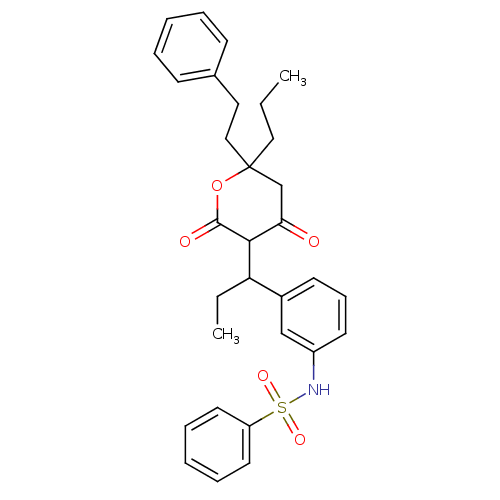
Report error Found 17 Enz. Inhib. hit(s) with all data for entry = 323
Affinity DataKi: 0.00700nM ΔG°: -63.0kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.00800nM ΔG°: -62.7kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.0180nM ΔG°: -60.7kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.0320nM ΔG°: -59.3kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.0400nM ΔG°: -58.8kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nM ΔG°: -57.8kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -56.5kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.120nM ΔG°: -56.1kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.120nM ΔG°: -56.1kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.220nM ΔG°: -54.6kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -53.8kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -50.9kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -50.9kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -48.2kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 6nM ΔG°: -46.5kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 8nM ΔG°: -45.7kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair
Affinity DataKi: 17nM ΔG°: -43.9kJ/molepH: 5.0 T: 2°CAssay Description:HIV-1 protease was purified and refolded from E. coli inclusion bodies. The substrate used spans the p17-p24 processing site (R-V-S-Q-N-Y-P-I-V-Q-N-K...More data for this Ligand-Target Pair