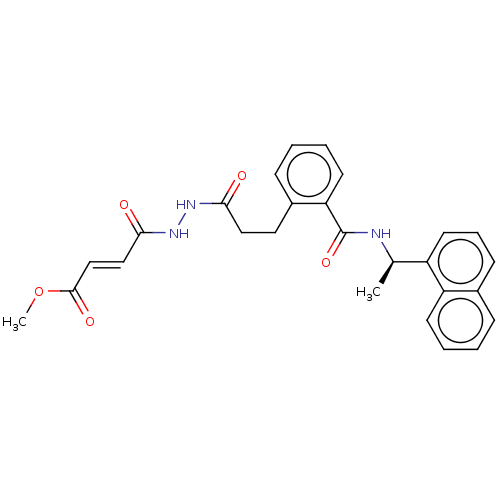
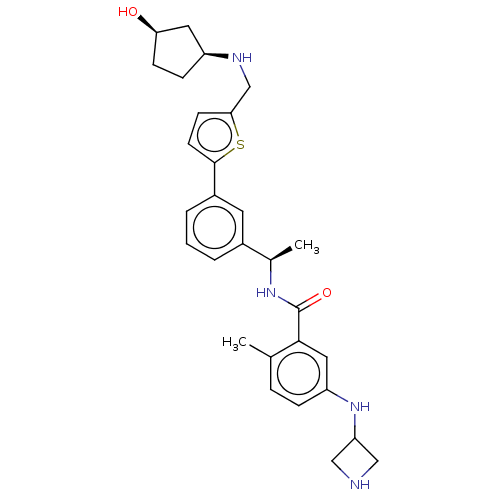
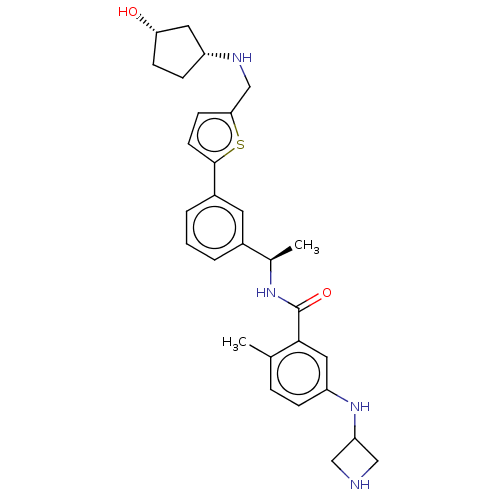
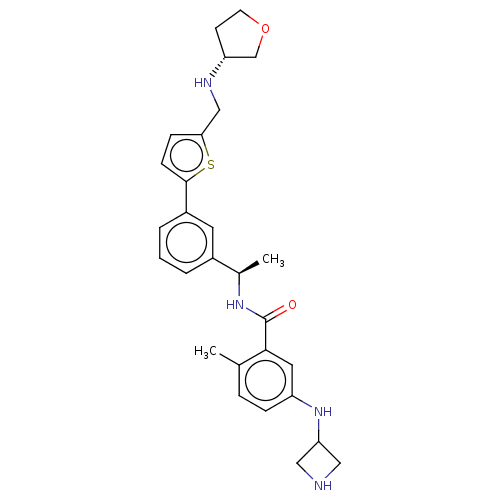
Report error Found 65 Enz. Inhib. hit(s) with all data for entry = 50021903
Affinity DataIC50: 94nMAssay Description:Inhibition of C-terminal his-6 tagged SARS CoV-2 PL protease expressed in Escherichia coli BL21 (DE3) cells incubated for 30 mins followed by substra...More data for this Ligand-Target Pair
Affinity DataIC50: 113nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataKd: 113nMAssay Description:Binding affinity to N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells assessed as dissociation const...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 260nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 320nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 330nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 330nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 360nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 370nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataKd: 370nMAssay Description:Binding affinity to N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells assessed as dissociation const...More data for this Ligand-Target Pair
Affinity DataIC50: 380nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 390nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 430nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 470nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 490nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 550nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 560nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 560nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 570nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 600nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 600nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 620nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 650nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 670nMAssay Description:Inhibition of C-terminal his-tagged SARS CoV-2 PL protease (1564 to 1876 residues) expressed in Escherichia coli BL21 (DE3) cells using ISG15-AMC as ...More data for this Ligand-Target Pair
Affinity DataIC50: 670nMAssay Description:Inhibition of C-terminal his-tagged SARS CoV-2 PL protease (1564 to 1876 residues) expressed in Escherichia coli BL21 (DE3) cells using ISG15-AMC as ...More data for this Ligand-Target Pair
Affinity DataIC50: 690nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 750nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 790nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 810nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 820nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 820nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 920nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 960nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 960nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 1.08E+3nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 1.13E+3nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 1.14E+3nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 1.19E+3nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 1.39E+3nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 1.39E+3nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 1.46E+3nMAssay Description:Inhibition of SARS CoV-2 PL proteaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.48E+3nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 1.52E+3nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 1.56E+3nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 1.61E+3nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair
Affinity DataIC50: 1.66E+3nMAssay Description:Inhibition of N-terminal his-tagged SARS CoV-2 PL protease (1564 to 1878 residues) expressed in BL21 (DE3) cells using Z-RLRGG-AMC as substrate prein...More data for this Ligand-Target Pair