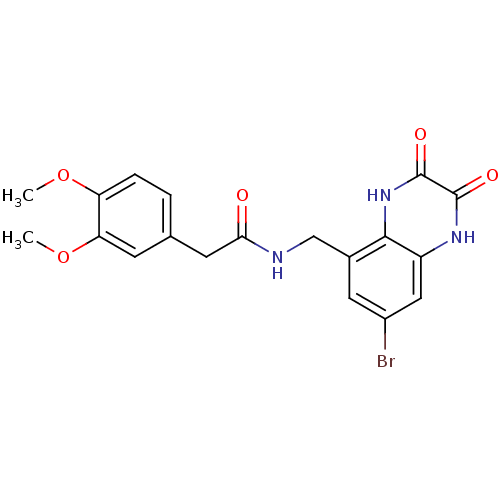
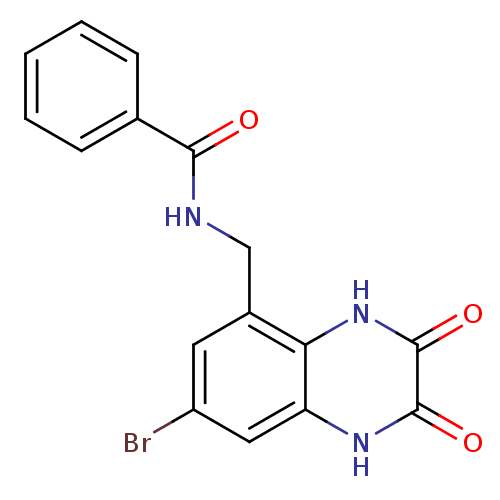
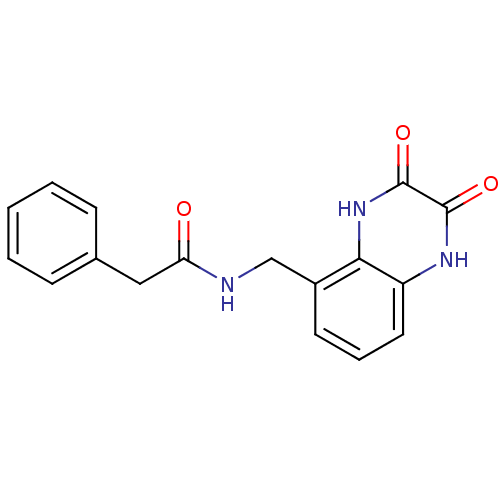
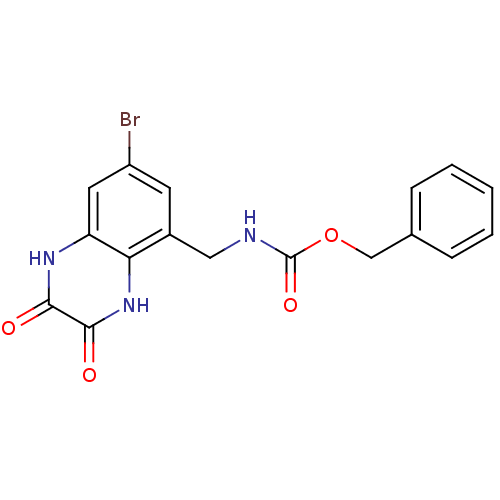
Report error Found 43 Enz. Inhib. hit(s) with all data for entry = 50034980
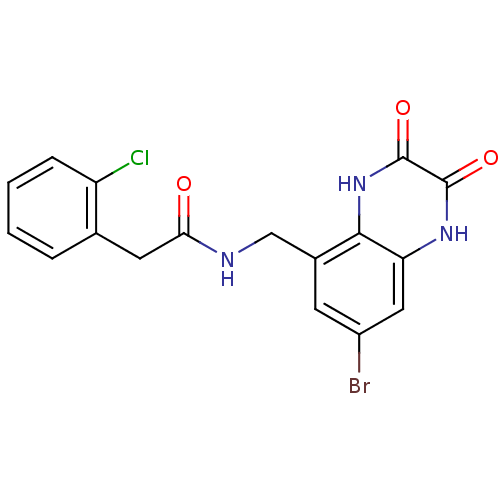
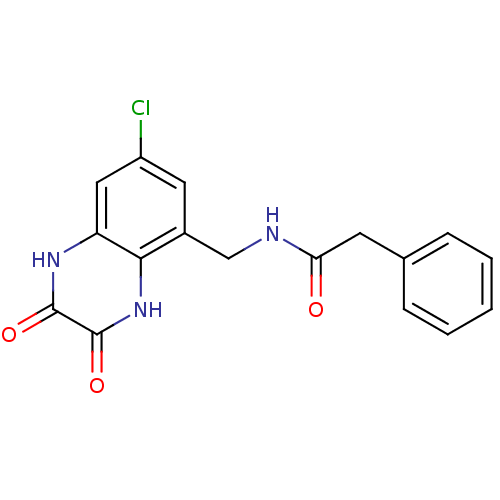
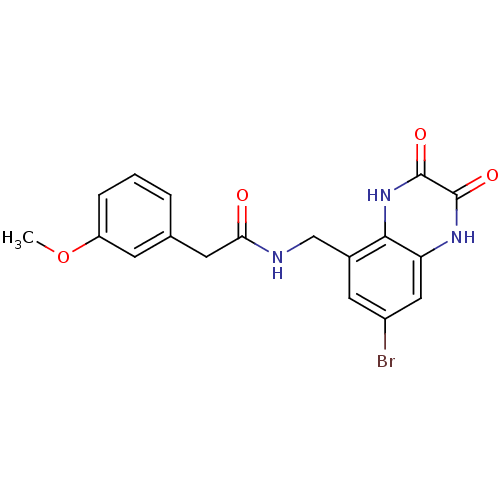
