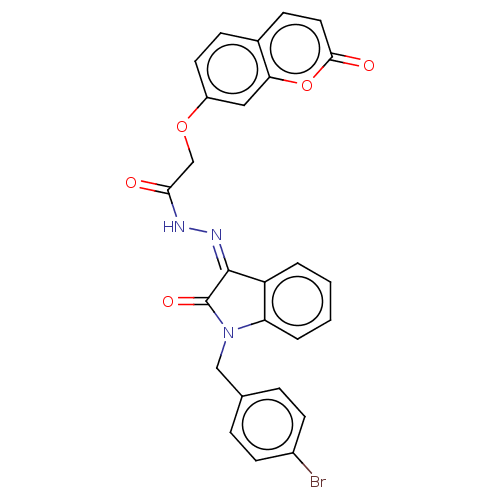
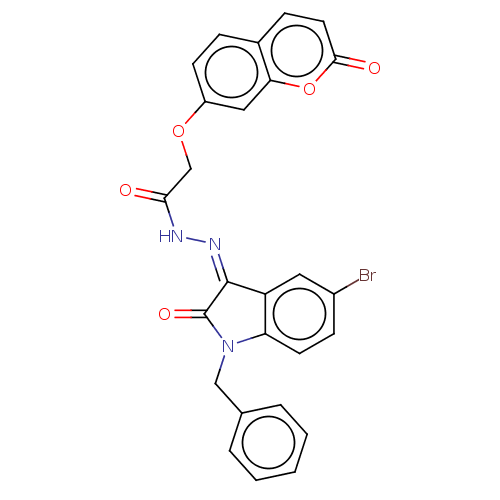
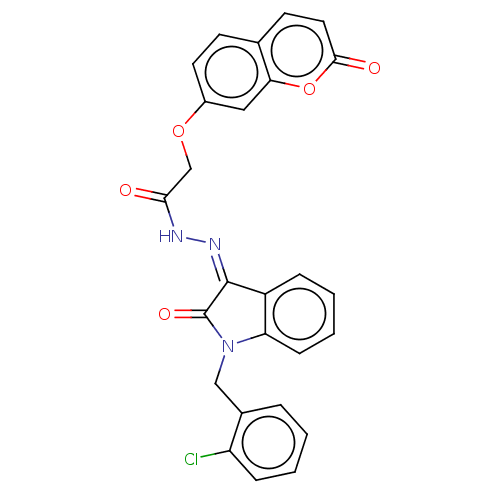
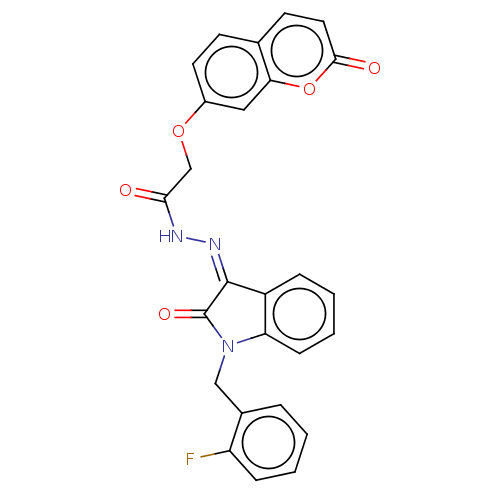
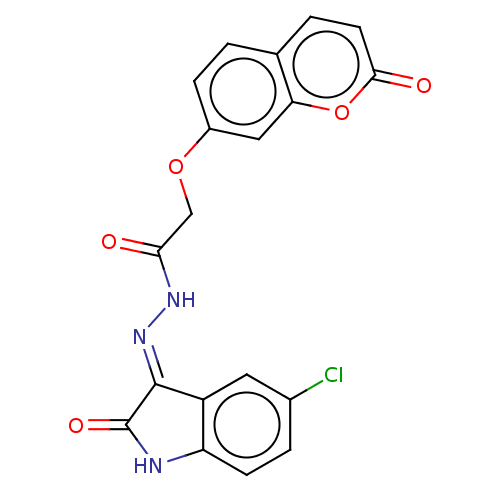
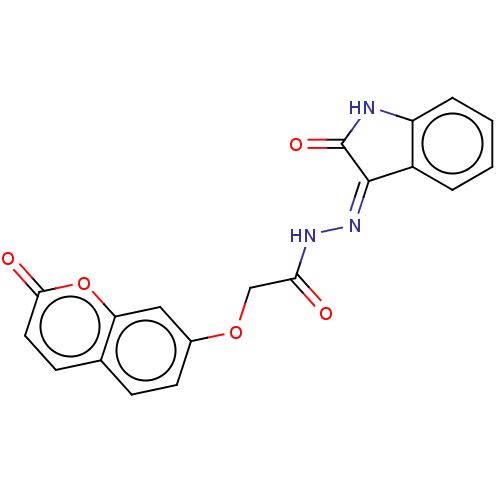
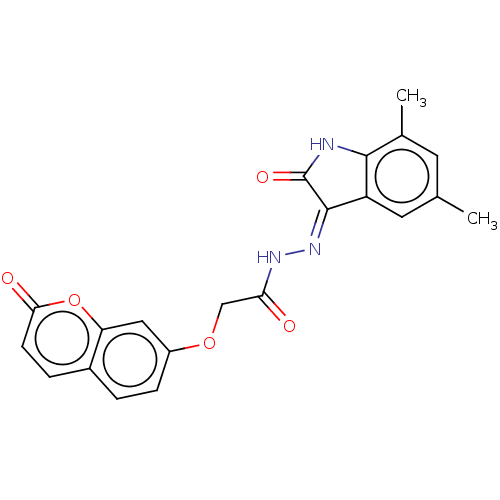
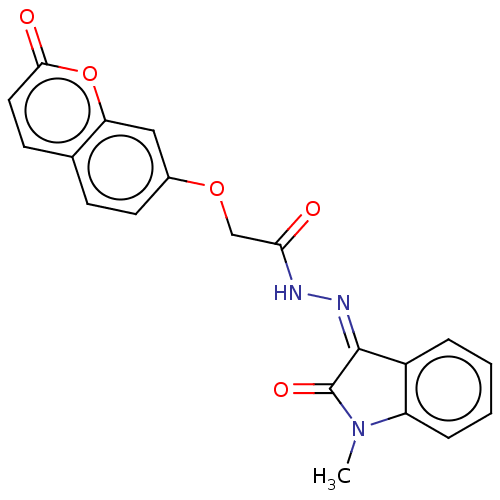
Report error Found 21 Enz. Inhib. hit(s) with all data for entry = 7889
Affinity DataIC50: 2.56E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 2.85E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 2.88E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 3.98E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 4.07E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 4.48E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 7.31E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 7.37E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 9.33E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 9.47E+3nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.41E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.42E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 6.68E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 7.78E+4nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.14E+5nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.53E+5nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.87E+5nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 1.97E+5nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 2.69E+5nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair
Affinity DataIC50: 8.17E+5nMpH: 6.8Assay Description:The test compounds were dissolved in DMSO to prepare the required distributing concentration. α-Glucosidase inhibitory activity was assayed usin...More data for this Ligand-Target Pair