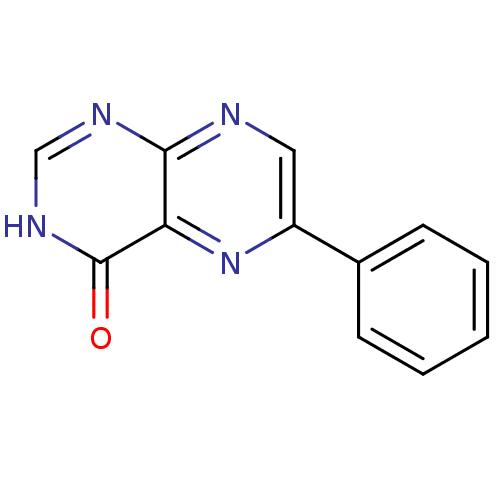
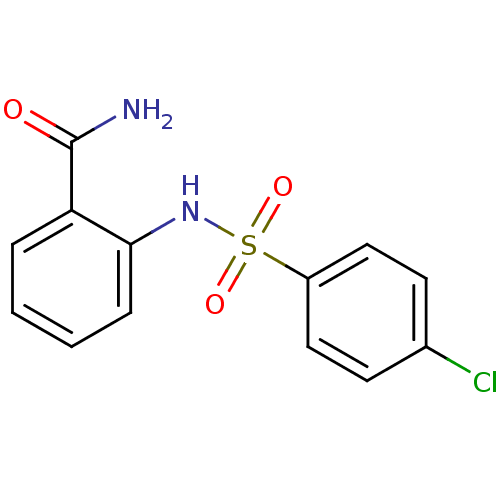
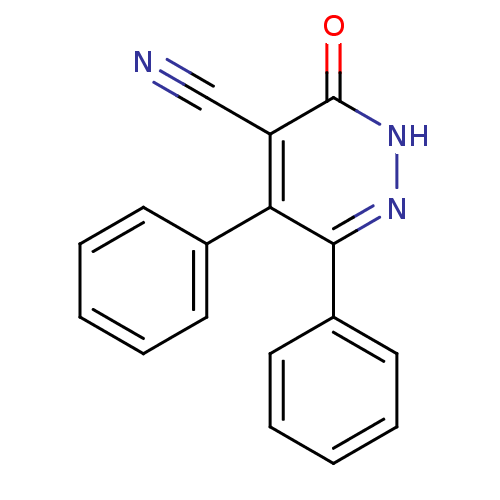
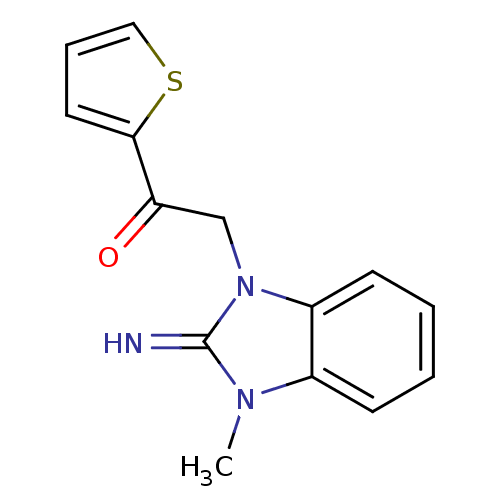
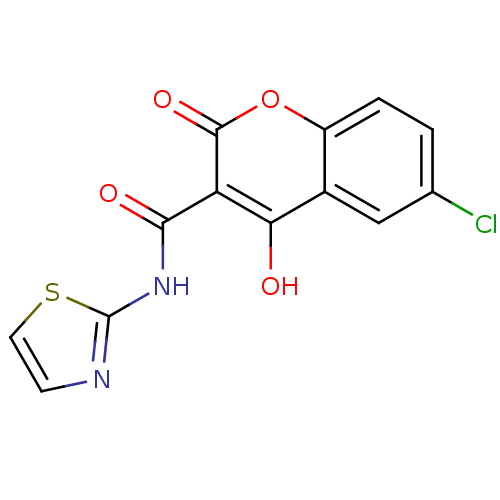
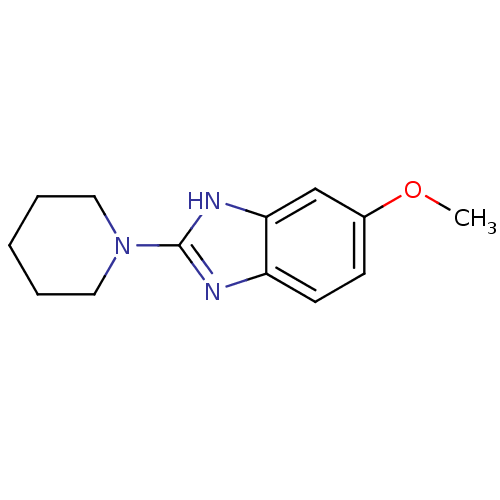
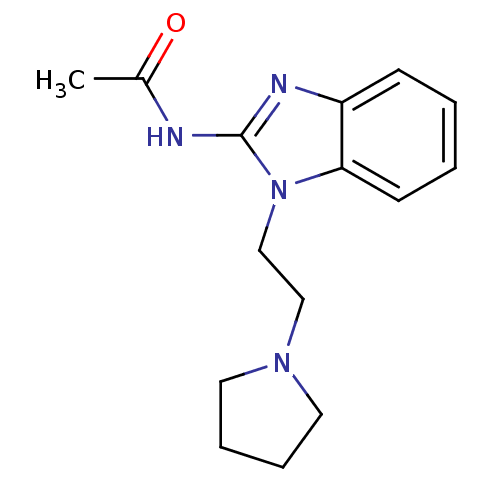
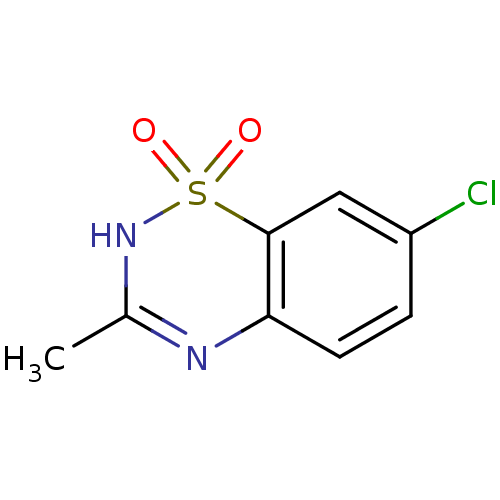
Report error Found 14 Enz. Inhib. hit(s) with all data for entry = 50020105
TargetATP-binding cassette sub-family C member 8/ATP-sensitive inward rectifier potassium channel 11(Human)
University of Perugia
Curated by ChEMBL
University of Perugia
Curated by ChEMBL
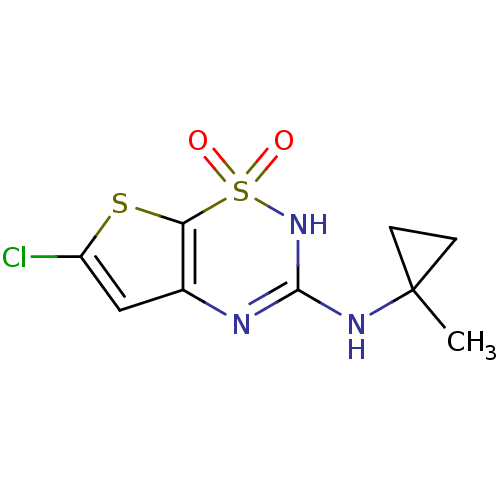
Affinity DataEC50: 190nMAssay Description:Activity at Kir6.2/SUR1 KATP channels expressed in HEK293 cells assessed as repolarization of tolbutamide-induced membrane depolarizationMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 8/ATP-sensitive inward rectifier potassium channel 11(Human)
University of Perugia
Curated by ChEMBL
University of Perugia
Curated by ChEMBL
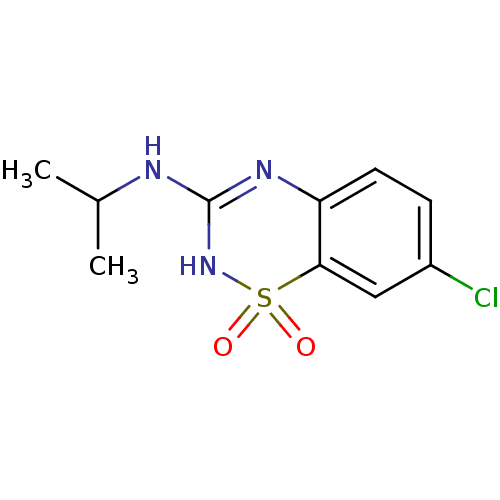
Affinity DataEC50: 250nMAssay Description:Activity at Kir6.2/SUR1 KATP channels expressed in HEK293 cells assessed as repolarization of tolbutamide-induced membrane depolarizationMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 8/ATP-sensitive inward rectifier potassium channel 11(Human)
University of Perugia
Curated by ChEMBL
University of Perugia
Curated by ChEMBL
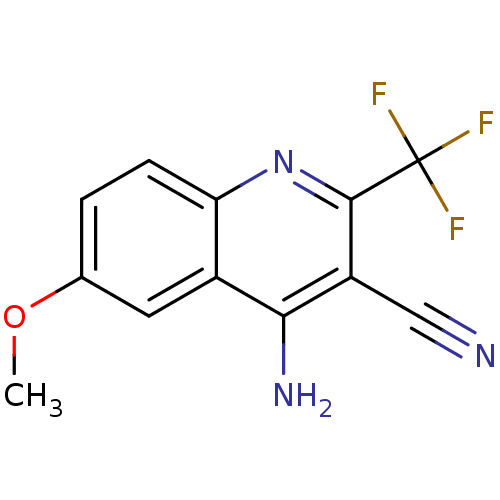
Affinity DataEC50: 450nMAssay Description:Activity at Kir6.2/SUR1 KATP channels expressed in HEK293 cells assessed as activation of K+ currentsMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 8/ATP-sensitive inward rectifier potassium channel 11(Human)
University of Perugia
Curated by ChEMBL
University of Perugia
Curated by ChEMBL
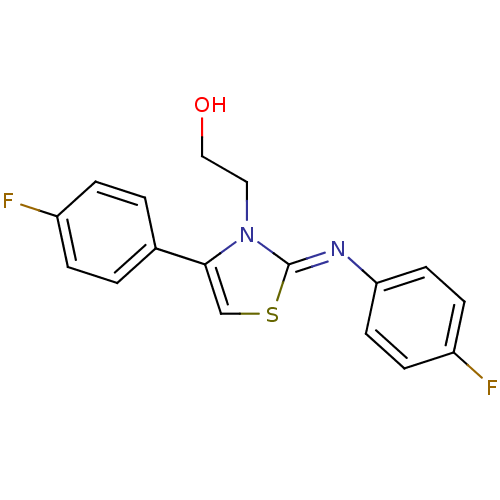