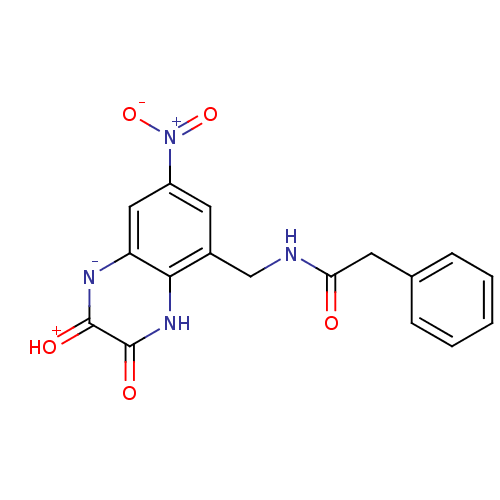
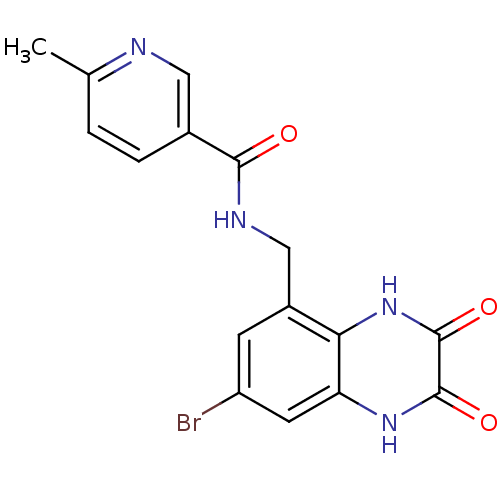
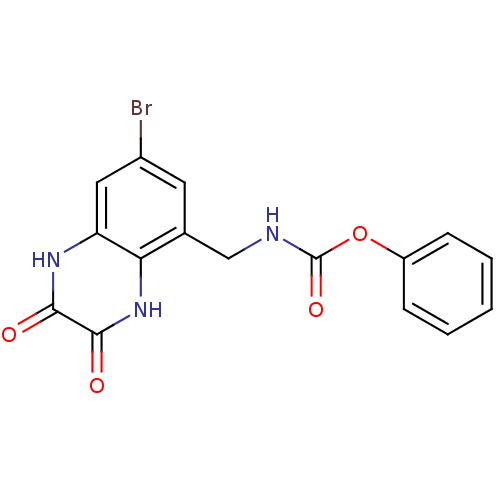
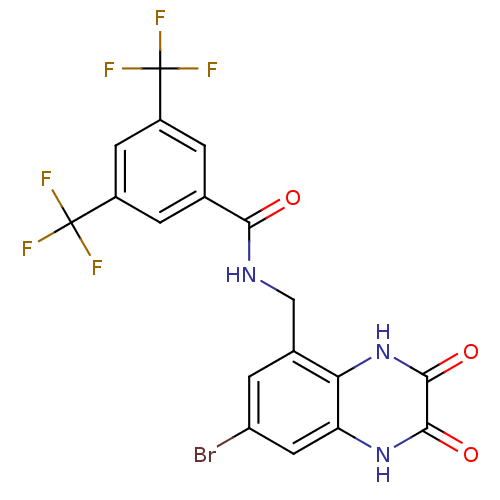
Affinity DataIC50: 7nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
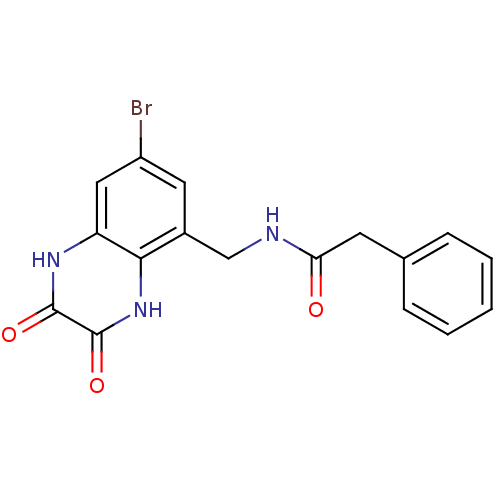
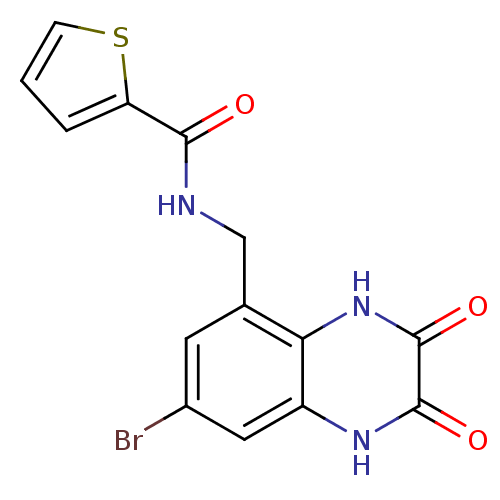
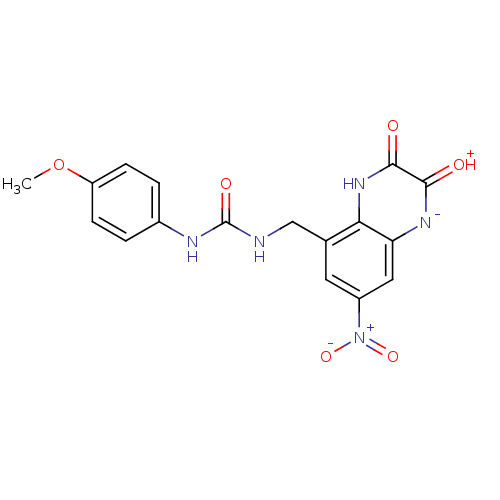
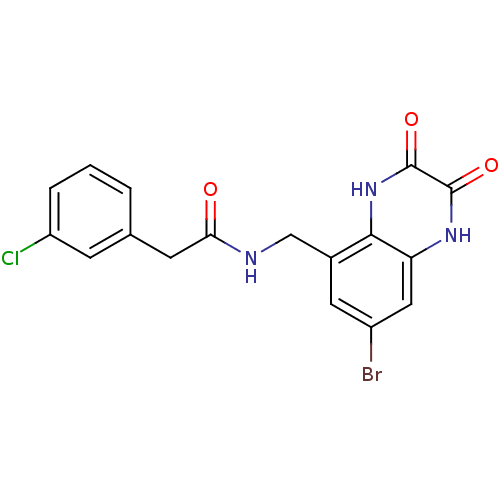
