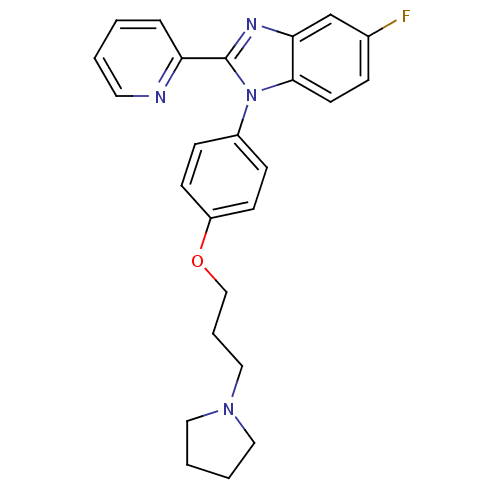
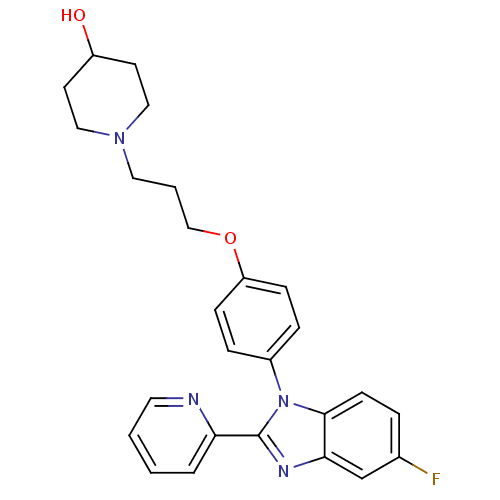
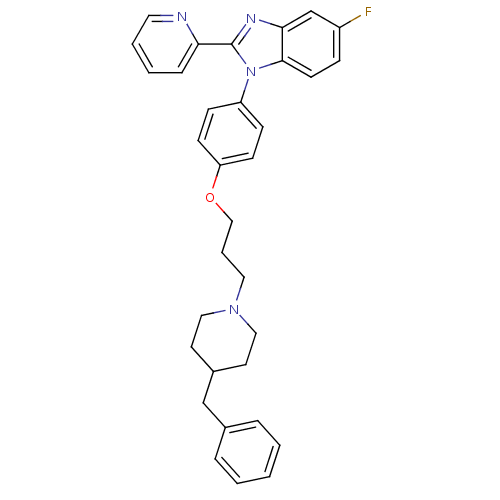
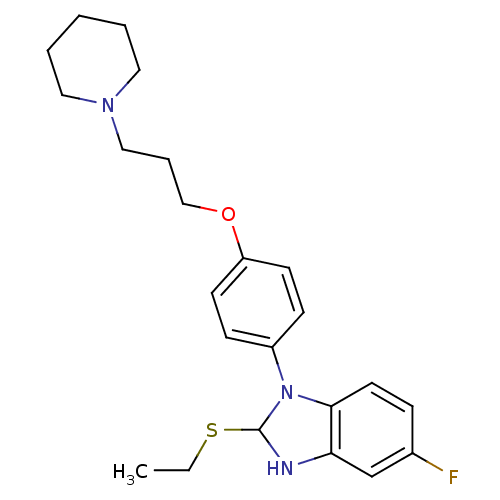
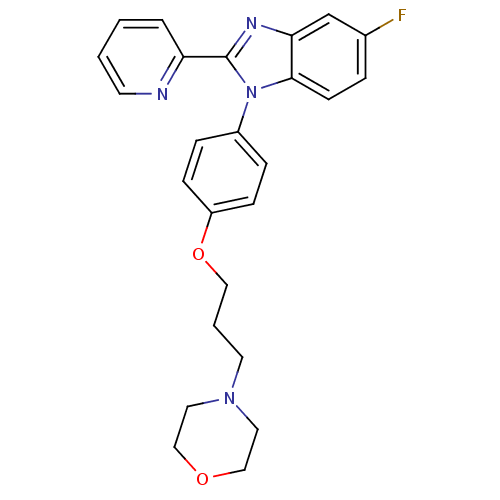
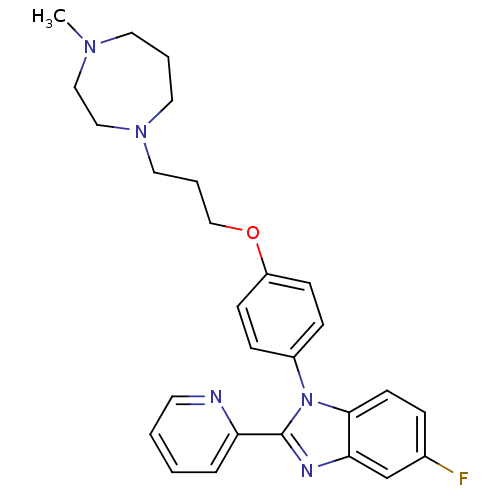
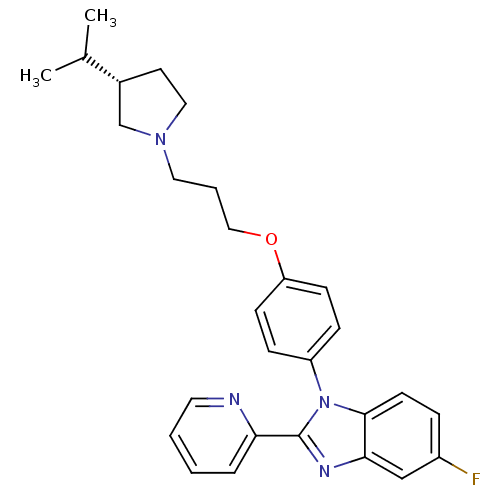
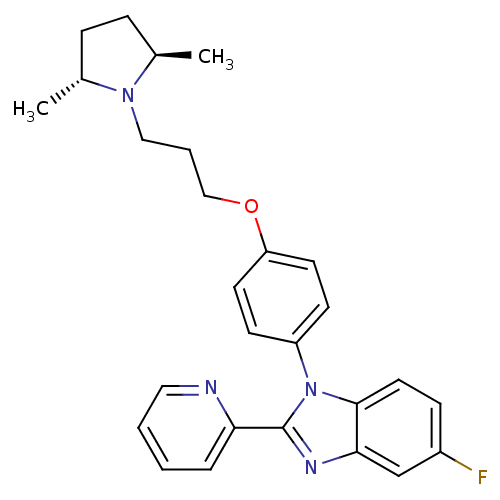
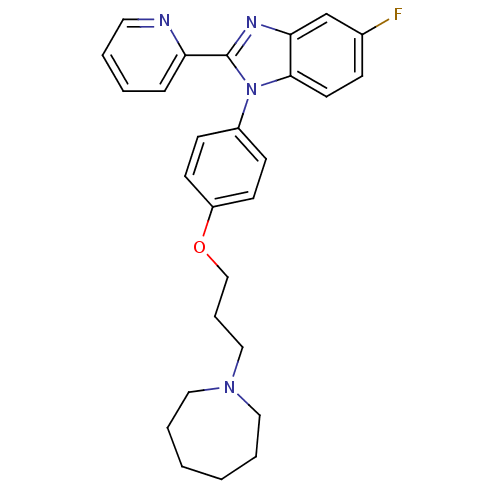
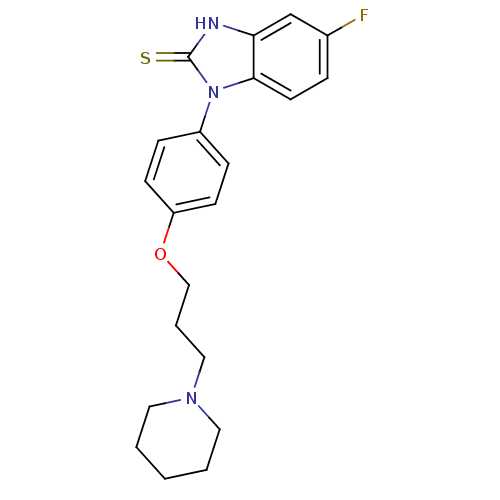
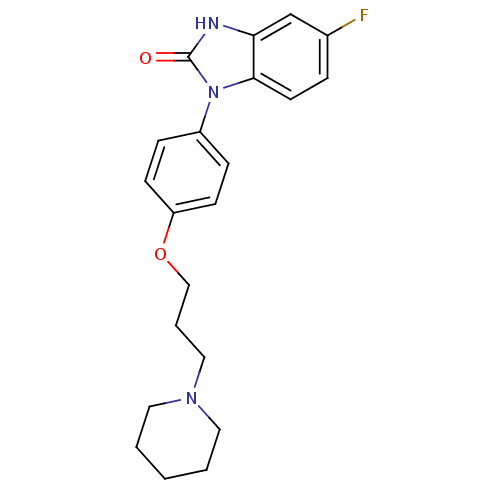
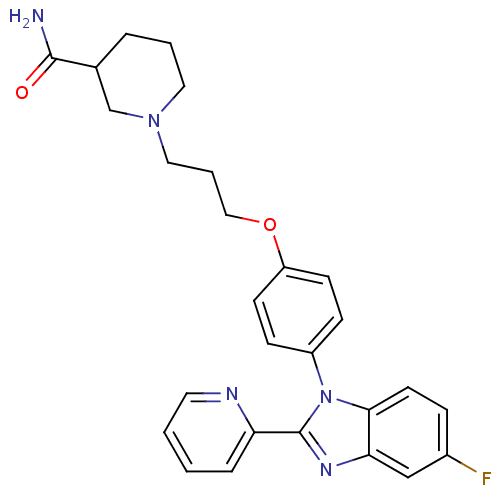
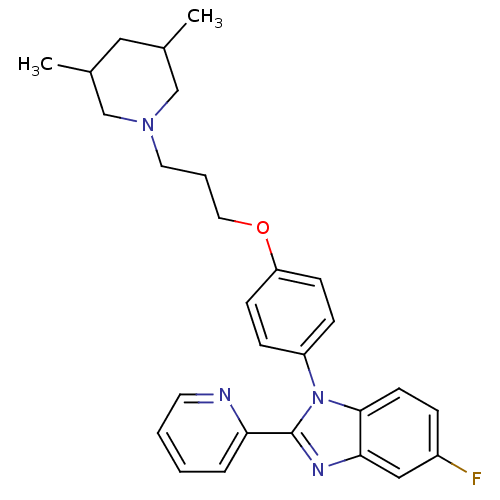
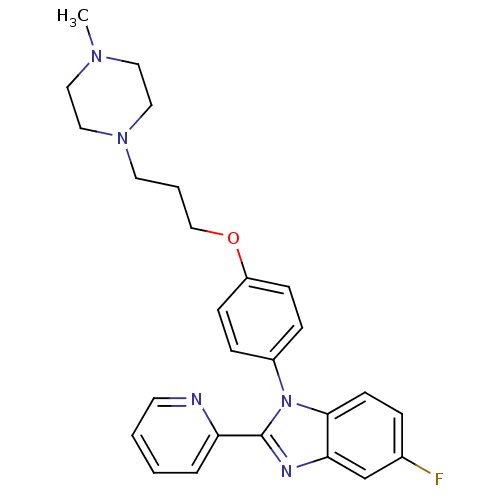
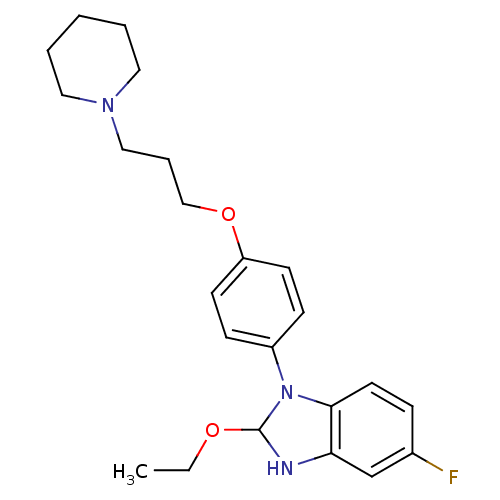
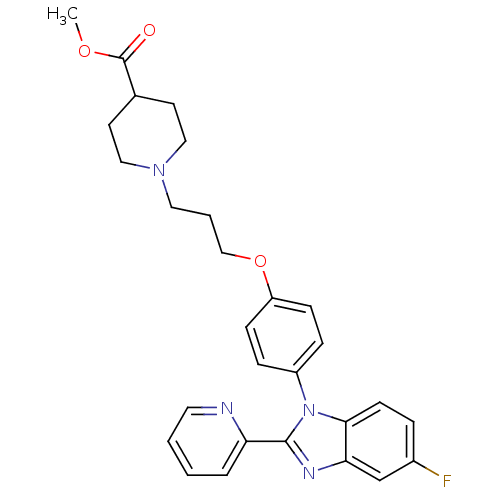
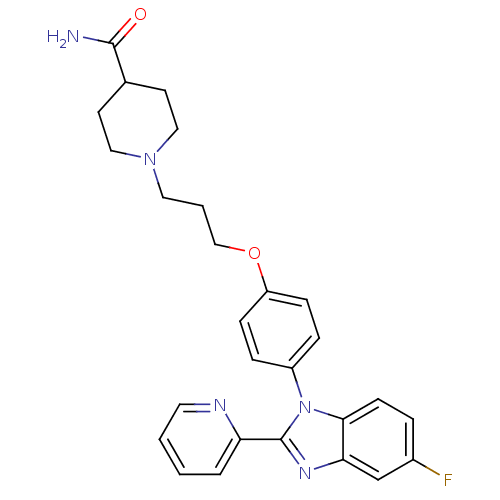
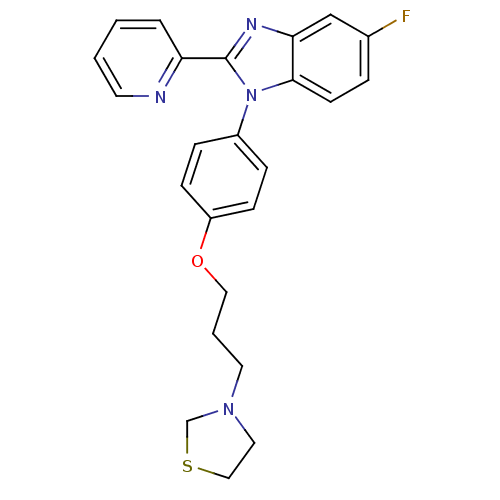
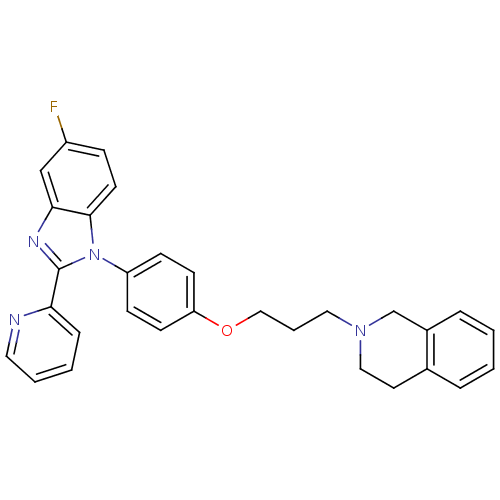
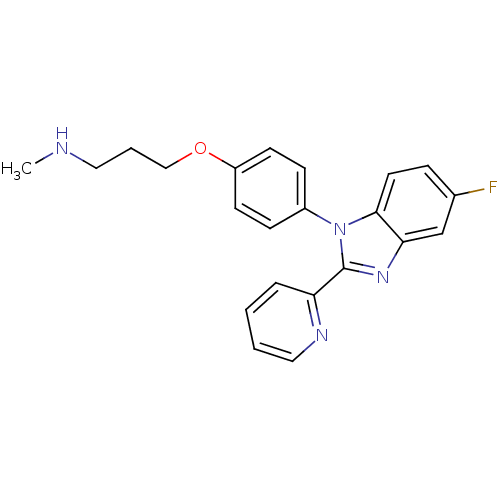
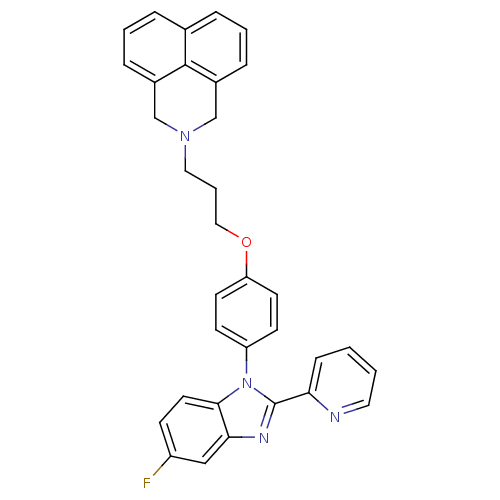
