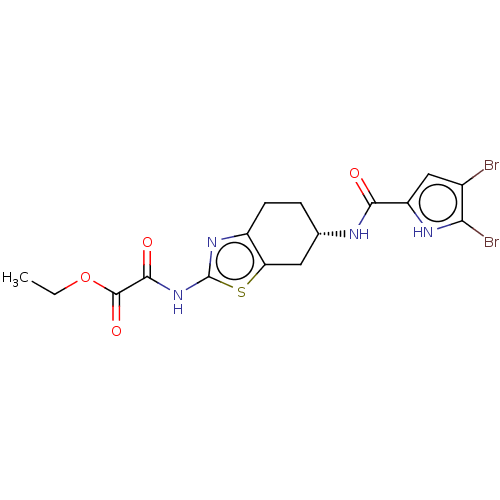
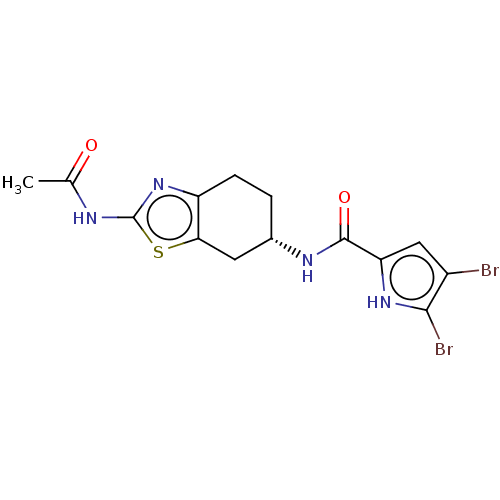
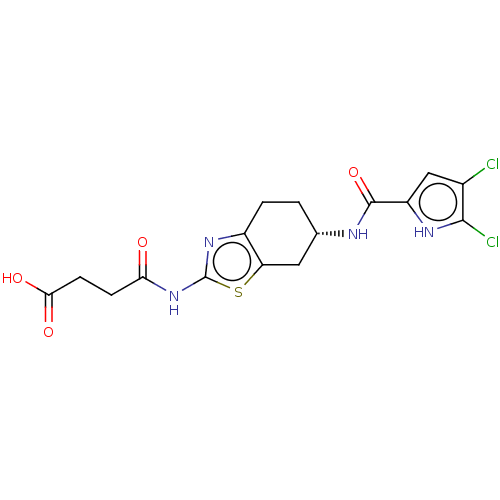
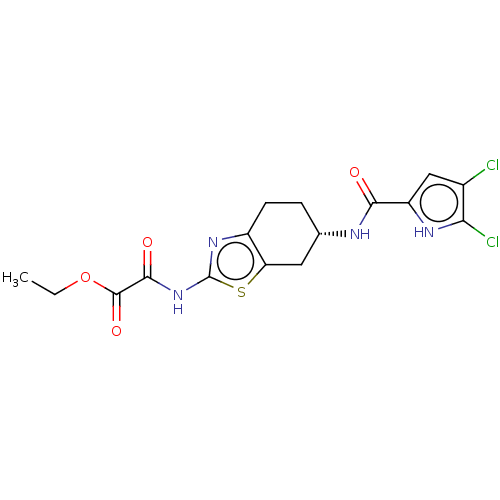
Report error Found 66 Enz. Inhib. hit(s) with all data for entry = 50046113
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataKd: 10nMAssay Description:Binding affinity to Escherichia coli DNA gyrase subunit B N-terminal 24 kDa domain by SPR assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase using relaxed pBR322 DNA as substrate after 3 hrs by DNA super-coiling assayMore data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence base...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataKd: 42nMAssay Description:Binding affinity to Escherichia coli DNA gyrase subunit B N-terminal 24 kDa domain by SPR assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 49nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataKd: 49nMAssay Description:Binding affinity to Escherichia coli DNA gyrase subunit B N-terminal 24 kDa domain by SPR assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataKd: 50nMAssay Description:Binding affinity to Escherichia coli DNA gyrase subunit B N-terminal 24 kDa domain by SPR assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 58nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 69nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataKd: 72nMAssay Description:Binding affinity to Escherichia coli DNA gyrase subunit B N-terminal 24 kDa domain by SPR assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 80nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pBR322 DNA as substrate after 3 hrs by DNA super-coiling assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 93nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 96nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 130nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataKd: 150nMAssay Description:Binding affinity to Escherichia coli DNA gyrase subunit B N-terminal 24 kDa domain by SPR assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 150nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 170nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 300nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 400nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 400nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 500nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 590nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 890nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 2.60E+3nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 2.90E+3nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataKd: 3.00E+3nMAssay Description:Binding affinity to Escherichia coli DNA gyrase subunit B N-terminal 24 kDa domain by SPR assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 5.70E+3nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 7.60E+3nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 7.70E+3nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 8.10E+3nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 9.10E+3nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 9.10E+3nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence base...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA topoisomerase 4 subunit A/B(Staphylococcus aureus)
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of Staphylococcus aureus DNA topoisomerase 4 using kinetoplast DNA as substrate by decatenation assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 2.10E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
Affinity DataIC50: 2.60E+4nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence base...More data for this Ligand-Target Pair
TargetDNA topoisomerase 4 subunit A/B(Staphylococcus aureus)
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 2.70E+4nMAssay Description:Inhibition of Staphylococcus aureus DNA topoisomerase 4 using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluoresc...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 2.90E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 3.50E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 3.50E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 4.10E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluorescence based DNA...More data for this Ligand-Target Pair
TargetDNA topoisomerase 4 subunit A/B(Staphylococcus aureus)
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 7.60E+4nMAssay Description:Inhibition of Staphylococcus aureus DNA topoisomerase 4 using relaxed pNO1 plasmid and biotinylated oligonucelotide incubated for 30 mins by fluoresc...More data for this Ligand-Target Pair