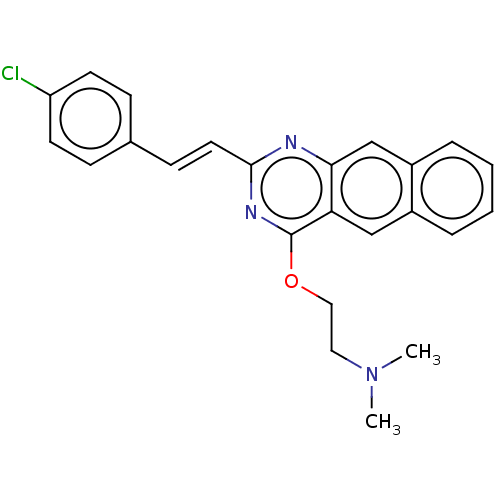
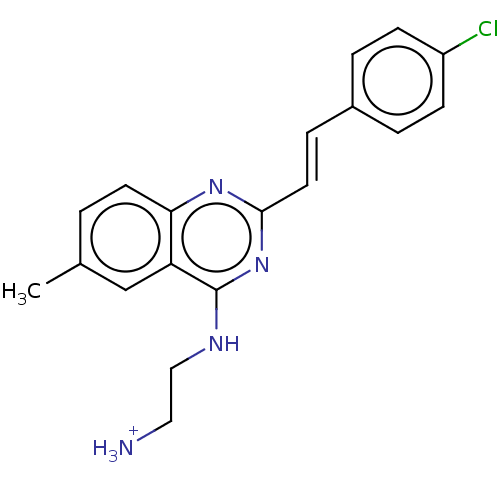
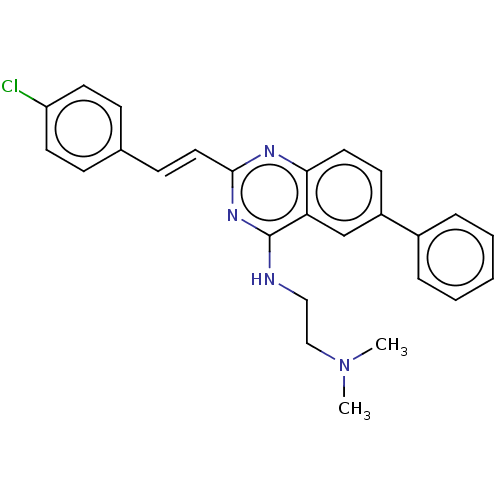
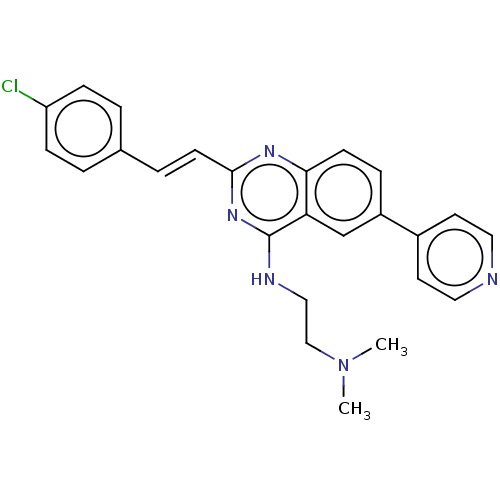
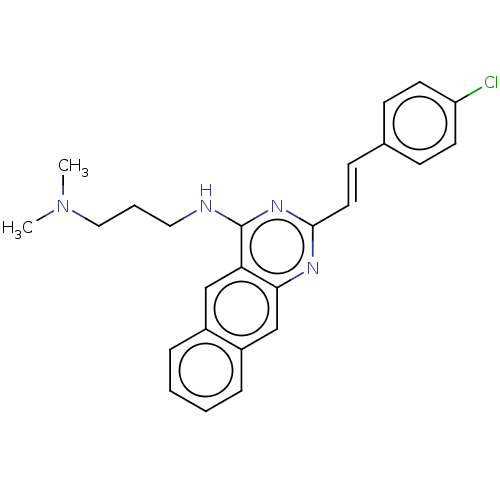
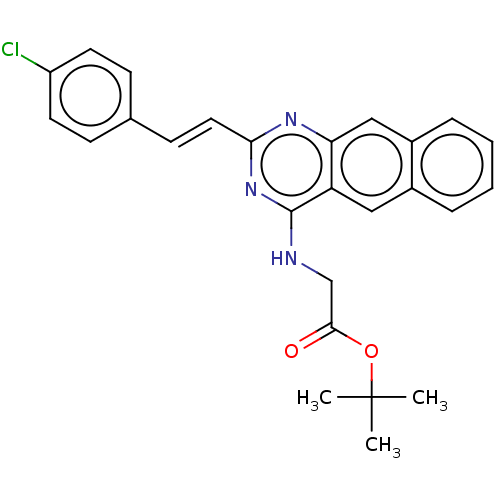
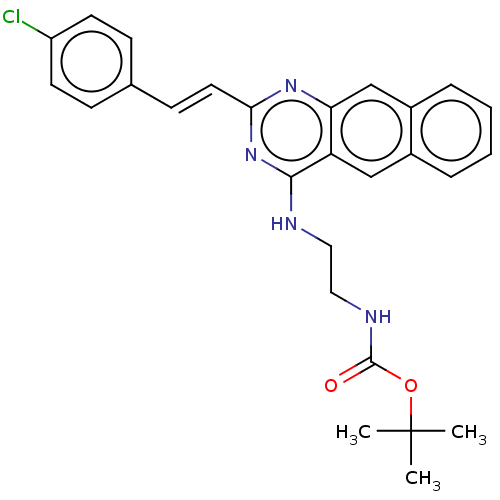
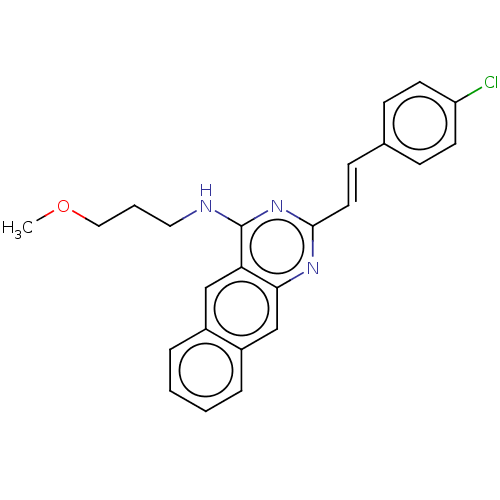
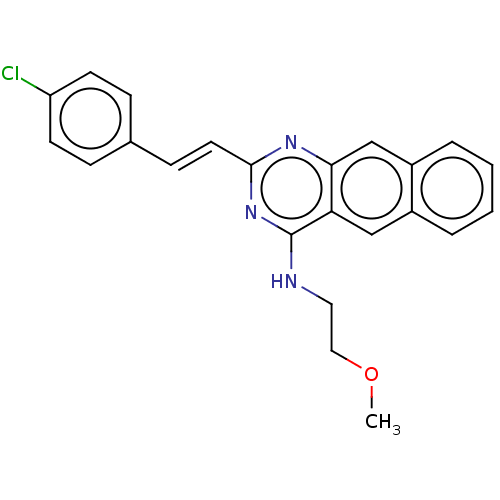
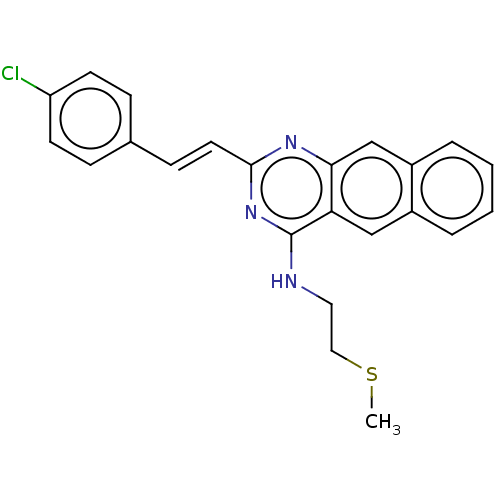
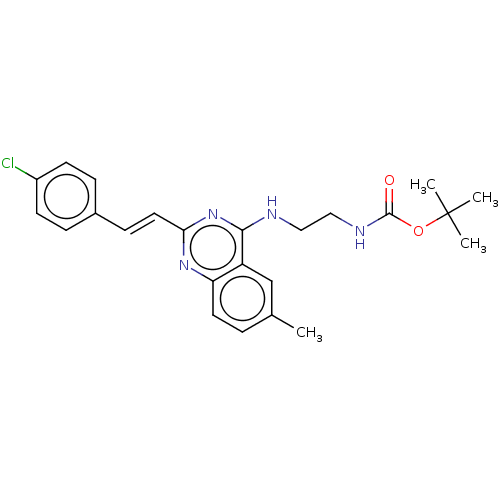
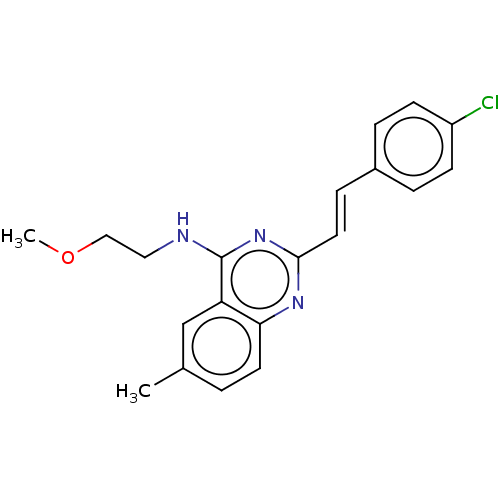
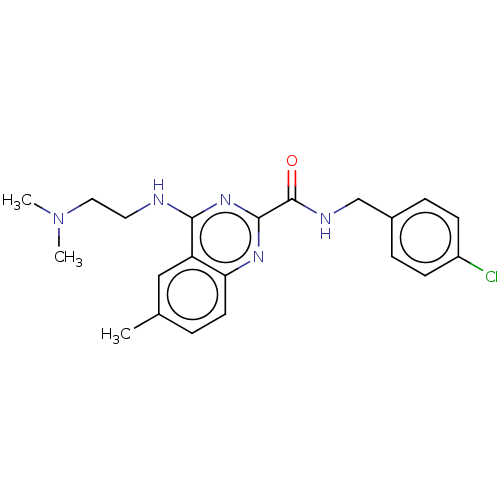
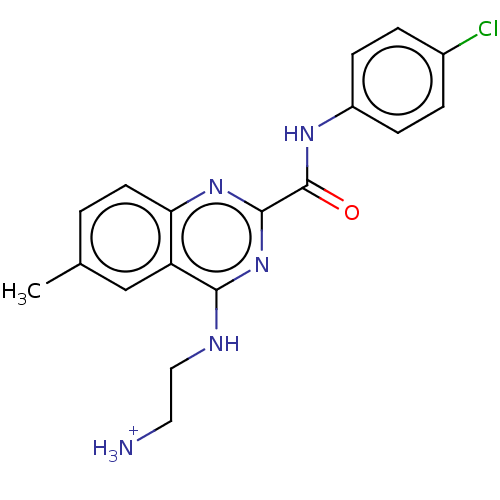
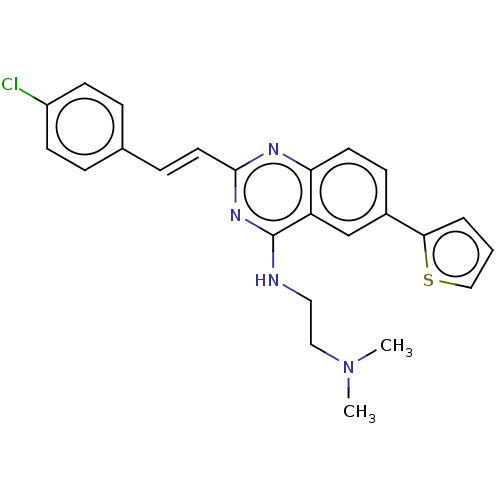
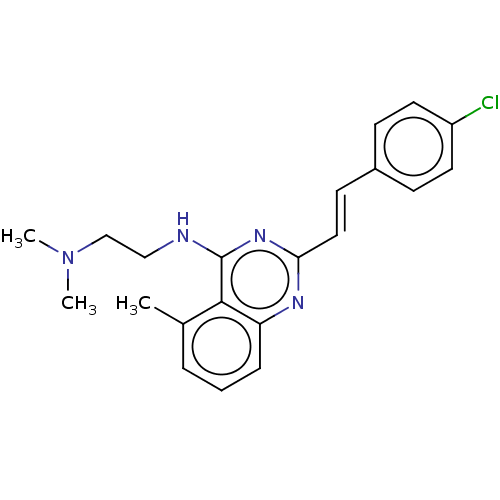
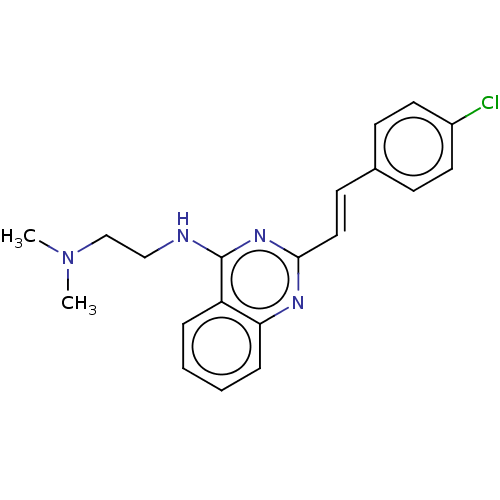
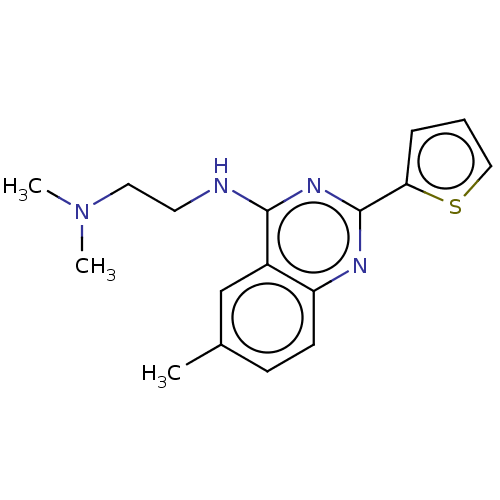
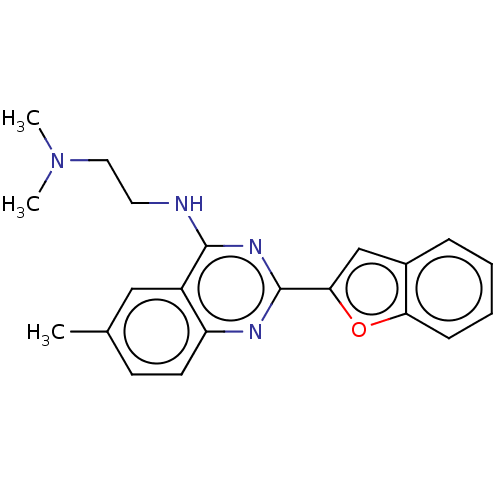
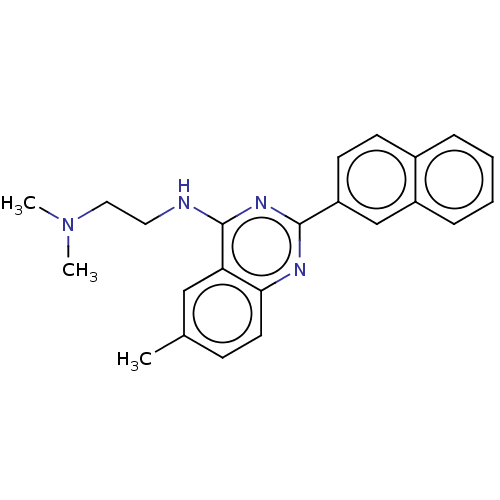
