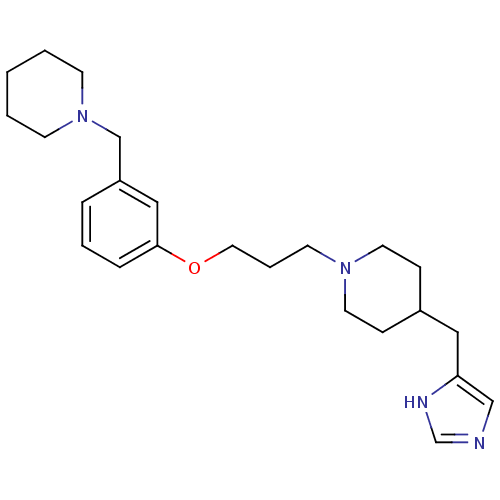
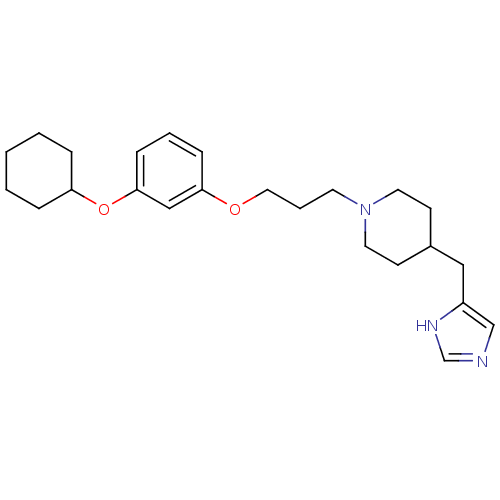
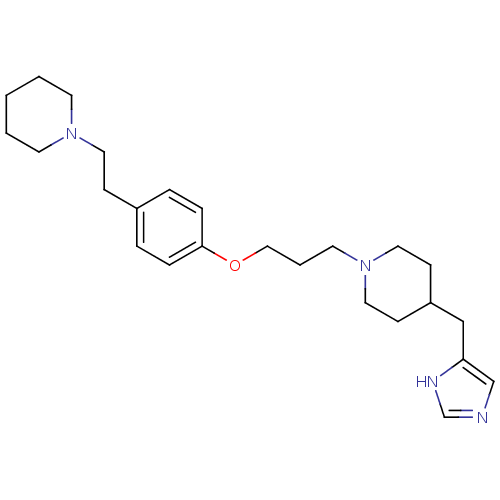
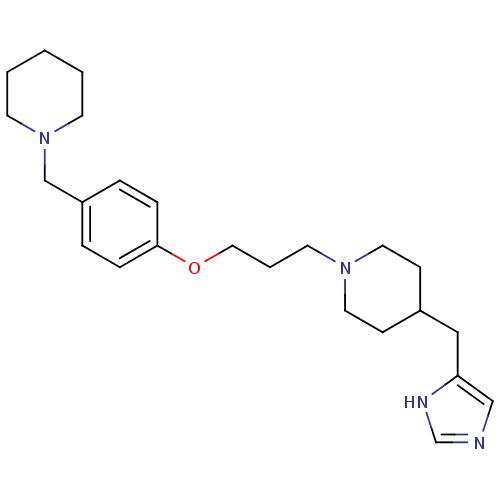
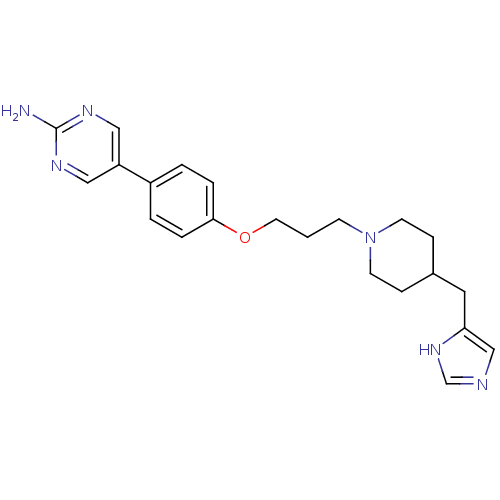
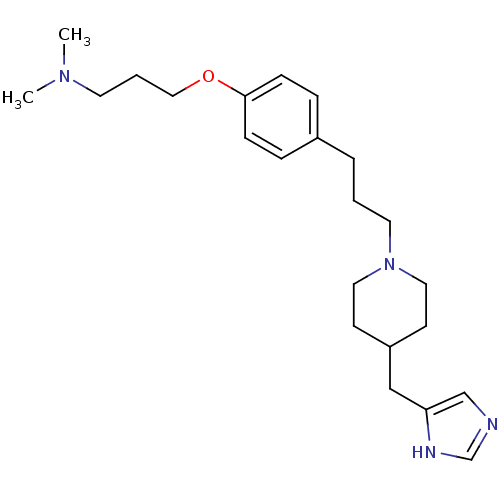
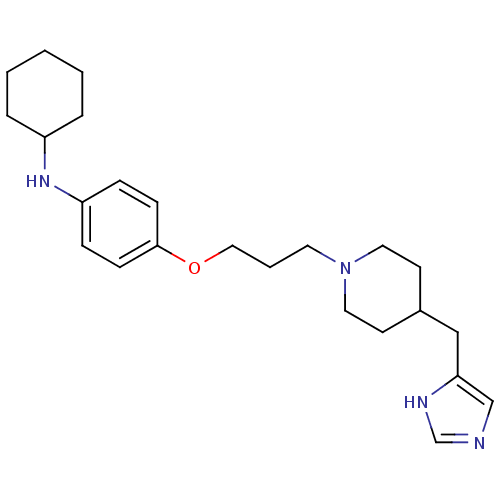
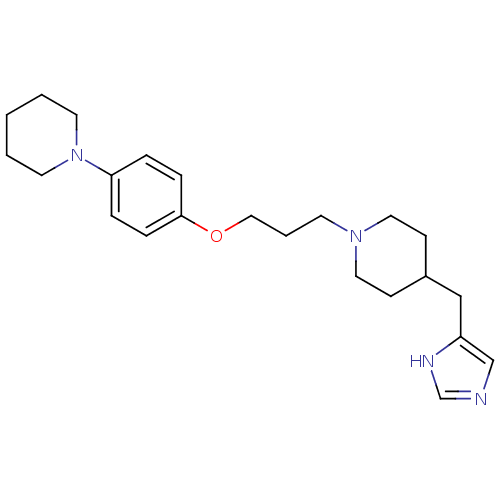
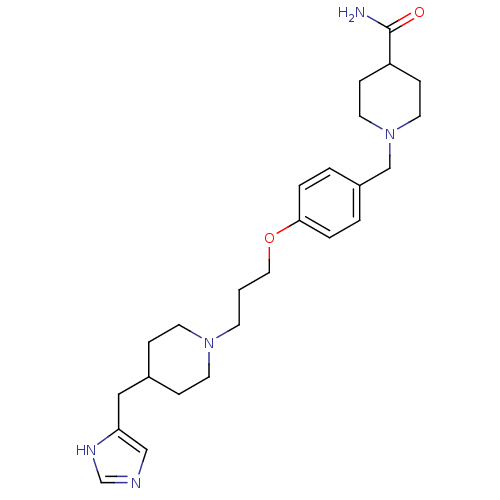
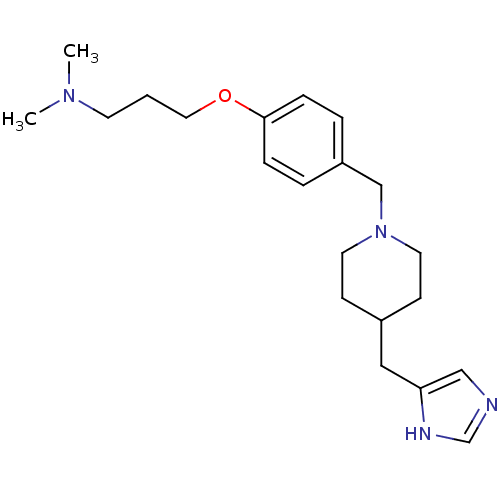
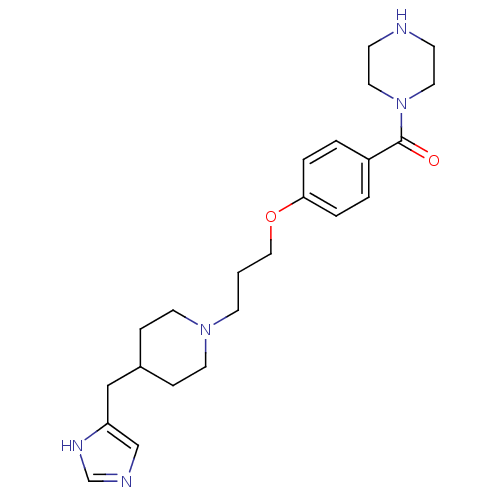
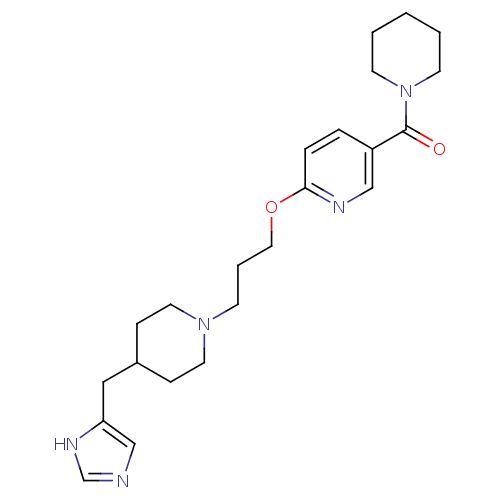
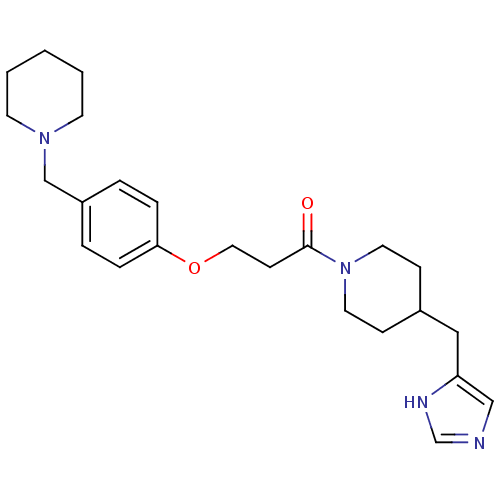
Report error Found 70 Enz. Inhib. hit(s) with all data for entry = 50017106
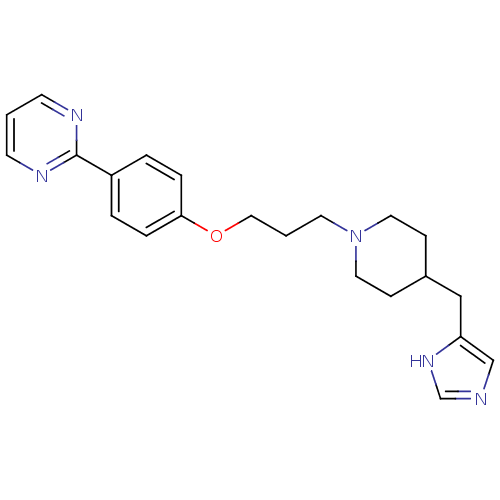
Affinity DataKi: 1nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
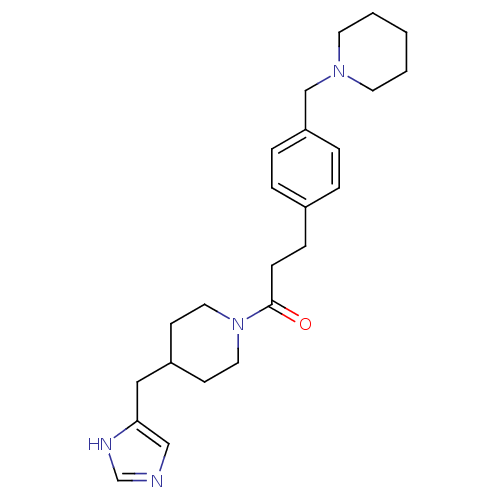
Affinity DataKi: 3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
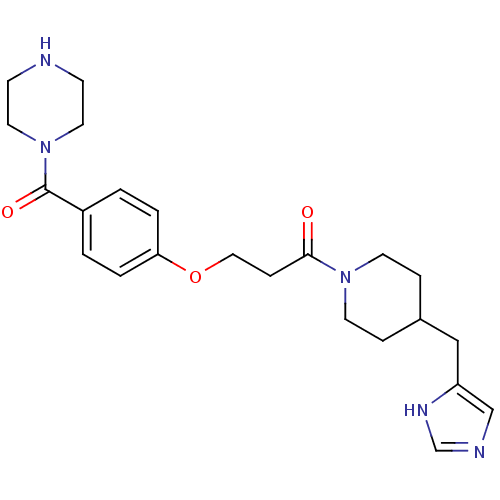
Affinity DataKi: 3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
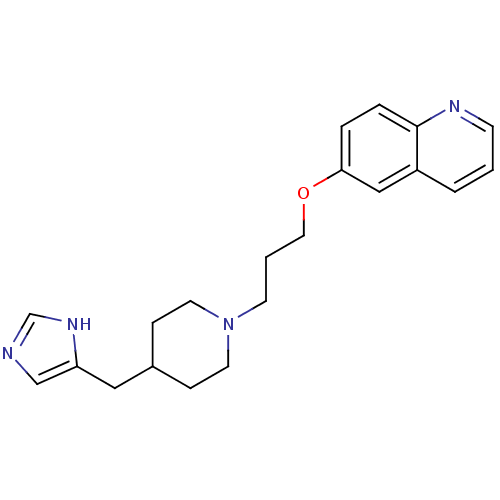
Affinity DataKi: 3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 24nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 26nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of CYP2D6 by human liver microsome assayMore data for this Ligand-Target Pair
Affinity DataKi: 33nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 34nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 40nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 45nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 45nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 51nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 52nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of CYP2D6 by human liver microsome assayMore data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of CYP2D6 by human liver microsome assayMore data for this Ligand-Target Pair
Affinity DataKi: 315nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from guinea pig histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 900nMAssay Description:Inhibition of CYP2D6 by human liver microsome assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of CYP2D6 by supersome assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of CYP2D6 by human liver microsome assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of CYP2D6 by human liver microsome assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of CYP2D6 by human liver microsome assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of CYP2D6 by human liver microsome assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of CYP2D6 by human liver microsome assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of CYP2D6 by human liver microsome assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of CYP2D6 by supersome assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of CYP2D6 by human liver microsome assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of CYP2D6 by human liver microsome assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of CYP2D6 by human liver microsome assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of CYP2D6 by human liver microsome assayMore data for this Ligand-Target Pair