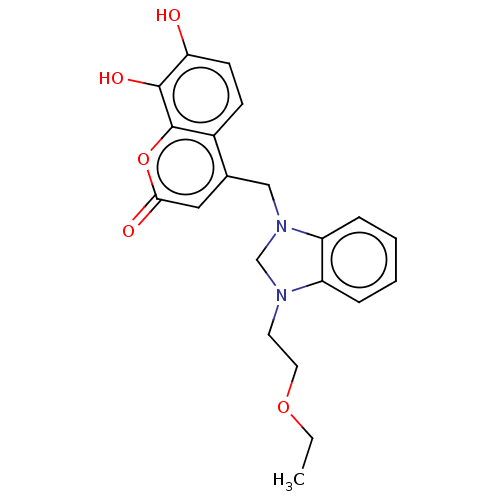
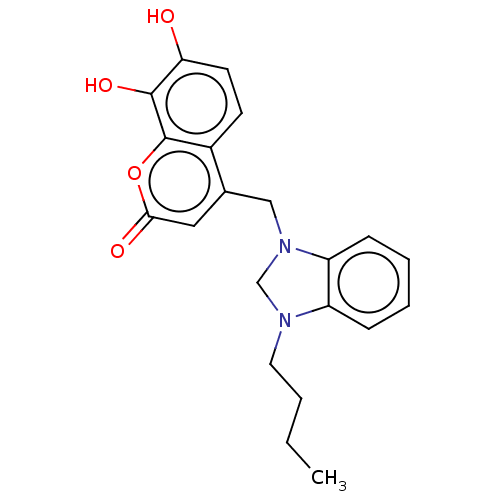
Report error Found 20 Enz. Inhib. hit(s) with all data for entry = 8246
Affinity DataIC50: 2.03E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.21E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.56E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.86E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.89E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.95E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.97E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.12E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.26E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.29E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.31E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.32E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.41E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.14E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.81E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.93E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.94E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.15E+4nMpH: 10.0Assay Description:CA activity was measured by the Maren method which is based on determination of the time required for the pH to decrease from 10.0 to 7.4 due to CO2 ...More data for this Ligand-Target Pair