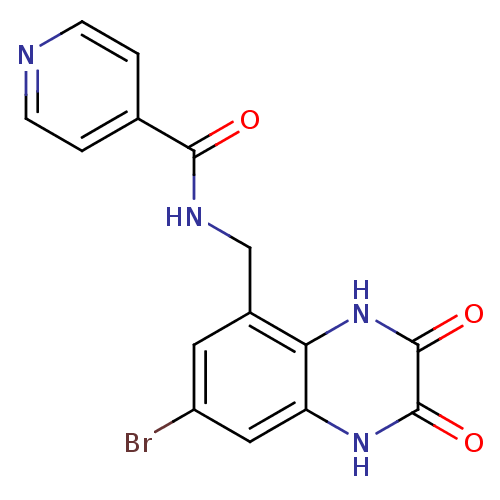
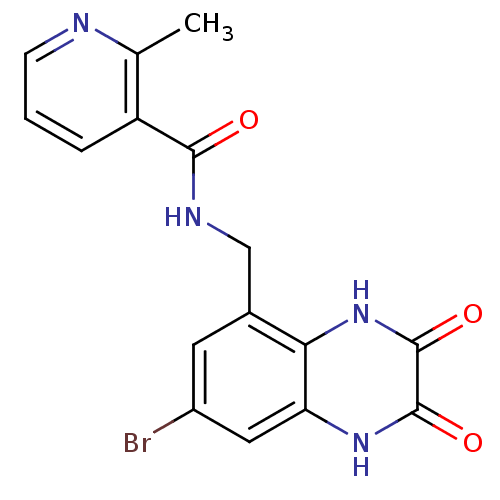
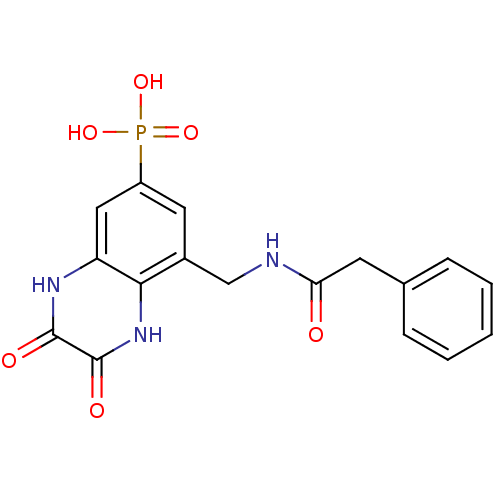
Affinity DataIC50: 10nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
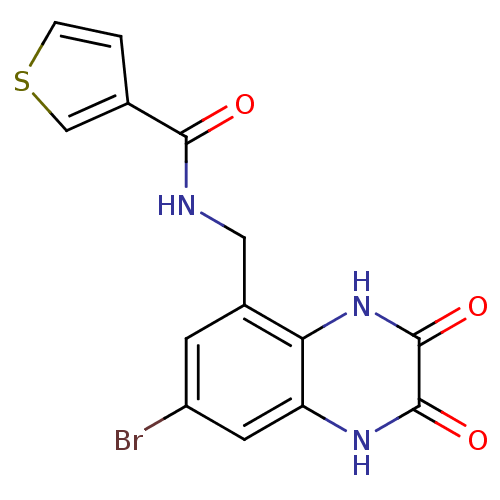
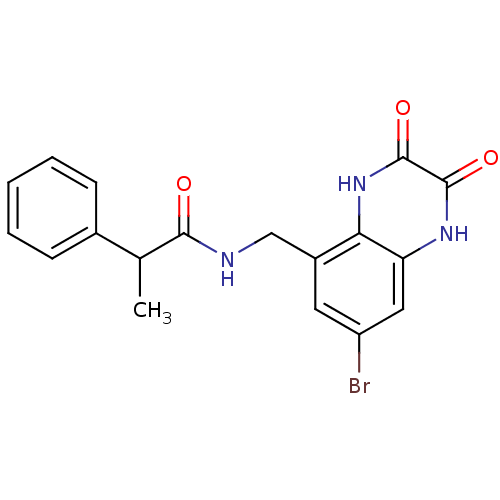
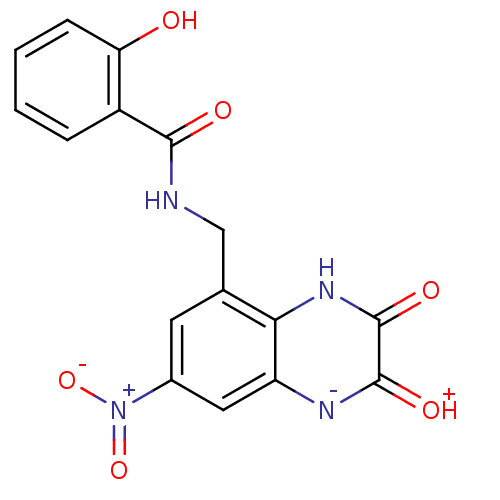
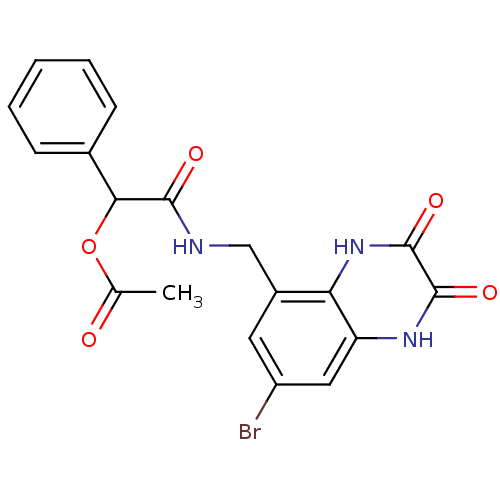
