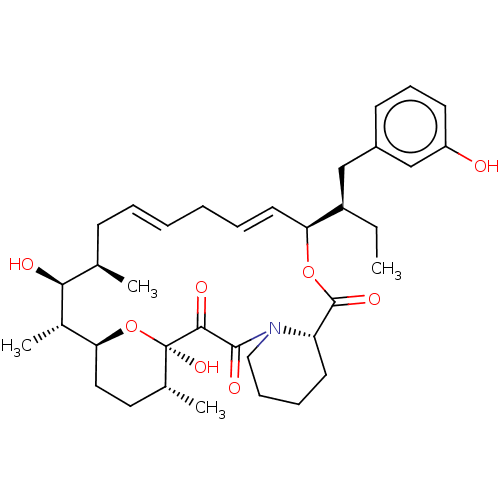
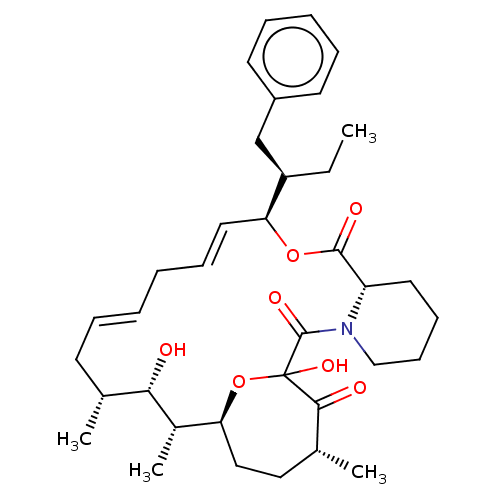
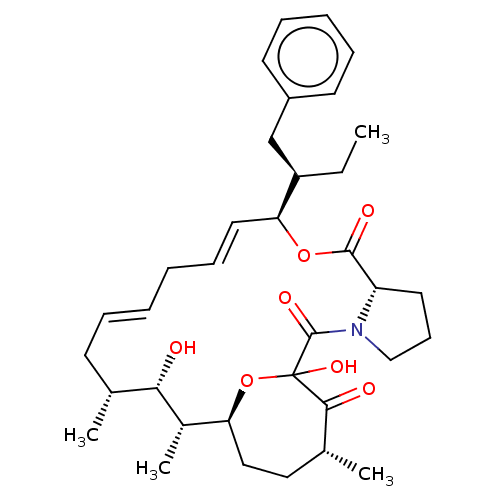
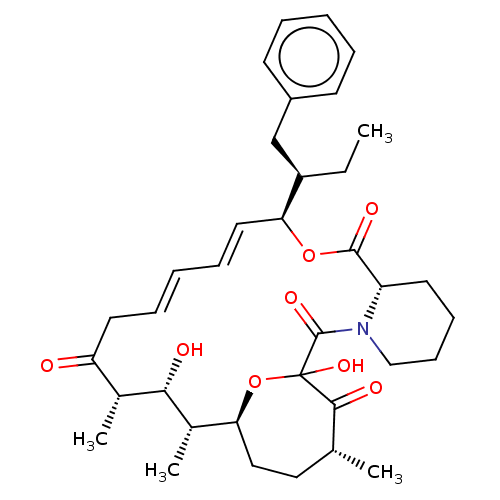
Report error Found 16 Enz. Inhib. hit(s) with all data for entry = 10102
Affinity DataKd: 0.210nMAssay Description:The binding of compounds of the invention to FKBP12 can be determined using the following protocol.General ProtocolThis protocol utilizes Perkin Elme...More data for this Ligand-Target Pair
Affinity DataKd: 0.340nMAssay Description:The binding of compounds of the invention to FKBP12 can be determined using the following protocol.General ProtocolThis protocol utilizes Perkin Elme...More data for this Ligand-Target Pair
Affinity DataKd: 0.510nMAssay Description:The binding of compounds of the invention to FKBP12 can be determined using the following protocol.General ProtocolThis protocol utilizes Perkin Elme...More data for this Ligand-Target Pair
Affinity DataKd: 0.950nMAssay Description:The binding of compounds of the invention to FKBP12 can be determined using the following protocol.General ProtocolThis protocol utilizes Perkin Elme...More data for this Ligand-Target Pair
Affinity DataKd: 1.10nMAssay Description:The binding of compounds of the invention to FKBP12 can be determined using the following protocol.General ProtocolThis protocol utilizes Perkin Elme...More data for this Ligand-Target Pair
Affinity DataKd: 1.30nMAssay Description:This protocol utilizes Surface Plasmon Resonance (SPR) as a method to determine kinetics (KD, Ka, Kd) for the binding of compound (analyte) to immobi...More data for this Ligand-Target Pair
Affinity DataKd: 1.5nMAssay Description:The binding of compounds of the invention to FKBP12 can be determined using the following protocol.General ProtocolThis protocol utilizes Perkin Elme...More data for this Ligand-Target Pair
Affinity DataKd: 4.80nMAssay Description:The binding of compounds of the invention to FKBP12 can be determined using the following protocol.General ProtocolThis protocol utilizes Perkin Elme...More data for this Ligand-Target Pair
Affinity DataKd: 7nMAssay Description:This protocol utilizes Surface Plasmon Resonance (SPR) as a method to determine kinetics (KD, Ka, Kd) for the binding of compound (analyte) to immobi...More data for this Ligand-Target Pair
Affinity DataKd: 12.6nMAssay Description:This protocol utilizes Surface Plasmon Resonance (SPR) as a method to determine kinetics (KD, Ka, Kd) for the binding of compound (analyte) to immobi...More data for this Ligand-Target Pair
Affinity DataKd: 18.8nMAssay Description:The binding of compounds of the invention to FKBP12 can be determined using the following protocol.General ProtocolThis protocol utilizes Perkin Elme...More data for this Ligand-Target Pair
Affinity DataKd: 21.5nMAssay Description:This protocol utilizes Surface Plasmon Resonance (SPR) as a method to determine kinetics (KD, Ka, Kd) for the binding of compound (analyte) to immobi...More data for this Ligand-Target Pair
Affinity DataKd: 23.1nMAssay Description:This protocol utilizes Surface Plasmon Resonance (SPR) as a method to determine kinetics (KD, Ka, Kd) for the binding of compound (analyte) to immobi...More data for this Ligand-Target Pair
Affinity DataKd: 71.5nMAssay Description:This protocol utilizes Surface Plasmon Resonance (SPR) as a method to determine kinetics (KD, Ka, Kd) for the binding of compound (analyte) to immobi...More data for this Ligand-Target Pair
Affinity DataKd: 355nMAssay Description:The binding of compounds of the invention to FKBP12 can be determined using the following protocol.General ProtocolThis protocol utilizes Perkin Elme...More data for this Ligand-Target Pair
Affinity DataKd: 785nMAssay Description:The binding of compounds of the invention to FKBP12 can be determined using the following protocol.General ProtocolThis protocol utilizes Perkin Elme...More data for this Ligand-Target Pair