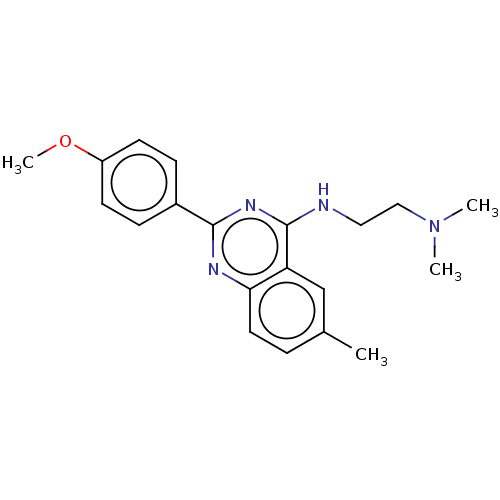
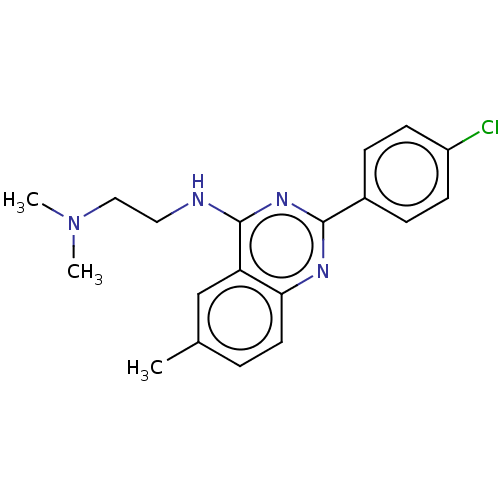
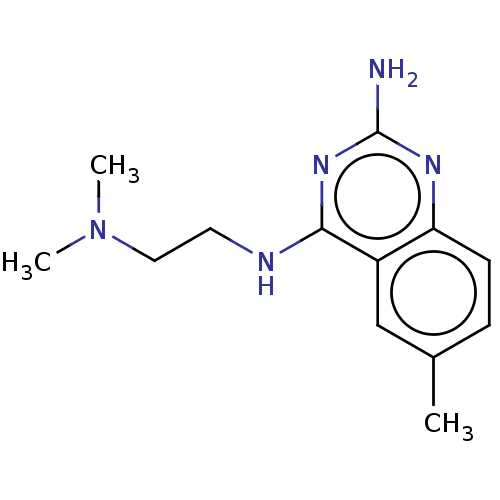
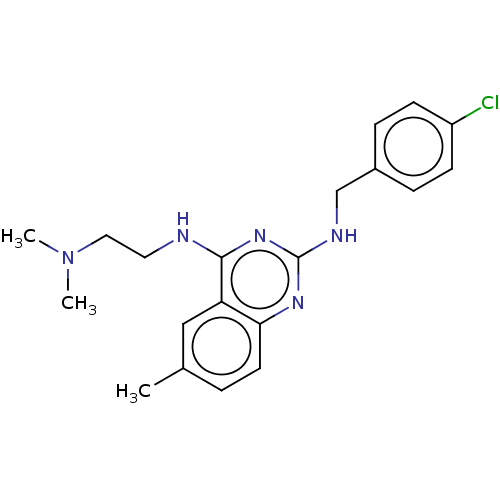
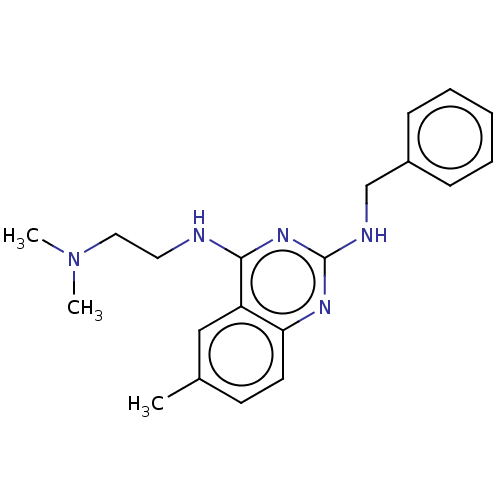
Report error Found 43 Enz. Inhib. hit(s) with all data for entry = 50045852
TargetCell division protein FtsZ(Staphylococcus aureus)
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataEC50: 4.00E+3nMAssay Description:Activation of Staphylococcus aureus FtsZ GTPase activityMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 9.00E+3nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 2.40E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 2.40E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 2.60E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 2.70E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 