Report error Found 31 Enz. Inhib. hit(s) with all data for entry = 50027016
Affinity DataKi: 0.100nMAssay Description:Antagonist activity at human histamine H3 receptor expressed in HEK293 cells assessed as reversal of N-alpha-methylhistamine-induced inhibition of fo...More data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Binding affinity to mouse histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.60nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 3.40nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 3.60nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 4.80nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 5.10nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 5.30nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 8.70nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
The Schering Plough Research Institute
Curated by ChEMBL
The Schering Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 16nMAssay Description:Binding affinity to human ERG assessed as rubidium efflux at 5 ug/mL pre-equilibrated for 30 mins by DiBAC4(3)-based flame atomic absorbance spectros...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
The Schering Plough Research Institute
Curated by ChEMBL
The Schering Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 26nMAssay Description:Binding affinity to human ERG assessed as rubidium efflux at 5 ug/mL pre-equilibrated for 30 mins by DiBAC4(3)-based flame atomic absorbance spectros...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
The Schering Plough Research Institute
Curated by ChEMBL
The Schering Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 42nMAssay Description:Binding affinity to human ERG assessed as rubidium efflux at 5 ug/mL pre-equilibrated for 30 mins by DiBAC4(3)-based flame atomic absorbance spectros...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
The Schering Plough Research Institute
Curated by ChEMBL
The Schering Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 49nMAssay Description:Binding affinity to human ERG assessed as rubidium efflux at 5 ug/mL pre-equilibrated for 30 mins by DiBAC4(3)-based flame atomic absorbance spectros...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
The Schering Plough Research Institute
Curated by ChEMBL
The Schering Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 62nMAssay Description:Binding affinity to human ERG assessed as rubidium efflux at 5 ug/mL pre-equilibrated for 30 mins by DiBAC4(3)-based flame atomic absorbance spectros...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
The Schering Plough Research Institute
Curated by ChEMBL
The Schering Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 70nMAssay Description:Binding affinity to human ERG assessed as rubidium efflux at 5 ug/mL pre-equilibrated for 30 mins by DiBAC4(3)-based flame atomic absorbance spectros...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
The Schering Plough Research Institute
Curated by ChEMBL
The Schering Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 94nMAssay Description:Binding affinity to human ERG assessed as rubidium efflux at 5 ug/mL pre-equilibrated for 30 mins by DiBAC4(3)-based flame atomic absorbance spectros...More data for this Ligand-Target Pair
Affinity DataKi: 101nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 107nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 713nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human histamine H3 receptor expressed in HEK cells by scintillation countingMore data for this Ligand-Target Pair