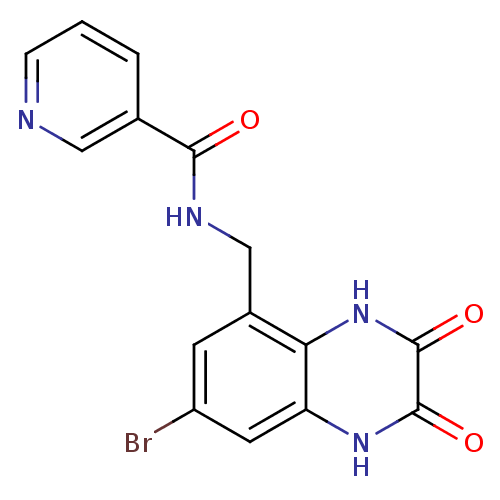
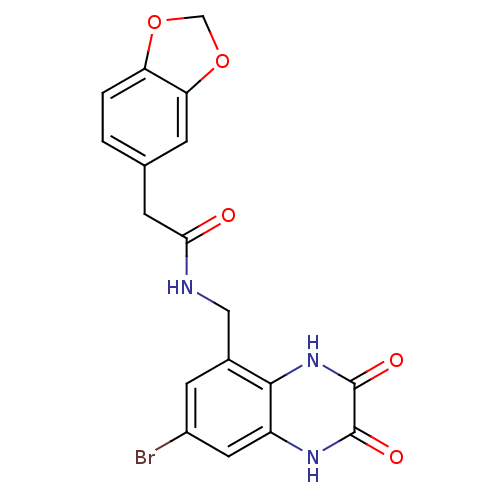
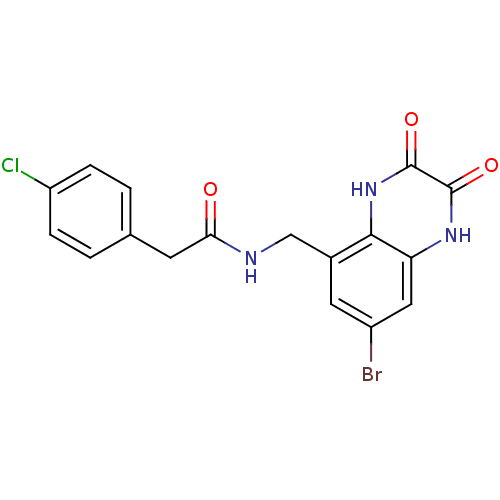
Affinity DataIC50: 10nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
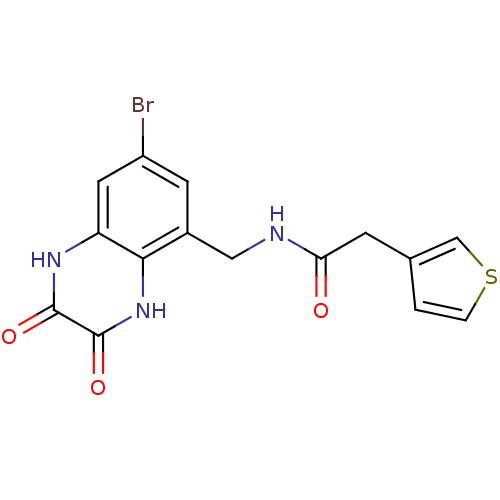
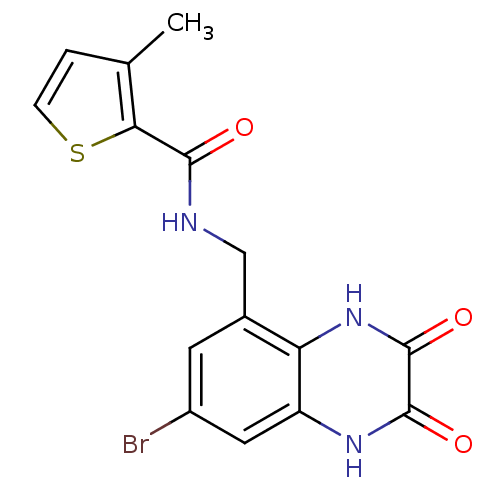
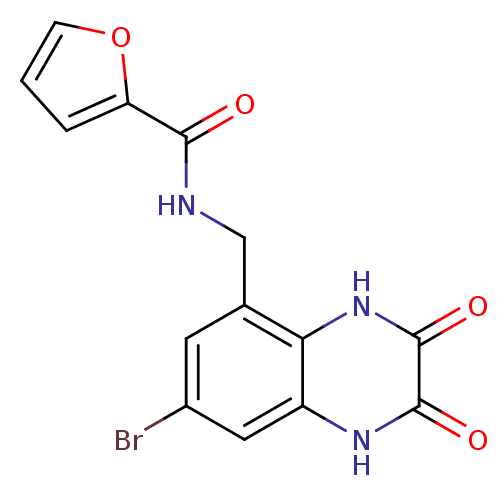
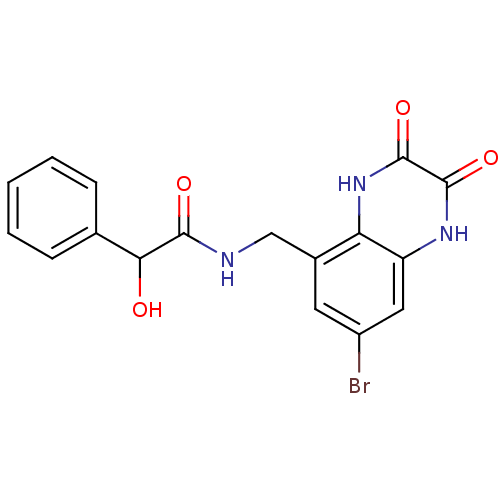
