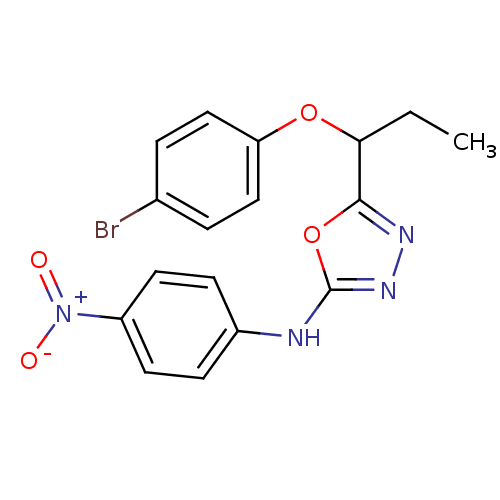
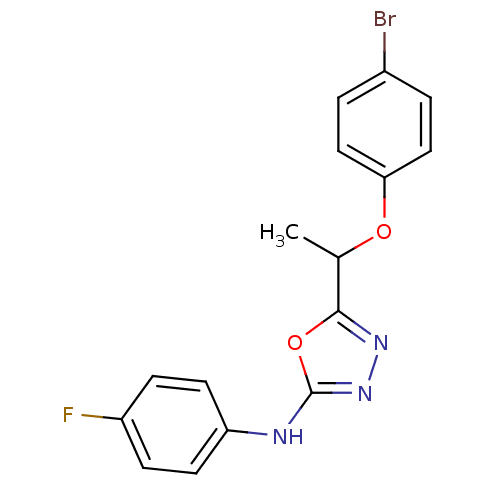
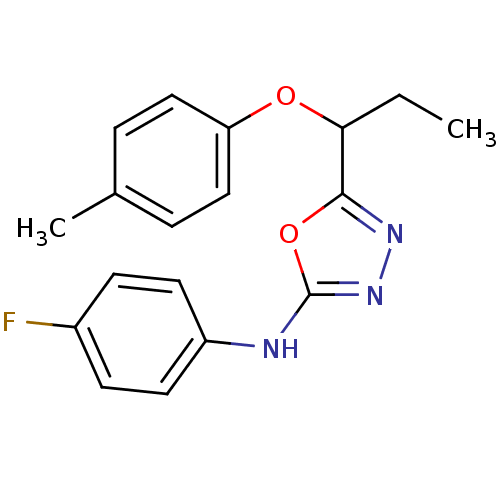
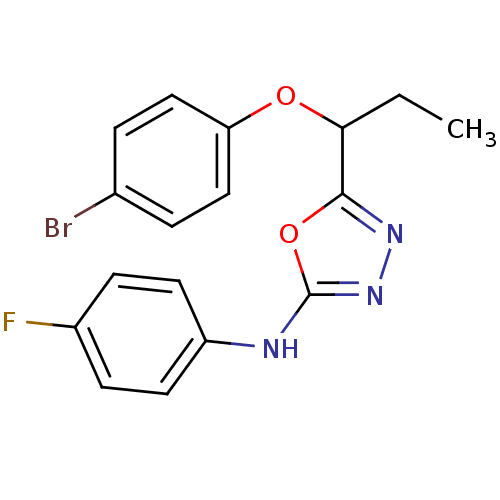
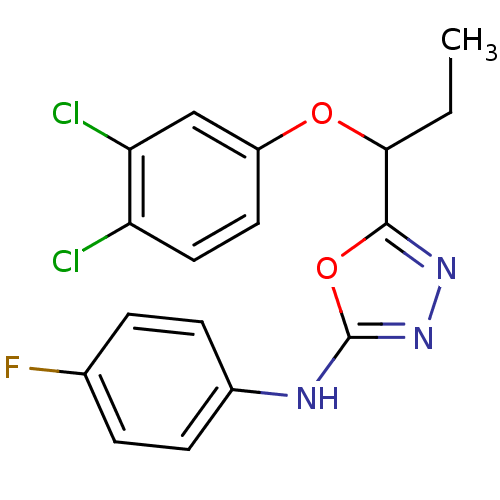
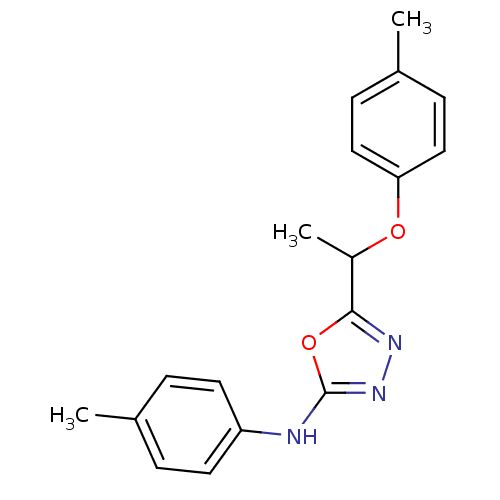
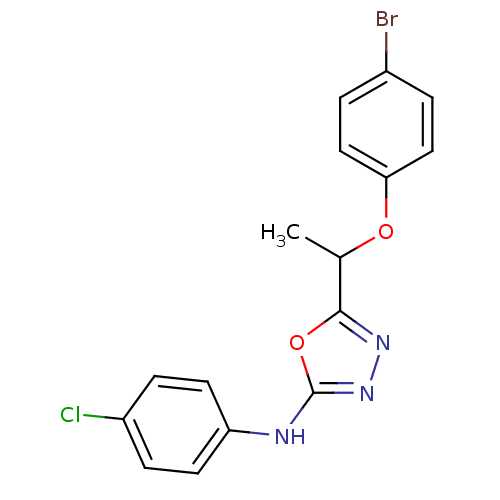
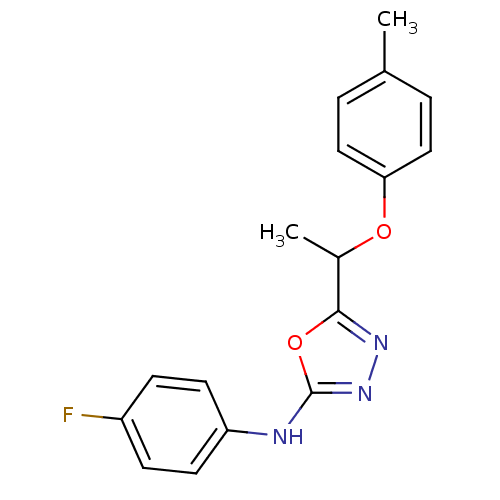
Report error Found 22 Enz. Inhib. hit(s) with all data for entry = 6336
Affinity DataIC50: 6.03E+3nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.21E+3nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.42E+3nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.65E+3nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.02E+3nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.82E+3nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.15E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.16E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.24E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.62E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.32E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.35E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.58E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.28E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.53E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.55E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.59E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.74E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.74E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.72E+4nMT: 2°CAssay Description:The assay mixture contained 40 uL of reaction buffer (100 mM urea, 0.01 M K2HPO4, 1 mM EDTA, and 0.01 M LiCl2, pH 8.2), 10 uL of urease (5 U/mL) and ...More data for this Ligand-Target Pair