Affinity DataEC50: 1.50E+4nMAssay Description:Activity at Kir6.2/SUR1 KATP channels expressed in HEK293 cells assessed as repolarization of tolbutamide-induced membrane depolarizationMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 8/ATP-sensitive inward rectifier potassium channel 11(Human)
University of Perugia
Curated by ChEMBL
University of Perugia
Curated by ChEMBL
Affinity DataEC50: 2.10E+4nMAssay Description:Activity at Kir6.2/SUR1 KATP channels expressed in HEK293 cells assessed as repolarization of tolbutamide-induced membrane depolarizationMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 8/ATP-sensitive inward rectifier potassium channel 11(Human)
University of Perugia
Curated by ChEMBL
University of Perugia
Curated by ChEMBL
Affinity DataEC50: >3.00E+4nMAssay Description:Activity at Kir6.2/SUR1 KATP channels expressed in HEK293 cells assessed as repolarization of tolbutamide-induced membrane depolarizationMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 8/ATP-sensitive inward rectifier potassium channel 11(Human)
University of Perugia
Curated by ChEMBL
University of Perugia
Curated by ChEMBL
Affinity DataEC50: >3.00E+4nMAssay Description:Activity at Kir6.2/SUR1 KATP channels expressed in HEK293 cells assessed as repolarization of tolbutamide-induced membrane depolarizationMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 8/ATP-sensitive inward rectifier potassium channel 11(Human)
University of Perugia
Curated by ChEMBL
University of Perugia
Curated by ChEMBL
Affinity DataEC50: >3.00E+4nMAssay Description:Activity at Kir6.2/SUR1 KATP channels expressed in HEK293 cells assessed as repolarization of tolbutamide-induced membrane depolarizationMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 8/ATP-sensitive inward rectifier potassium channel 11(Human)
University of Perugia
Curated by ChEMBL
University of Perugia
Curated by ChEMBL
Affinity DataEC50: >3.00E+4nMAssay Description:Activity at Kir6.2/SUR1 KATP channels expressed in HEK293 cells assessed as repolarization of tolbutamide-induced membrane depolarizationMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 8/ATP-sensitive inward rectifier potassium channel 11(Human)
