Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Serine/threonine-protein kinase B-raf
Ligand
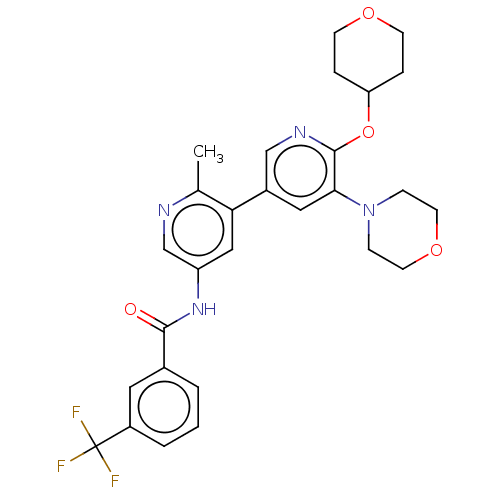
BDBM202784
Substrate
n/a
Meas. Tech.
ChEMBL_1663100 (CHEMBL4012781)
IC50
0.400000±n/a nM
Citation
 Nishiguchi, GA; Rico, A; Tanner, H; Aversa, RJ; Taft, BR; Subramanian, S; Setti, L; Burger, MT; Wan, L; Tamez, V; Smith, A; Lou, Y; Barsanti, PA; Appleton, BA; Mamo, M; Tandeske, L; Dix, I; Tellew, JE; Huang, S; Mathews Griner, LA; Cooke, VG; Van Abbema, A; Merritt, H; Ma, S; Gampa, K; Feng, F; Yuan, J; Wang, Y; Haling, JR; Vaziri, S; Hekmat-Nejad, M; Jansen, JM; Polyakov, V; Zang, R; Sethuraman, V; Amiri, P; Singh, M; Lees, E; Shao, W; Stuart, DD; Dillon, MP; Ramurthy, S Design and Discovery of N-(2-Methyl-5'-morpholino-6'-((tetrahydro-2H-pyran-4-yl)oxy)-[3,3'-bipyridin]-5-yl)-3-(trifluoromethyl)benzamide (RAF709): A Potent, Selective, and Efficacious RAF Inhibitor Targeting RAS Mutant Cancers. J Med Chem 60:4869-4881 (2017) [PubMed] Article
Nishiguchi, GA; Rico, A; Tanner, H; Aversa, RJ; Taft, BR; Subramanian, S; Setti, L; Burger, MT; Wan, L; Tamez, V; Smith, A; Lou, Y; Barsanti, PA; Appleton, BA; Mamo, M; Tandeske, L; Dix, I; Tellew, JE; Huang, S; Mathews Griner, LA; Cooke, VG; Van Abbema, A; Merritt, H; Ma, S; Gampa, K; Feng, F; Yuan, J; Wang, Y; Haling, JR; Vaziri, S; Hekmat-Nejad, M; Jansen, JM; Polyakov, V; Zang, R; Sethuraman, V; Amiri, P; Singh, M; Lees, E; Shao, W; Stuart, DD; Dillon, MP; Ramurthy, S Design and Discovery of N-(2-Methyl-5'-morpholino-6'-((tetrahydro-2H-pyran-4-yl)oxy)-[3,3'-bipyridin]-5-yl)-3-(trifluoromethyl)benzamide (RAF709): A Potent, Selective, and Efficacious RAF Inhibitor Targeting RAS Mutant Cancers. J Med Chem 60:4869-4881 (2017) [PubMed] Article More Info.:
Target
Name:
Serine/threonine-protein kinase B-raf
Synonyms:
B-RAF | B-Raf Protein Kinase | B-Raf proto-oncogene serine/threonine-protein kinase | BRAF | BRAF1 | BRAF_HUMAN | RAFB1 | p94 | v-Raf murine sarcoma viral oncogene homolog B1
Type:
Serine/threonine-protein kinase
Mol. Mass.:
84446.00
Organism:
Homo sapiens (Human)
Description:
P15056
Residue:
766
Sequence:
MAALSGGGGGGAEPGQALFNGDMEPEAGAGAGAAASSAADPAIPEEVWNIKQMIKLTQEHIEALLDKFGGEHNPPSIYLEAYEEYTSKLDALQQREQQLLESLGNGTDFSVSSSASMDTVTSSSSSSLSVLPSSLSVFQNPTDVARSNPKSPQKPIVRVFLPNKQRTVVPARCGVTVRDSLKKALMMRGLIPECCAVYRIQDGEKKPIGWDTDISWLTGEELHVEVLENVPLTTHNFVRKTFFTLAFCDFCRKLLFQGFRCQTCGYKFHQRCSTEVPLMCVNYDQLDLLFVSKFFEHHPIPQEEASLAETALTSGSSPSAPASDSIGPQILTSPSPSKSIPIPQPFRPADEDHRNQFGQRDRSSSAPNVHINTIEPVNIDDLIRDQGFRGDGGSTTGLSATPPASLPGSLTNVKALQKSPGPQRERKSSSSSEDRNRMKTLGRRDSSDDWEIPDGQITVGQRIGSGSFGTVYKGKWHGDVAVKMLNVTAPTPQQLQAFKNEVGVLRKTRHVNILLFMGYSTKPQLAIVTQWCEGSSLYHHLHIIETKFEMIKLIDIARQTAQGMDYLHAKSIIHRDLKSNNIFLHEDLTVKIGDFGLATVKSRWSGSHQFEQLSGSILWMAPEVIRMQDKNPYSFQSDVYAFGIVLYELMTGQLPYSNINNRDQIIFMVGRGYLSPDLSKVRSNCPKAMKRLMAECLKKKRDERPLFPQILASIELLARSLPKIHRSASEPSLNRAGFQTEDFSLYACASPKTPIQAGGYGAFPVH