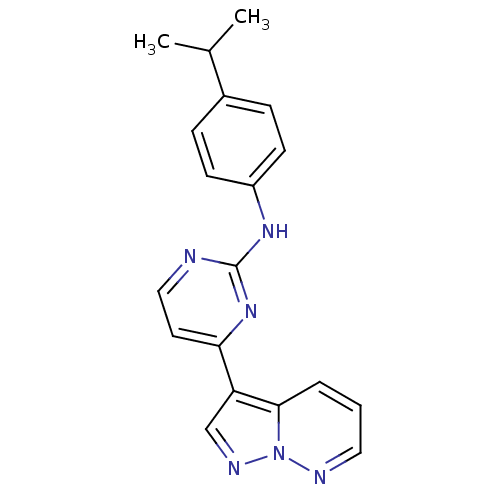
Report error Found 213 Enz. Inhib. hit(s) with all data for entry = 1025
Affinity DataKi: 0.300nM IC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataKi: 1nM IC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataKi: 1nM IC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMpH: 7.2 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataKi: 1nM IC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataKi: 1nM IC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMpH: 7.2 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataKi: 2nM IC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataKi: 2nM IC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMpH: 7.2 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataKi: 2nM IC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataKi: 2nM IC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataKi: 2nM IC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMpH: 7.2 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataKi: 2nM IC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataKi: 2nM IC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMpH: 7.2 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMpH: 7.2 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMpH: 7.2 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMpH: 7.2 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMpH: 7.2 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 19nMpH: 7.2 T: 2°CAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:The biochemical activity of compounds was determined by incubation with specific enzyme and substrate in the presence 2.5 uM ATP/ [gamma-32P] ATP. Af...More data for this Ligand-Target Pair