Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Voltage-dependent N-type calcium channel subunit alpha-1B
Ligand
BDBM50111933
Substrate
n/a
Meas. Tech.
ChEBML_141440
IC50
610±n/a nM
Citation
 Seko, T; Kato, M; Kohno, H; Ono, S; Hashimura, K; Takimizu, H; Nakai, K; Maegawa, H; Katsube, N; Toda, M Structure-activity study of L-cysteine-based N-type calcium channel blockers: optimization of N- and C-terminal substituents. Bioorg Med Chem Lett 12:915-8 (2002) [PubMed] Article
Seko, T; Kato, M; Kohno, H; Ono, S; Hashimura, K; Takimizu, H; Nakai, K; Maegawa, H; Katsube, N; Toda, M Structure-activity study of L-cysteine-based N-type calcium channel blockers: optimization of N- and C-terminal substituents. Bioorg Med Chem Lett 12:915-8 (2002) [PubMed] Article More Info.:
Target
Name:
Voltage-dependent N-type calcium channel subunit alpha-1B
Synonyms:
BIII | Brain calcium channel III | CAC1B_HUMAN | CACH5 | CACNA1B | CACNL1A5 | Calcium channel (Type N) | Calcium channel, L type, alpha-1 polypeptide isoform 5 | Voltage-dependent N-type calcium channel subunit alpha-1B | Voltage-dependent N-type calcium channel subunit alpha-1B/Voltage-dependent calcium channel subunit alpha-2/delta-1/Voltage-dependent L-type calcium channel subunit beta-3 | Voltage-gated N-type calcium channel alpha-1B subunit | Voltage-gated N-type calcium channel alpha-1B subunit/Amyloid beta A4 precursor protein-binding family A member 1 | Voltage-gated calcium channel | Voltage-gated calcium channel subunit alpha Cav2.2 | Voltage-gated calcium channel subunit alpha Cav2.2 ((alpha 1B, beta 1b, alpha 2 delta-1) | calcium channel, voltage-dependent, N type, alpha 1B subunit
Type:
Enzyme
Mol. Mass.:
262548.16
Organism:
Human
Description:
Q00975
Residue:
2339
Sequence:
MVRFGDELGGRYGGPGGGERARGGGAGGAGGPGPGGLQPGQRVLYKQSIAQRARTMALYNPIPVKQNCFTVNRSLFVFSEDNVVRKYAKRITEWPPFEYMILATIIANCIVLALEQHLPDGDKTPMSERLDDTEPYFIGIFCFEAGIKIIALGFVFHKGSYLRNGWNVMDFVVVLTGILATAGTDFDLRTLRAVRVLRPLKLVSGIPSLQVVLKSIMKAMVPLLQIGLLLFFAILMFAIIGLEFYMGKFHKACFPNSTDAEPVGDFPCGKEAPARLCEGDTECREYWPGPNFGITNFDNILFAILTVFQCITMEGWTDILYNTNDAAGNTWNWLYFIPLIIIGSFFMLNLVLGVLSGEFAKERERVENRRAFLKLRRQQQIERELNGYLEWIFKAEEVMLAEEDRNAEEKSPLDVLKRAATKKSRNDLIHAEEGEDRFADLCAVGSPFARASLKSGKTESSSYFRRKEKMFRFFIRRMVKAQSFYWVVLCVVALNTLCVAMVHYNQPRRLTTTLYFAEFVFLGLFLTEMSLKMYGLGPRSYFRSSFNCFDFGVIVGSVFEVVWAAIKPGSSFGISVLRALRLLRIFKVTKYWSSLRNLVVSLLNSMKSIISLLFLLFLFIVVFALLGMQLFGGQFNFQDETPTTNFDTFPAAILTVFQILTGEDWNAVMYHGIESQGGVSKGMFSSFYFIVLTLFGNYTLLNVFLAIAVDNLANAQELTKDEEEMEEAANQKLALQKAKEVAEVSPMSAANISIAARQQNSAKARSVWEQRASQLRLQNLRASCEALYSEMDPEERLRFATTRHLRPDMKTHLDRPLVVELGRDGARGPVGGKARPEAAEAPEGVDPPRRHHRHRDKDKTPAAGDQDRAEAPKAESGEPGAREERPRPHRSHSKEAAGPPEARSERGRGPGPEGGRRHHRRGSPEEAAEREPRRHRAHRHQDPSKECAGAKGERRARHRGGPRAGPREAESGEEPARRHRARHKAQPAHEAVEKETTEKEATEKEAEIVEADKEKELRNHQPREPHCDLETSGTVTVGPMHTLPSTCLQKVEEQPEDADNQRNVTRMGSQPPDPNTIVHIPVMLTGPLGEATVVPSGNVDLESQAEGKKEVEADDVMRSGPRPIVPYSSMFCLSPTNLLRRFCHYIVTMRYFEVVILVVIALSSIALAAEDPVRTDSPRNNALKYLDYIFTGVFTFEMVIKMIDLGLLLHPGAYFRDLWNILDFIVVSGALVAFAFSGSKGKDINTIKSLRVLRVLRPLKTIKRLPKLKAVFDCVVNSLKNVLNILIVYMLFMFIFAVIAVQLFKGKFFYCTDESKELERDCRGQYLDYEKEEVEAQPRQWKKYDFHYDNVLWALLTLFTVSTGEGWPMVLKHSVDATYEEQGPSPGYRMELSIFYVVYFVVFPFFFVNIFVALIIITFQEQGDKVMSECSLEKNERACIDFAISAKPLTRYMPQNRQSFQYKTWTFVVSPPFEYFIMAMIALNTVVLMMKFYDAPYEYELMLKCLNIVFTSMFSMECVLKIIAFGVLNYFRDAWNVFDFVTVLGSITDILVTEIAETNNFINLSFLRLFRAARLIKLLRQGYTIRILLWTFVQSFKALPYVCLLIAMLFFIYAIIGMQVFGNIALDDDTSINRHNNFRTFLQALMLLFRSATGEAWHEIMLSCLSNQACDEQANATECGSDFAYFYFVSFIFLCSFLMLNLFVAVIMDNFEYLTRDSSILGPHHLDEFIRVWAEYDPAACGRISYNDMFEMLKHMSPPLGLGKKCPARVAYKRLVRMNMPISNEDMTVHFTSTLMALIRTALEIKLAPAGTKQHQCDAELRKEISVVWANLPQKTLDLLVPPHKPDEMTVGKVYAALMIFDFYKQNKTTRDQMQQAPGGLSQMGPVSLFHPLKATLEQTQPAVLRGARVFLRQKSSTSLSNGGAIQNQESGIKESVSWGTQRTQDAPHEARPPLERGHSTEIPVGRSGALAVDVQMQSITRRGPDGEPQPGLESQGRAASMPRLAAETQPVTDASPMKRSISTLAQRPRGTHLCSTTPDRPPPSQASSHHHHHRCHRRRDRKQRSLEKGPSLSADMDGAPSSAVGPGLPPGEGPTGCRRERERRQERGRSQERRQPSSSSSEKQRFYSCDRFGGREPPKPKPSLSSHPTSPTAGQEPGPHPQGSGSVNGSPLLSTSGASTPGRGGRRQLPQTPLTPRPSITYKTANSSPIHFAGAQTSLPAFSPGRLSRGLSEHNALLQRDPLSQPLAPGSRIGSDPYLGQRLDSEASVHALPEDTLTFEEAVATNSGRSSRTSYVSSLTSQSHPLRRVPNGYHCTLGLSSGGRARHSYHHPDQDHWC
Inhibitor
Name:
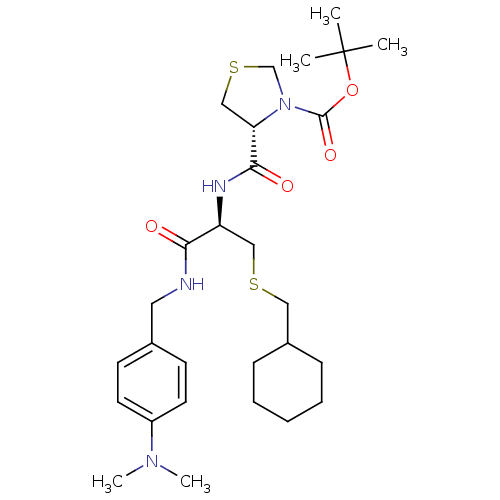
BDBM50111933
Synonyms:
(R)-4-[(R)-2-Cyclohexylmethylsulfanyl-1-(4-dimethylamino-benzylcarbamoyl)-ethylcarbamoyl]-thiazolidine-3-carboxylic acid tert-butyl ester | CHEMBL366590
Type:
Small organic molecule
Emp. Form.:
C28H44N4O4S2
Mol. Mass.:
564.803
SMILES:
CN(C)c1ccc(CNC(=O)[C@H](CSCC2CCCCC2)NC(=O)[C@@H]2CSCN2C(=O)OC(C)(C)C)cc1