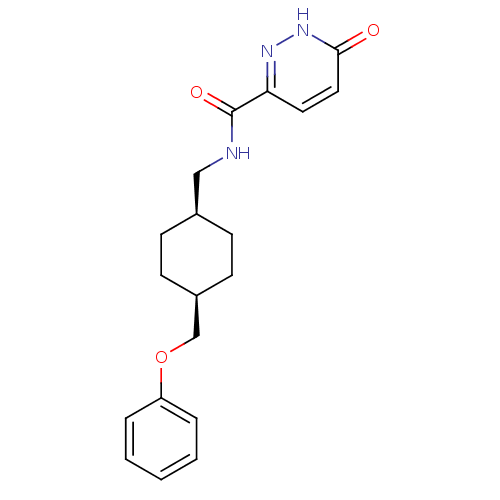
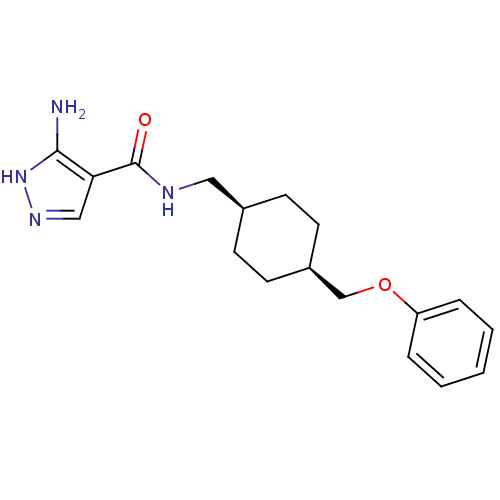
Report error Found 25 Enz. Inhib. hit(s) with all data for entry = 50021070
Affinity DataKi: 3.70nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 4.20nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 5.60nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 5.60nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 5.90nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 6.80nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 6.80nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 8.5nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 8.90nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: 84nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: >100nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: >100nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: >100nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: >100nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: >100nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataKi: >100nMAssay Description:Displacement of [3H]racemic CP-101606 from rat NR2B receptor in P2 membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of hERGMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of hERGMore data for this Ligand-Target Pair