University of Perugia
Curated by ChEMBL
University of Perugia
Curated by ChEMBL
Affinity DataEC50: >3.00E+4nMAssay Description:Activity at Kir6.2/SUR1 KATP channels expressed in HEK293 cells assessed as repolarization of tolbutamide-induced membrane depolarizationMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 8/ATP-sensitive inward rectifier potassium channel 11(Human)
University of Perugia
Curated by ChEMBL
University of Perugia
Curated by ChEMBL
Affinity DataEC50: >3.00E+4nMAssay Description:Activity at Kir6.2/SUR1 KATP channels expressed in HEK293 cells assessed as repolarization of tolbutamide-induced membrane depolarizationMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 8/ATP-sensitive inward rectifier potassium channel 11(Human)
University of Perugia
Curated by ChEMBL
University of Perugia
Curated by ChEMBL
Affinity DataEC50: >3.00E+4nMAssay Description:Activity at Kir6.2/SUR1 KATP channels expressed in HEK293 cells assessed as repolarization of tolbutamide-induced membrane depolarizationMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 8/ATP-sensitive inward rectifier potassium channel 11(Human)
University of Perugia
Curated by ChEMBL
University of Perugia
Curated by ChEMBL
Affinity DataEC50: 3.10E+4nMAssay Description:Activity at Kir6.2/SUR1 KATP channels expressed in HEK293 cells assessed as activation of K+ currentsMore data for this Ligand-Target Pair
TargetATP-binding cassette sub-family C member 8/ATP-sensitive inward rectifier potassium channel 11(Human)
University of Perugia
Curated by ChEMBL
University of Perugia
Curated by ChEMBL
Affinity DataEC50: 3.30E+4nMAssay Description:Activity at Kir6.2/SUR1 KATP channels expressed in HEK293 cells assessed as repolarization of tolbutamide-induced membrane depolarizationMore data for this Ligand-Target Pair