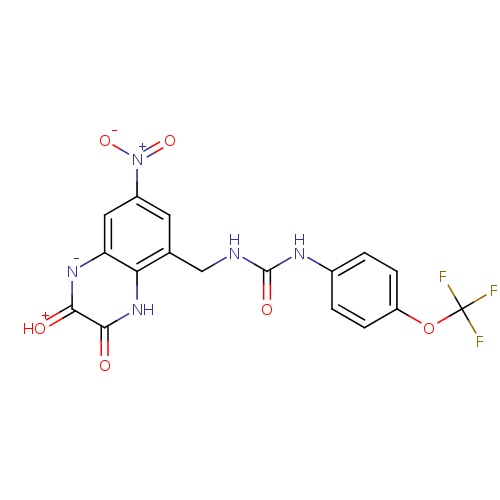
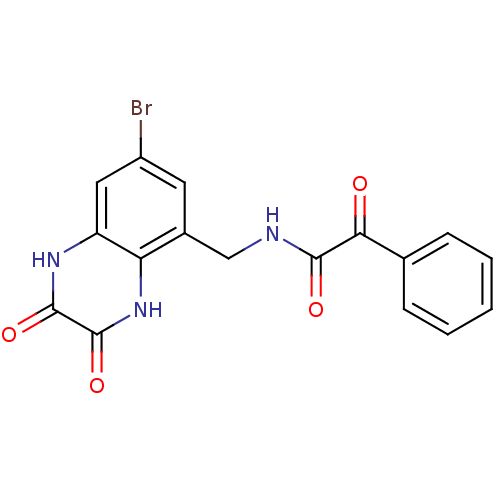
Affinity DataIC50: 10nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 90nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 250nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 280nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 350nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 630nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 860nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 900nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.90E+3nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:In vitro binding assay for the displacement of [3H]MDL-105519 from the glycine-site of NMDA receptorsMore data for this Ligand-Target Pair