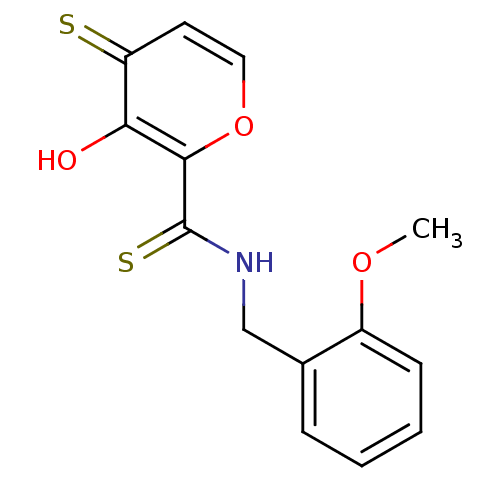
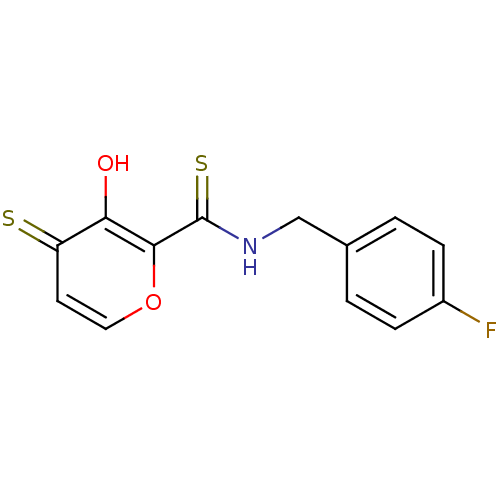
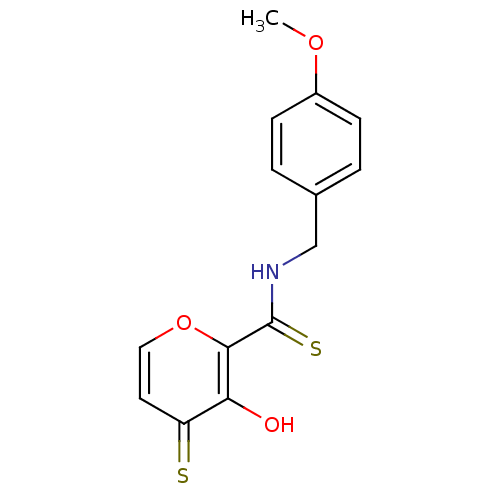
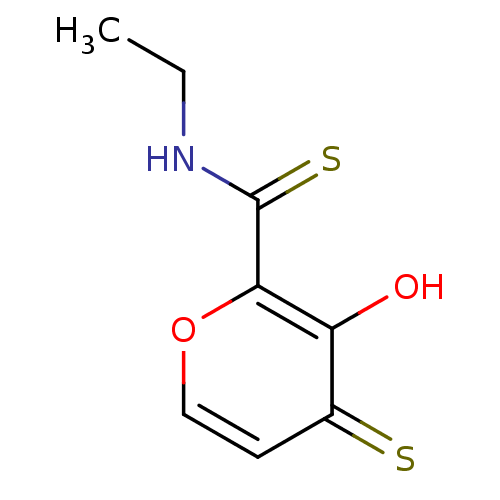
Report error Found 28 Enz. Inhib. hit(s) with all data for entry = 3088
Affinity DataIC50: 5.00E+3nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 6.60E+3nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 7.90E+3nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 8.30E+3nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 8.70E+3nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 9.20E+3nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.06E+4nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.14E+4nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.32E+4nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.41E+4nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 2.92E+4nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 3.35E+4nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 3.74E+4nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 5.04E+4nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 7.25E+4nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.0 T: 2°CAssay Description:Fluorescence peptide cleavage assay was performed in a 96-well plate. Each reaction contained fluorogenic peptide substrate, LF, and small-molecule i...More data for this Ligand-Target Pair