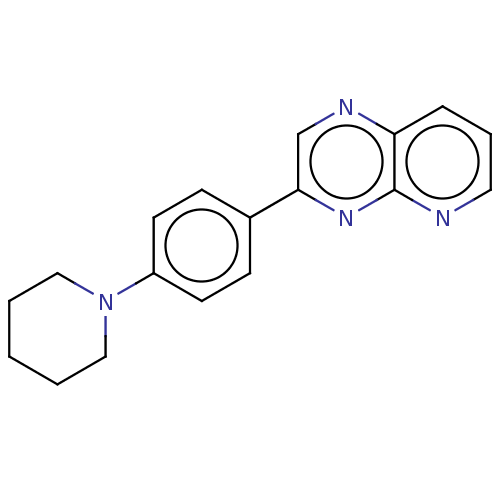
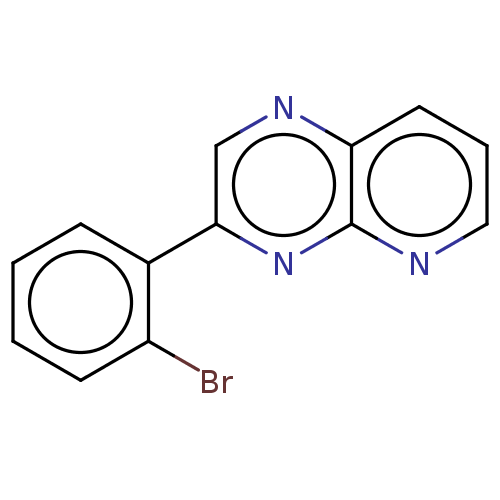
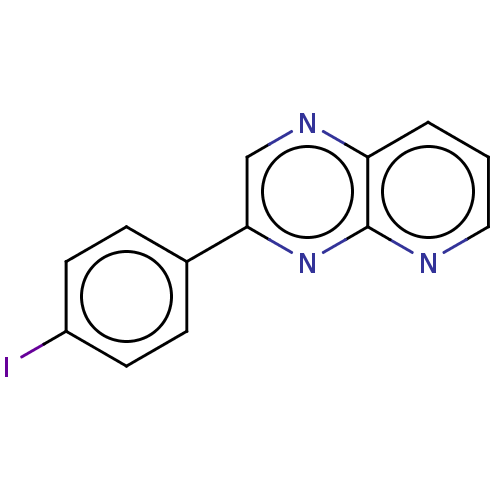
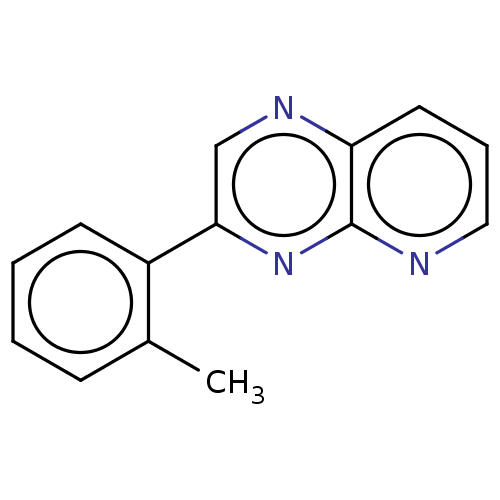
Report error Found 38 Enz. Inhib. hit(s) with all data for entry = 7033
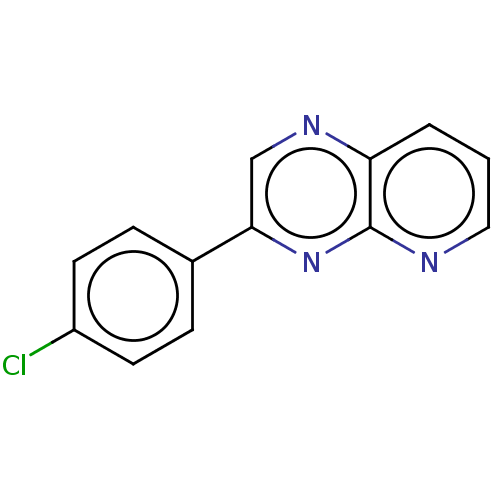
Affinity DataIC50: 32nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
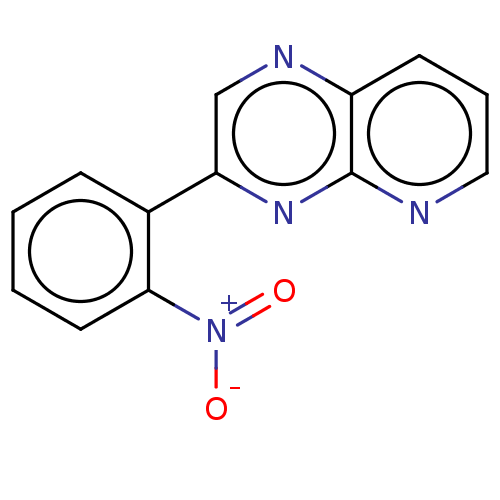
Affinity DataIC50: 466nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
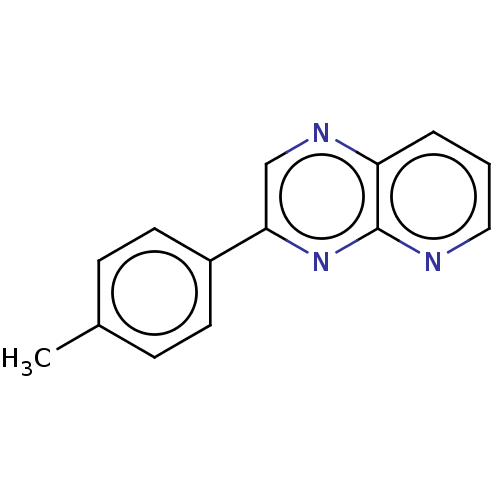
Affinity DataIC50: 583nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
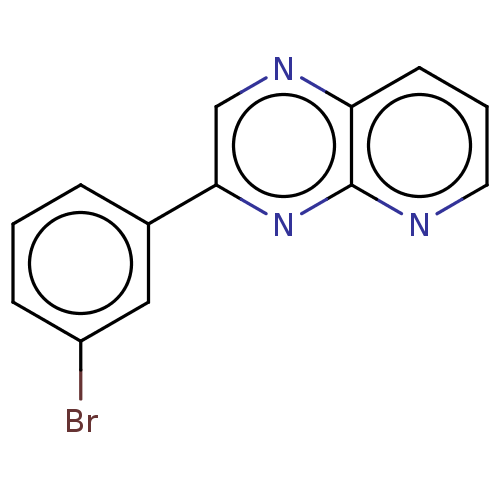
Affinity DataIC50: 899nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 1.26E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 1.89E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 1.97E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 2.45E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 2.52E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 2.59E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 2.63E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 3.06E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 3.12E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 3.91E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 4.03E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 4.05E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 4.75E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 5.39E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 5.78E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 5.88E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 6.41E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 6.44E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 7.45E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 9.65E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 9.67E+3nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 1.09E+4nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 1.12E+4nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 1.15E+4nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 1.21E+4nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 2.08E+4nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 2.12E+4nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 2.22E+4nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 2.23E+4nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 3.12E+4nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 3.54E+4nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 4.96E+4nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair
Affinity DataIC50: 7.98E+4nMpH: 8.0 T: 2°CAssay Description:A 96-microliter well plate was used for screening purpose. Each pyridopyrazine derivative was first dissolved in dimethyl sulfoxide to prepare the st...More data for this Ligand-Target Pair