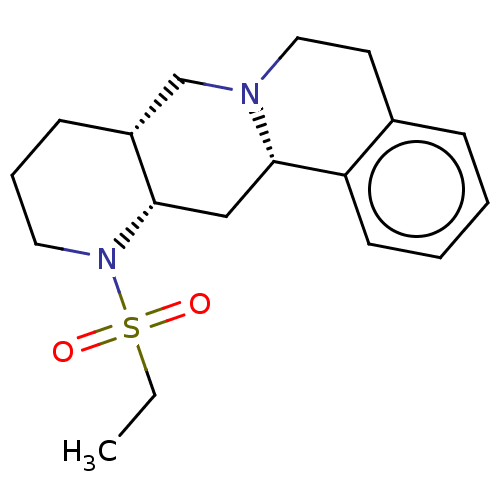
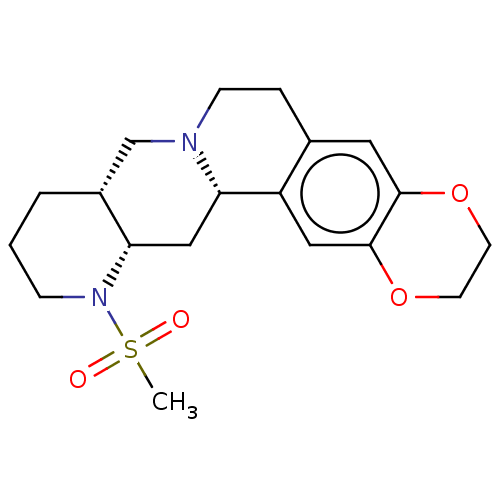
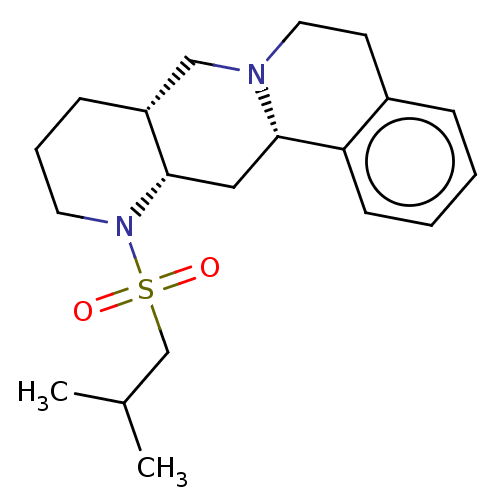
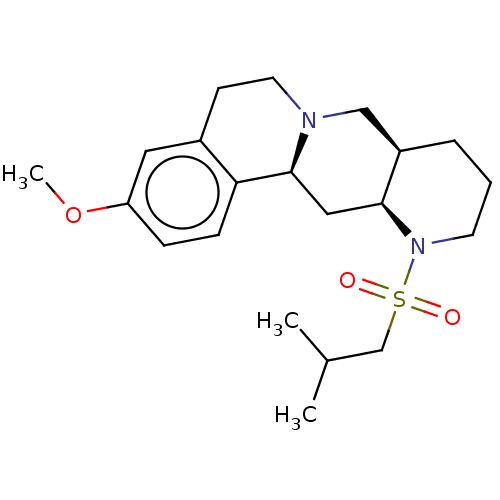
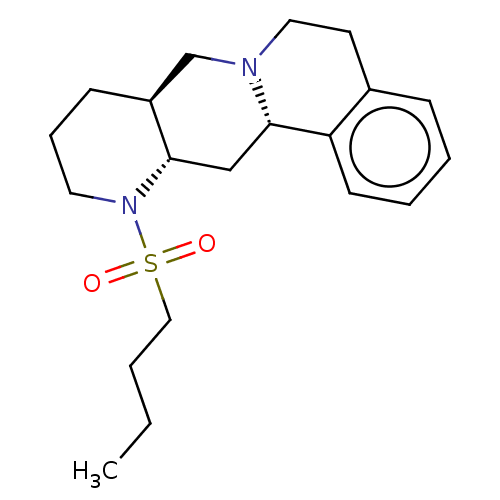
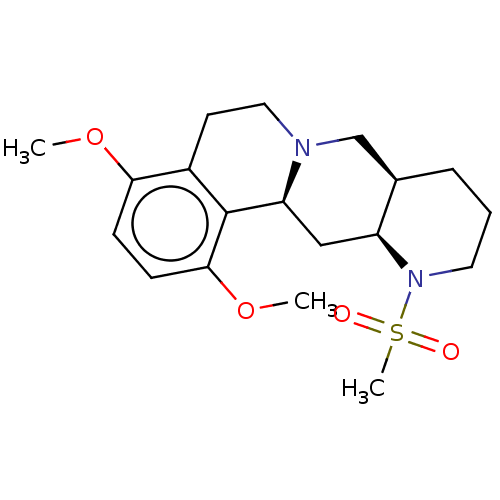
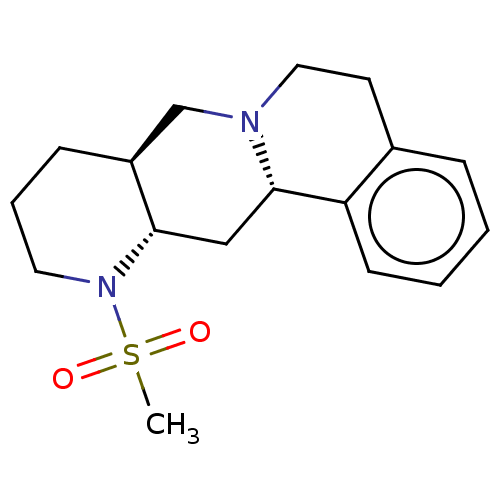
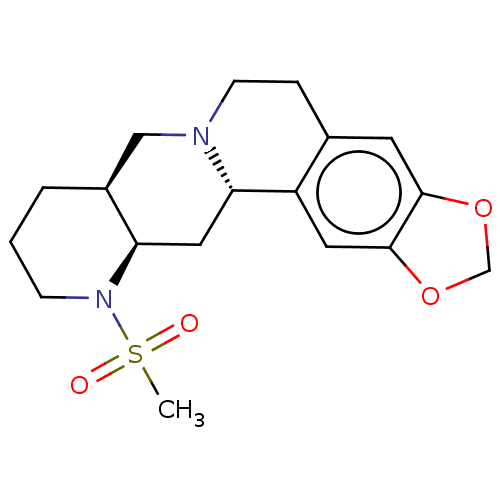
Report error Found 126 Enz. Inhib. hit(s) with all data for entry = 50048856
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
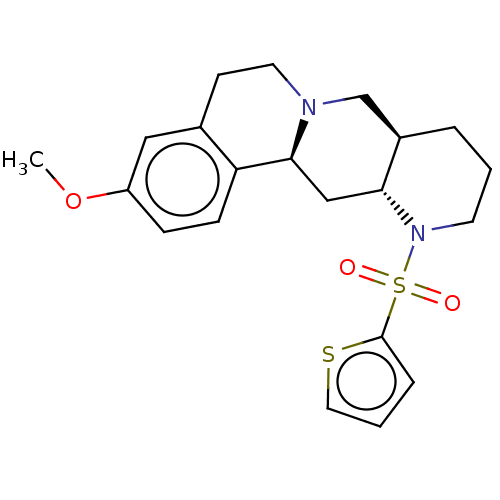
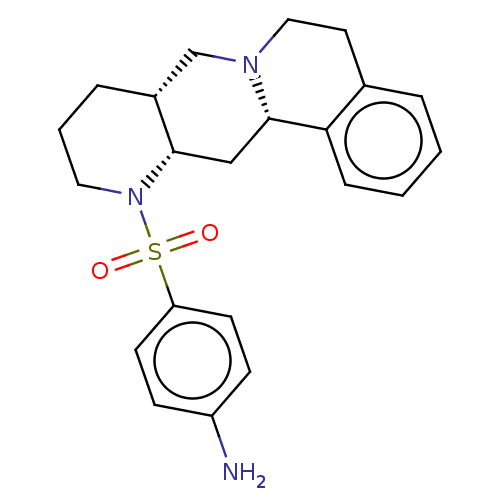
Affinity DataKi: 0.355nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
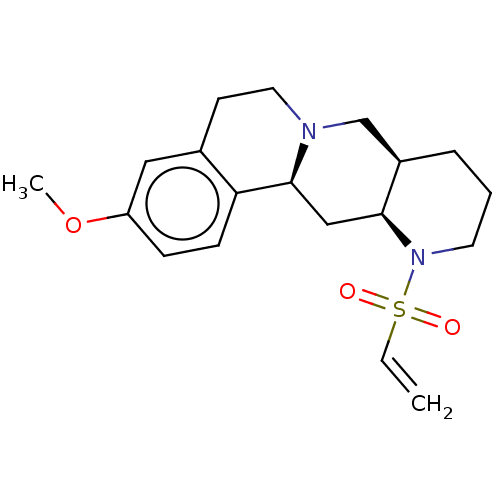
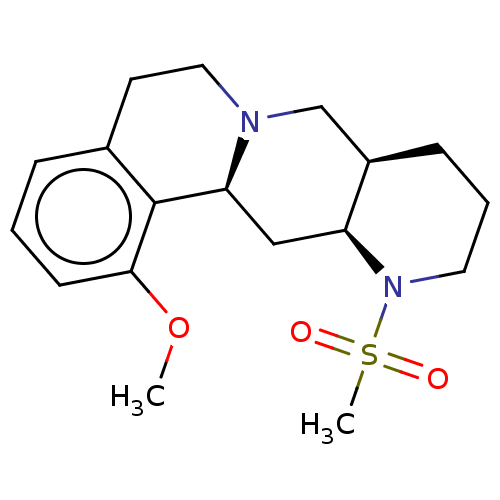
Affinity DataKi: 0.457nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
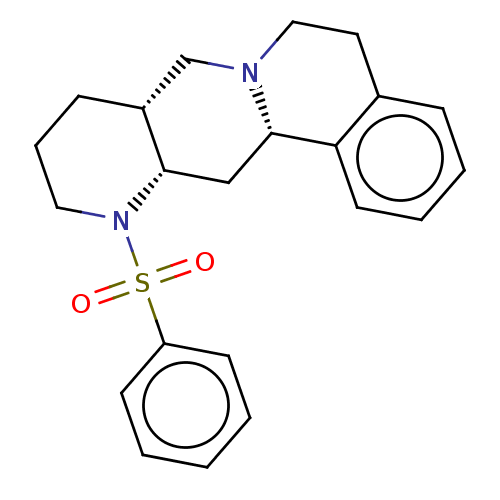
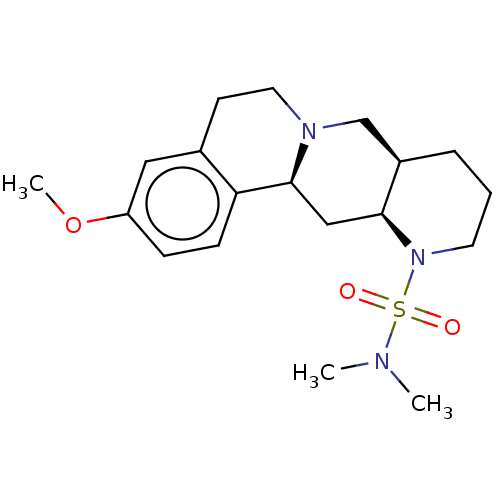
Affinity DataKi: 0.603nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
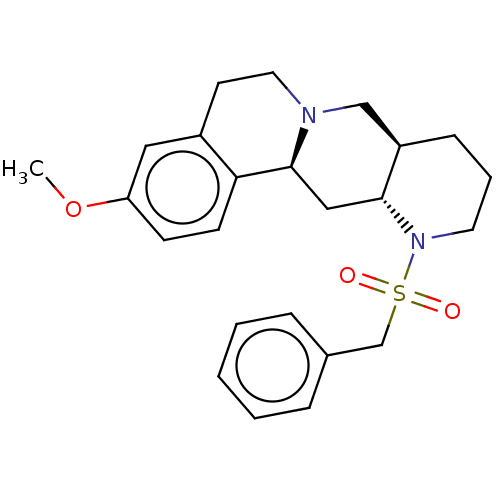
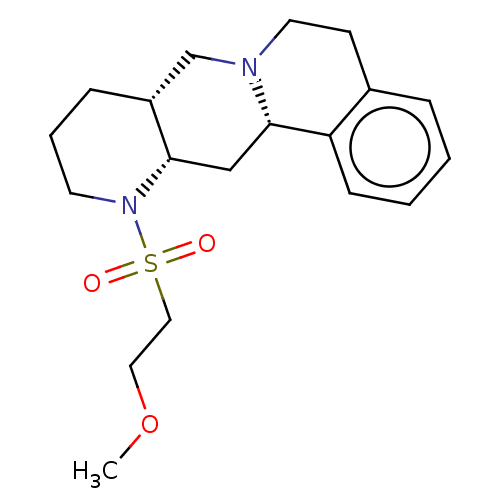
Affinity DataKi: 0.646nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 0.661nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 0.661nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 0.708nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 1.10nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 1.10nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 1.30nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 1.5nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 1.5nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 1.5nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 1.5nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 1.5nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 1.60nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 1.70nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 1.70nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 1.70nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 1.80nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 1.90nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 2nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 2.20nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 2.20nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 2.20nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 2.30nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 3.70nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 5nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 5nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 5nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 6.30nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 6.80nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 6.90nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 7.10nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 7.90nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 9.10nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 9.10nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 9.80nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 11nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 11nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 13nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 14nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 17nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 17nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 28nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 29nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 38nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 43nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 56nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
Syntex Research
Curated by ChEMBL
Syntex Research
Curated by ChEMBL
Affinity DataKi: 58nMAssay Description:Binding affinity to alpha-2 adrenergic receptor determined by measurement of [3H]yohimbine displacement from rat cortical membraneMore data for this Ligand-Target Pair