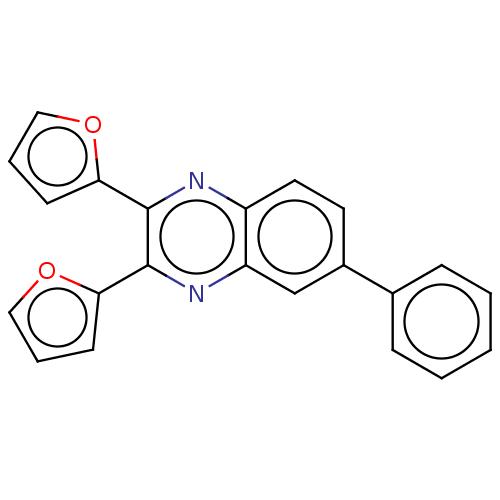
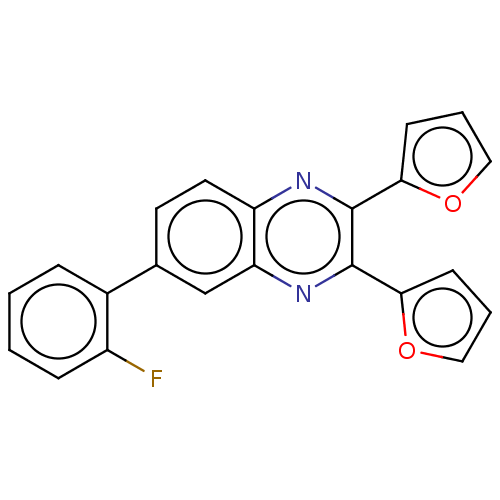
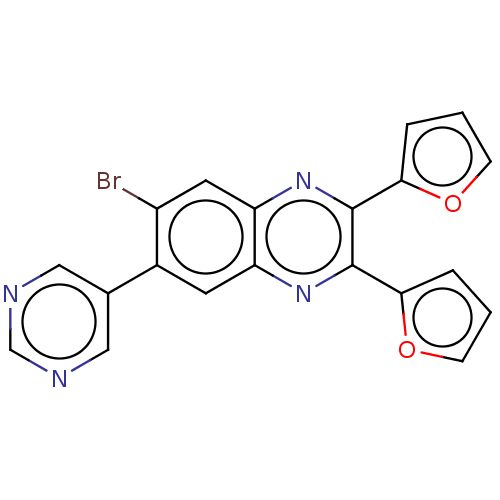
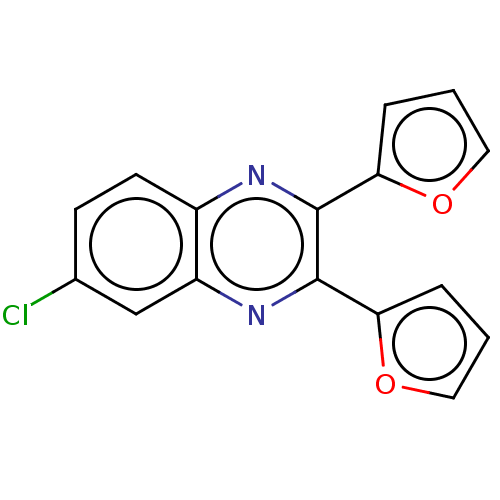
Report error Found 56 Enz. Inhib. hit(s) with all data for entry = 50003276
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 10nMAssay Description:Inhibition of PI3K-delta (unknown origin) after 40 mins by Kinase-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 15nMAssay Description:Inhibition of PI3K-beta (unknown origin) after 40 mins by Kinase-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 40nMAssay Description:Inhibition of recombinant full length PI3K p110alpha/p85alpha (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 as substrate pr...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 44nMAssay Description:Inhibition of PI3K-beta (unknown origin) after 40 mins by Kinase-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 50nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 50nMAssay Description:Inhibition of recombinant full length PI3K p110alpha/p85alpha (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 as substrate pr...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 51nMAssay Description:Inhibition of PI3K-gamma (unknown origin) after 40 mins by Kinase-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 67nMAssay Description:Inhibition of PI3K-gamma (unknown origin) after 40 mins by Kinase-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 93nMAssay Description:Inhibition of recombinant full length PI3K p110alpha/p85alpha (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 as substrate pr...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 93nMAssay Description:Inhibition of PI3K-delta (unknown origin) after 40 mins by Kinase-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 97nMAssay Description:Inhibition of recombinant full length PI3K p110alpha/p85alpha (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 as substrate pr...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 137nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 169nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 516nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 1.20E+3nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 1.90E+3nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 2.90E+3nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 3.20E+3nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 3.34E+3nMAssay Description:Inhibition of PI3K-beta (unknown origin) after 40 mins by Kinase-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 7.30E+3nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 9.89E+3nMAssay Description:Inhibition of PI3K-gamma (unknown origin) after 40 mins by Kinase-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 1.08E+4nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 1.18E+4nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 1.24E+4nMAssay Description:Inhibition of recombinant full length PI3K p110alpha/p85alpha (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 as substrate pr...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 1.27E+4nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 1.28E+4nMAssay Description:Inhibition of recombinant full length PI3K p110alpha/p85alpha (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 as substrate pr...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 1.35E+4nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 1.57E+4nMAssay Description:Inhibition of PI3K-delta (unknown origin) after 40 mins by Kinase-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 1.81E+4nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 1.86E+4nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 2.05E+4nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 2.76E+4nMAssay Description:Inhibition of recombinant full length PI3K p110alpha/p85alpha (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 as substrate pr...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 3.35E+4nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 3.93E+4nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 4.35E+4nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 4.78E+4nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 5.46E+4nMAssay Description:Inhibition of recombinant full length PI3K p110alpha/p85alpha (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 as substrate pr...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 7.60E+4nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: 8.73E+4nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 3-kinase regulatory subunit alpha/4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Chongqing University
Curated by ChEMBL
Chongqing University
Curated by ChEMBL
Affinity DataEC50: >1.00E+5nMAssay Description:Inhibition of recombinant full length PI3K p85alpha/p110alpha H1047R mutant (unknown origin) expressed in baculovirus infected sf9 cells using PIP2 a...More data for this Ligand-Target Pair