2.80E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 4.90E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 5.20E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 5.80E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 6.50E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 8.50E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 9.50E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Bacillus subtilis (strain 168))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Bacillus subtilis FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Bacillus subtilis (strain 168))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Bacillus subtilis FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Bacillus subtilis (strain 168))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Bacillus subtilis FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Bacillus subtilis (strain 168))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Bacillus subtilis FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Bacillus subtilis (strain 168))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Bacillus subtilis FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Bacillus subtilis (strain 168))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Bacillus subtilis FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Bacillus subtilis (strain 168))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Bacillus subtilis FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Bacillus subtilis (strain 168))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Bacillus subtilis FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Bacillus subtilis (strain 168))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Bacillus subtilis FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Bacillus subtilis (strain 168))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Bacillus subtilis FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Bacillus subtilis (strain 168))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Bacillus subtilis FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Bacillus subtilis (strain 168))
University of California Davis
Curated by ChEMBL
University of California Davis
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Bacillus subtilis FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
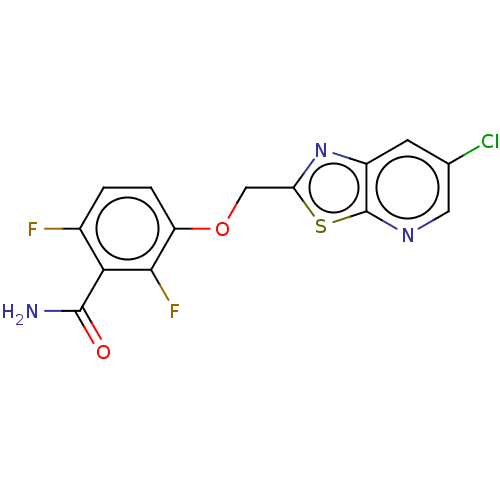
 3D Structure (crystal)
3D Structure (crystal)