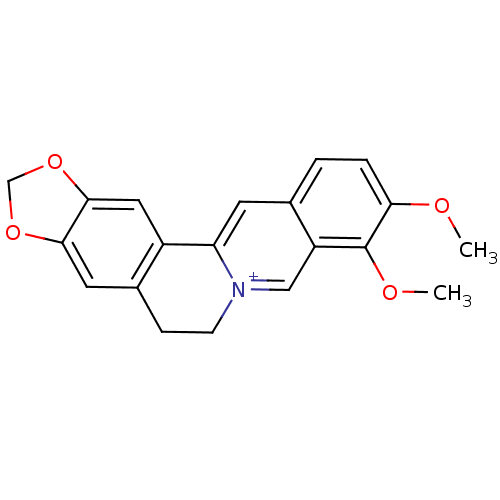
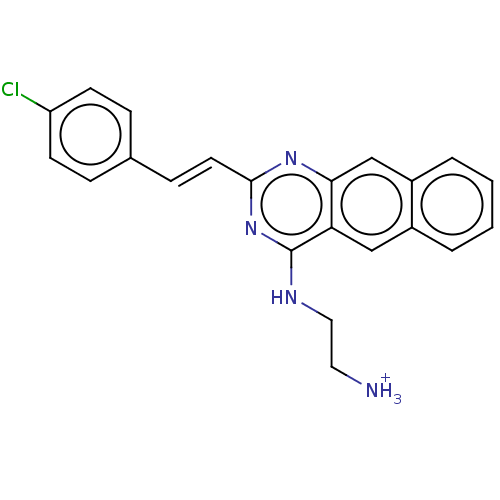
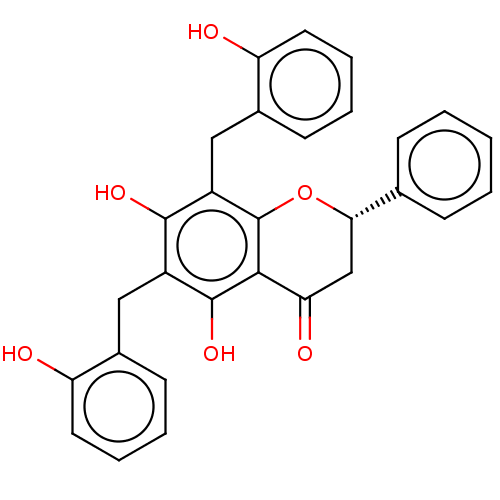
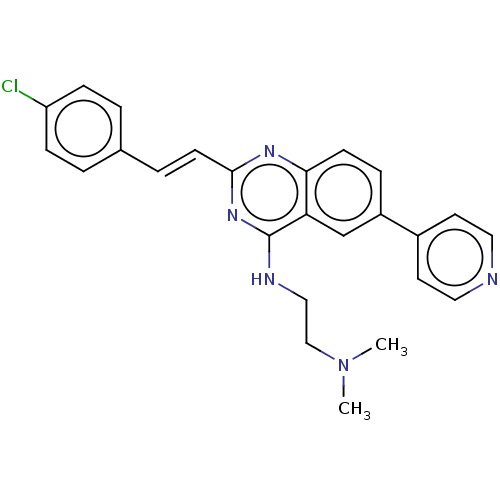
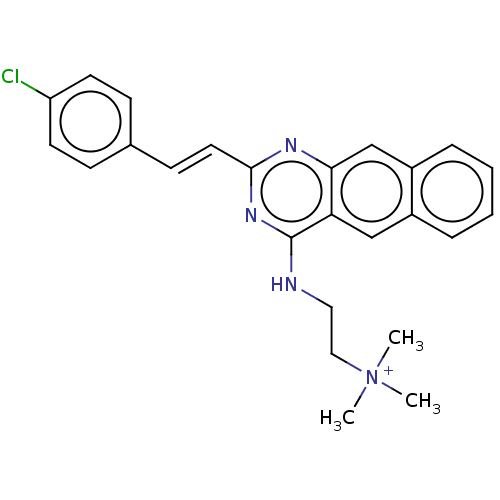
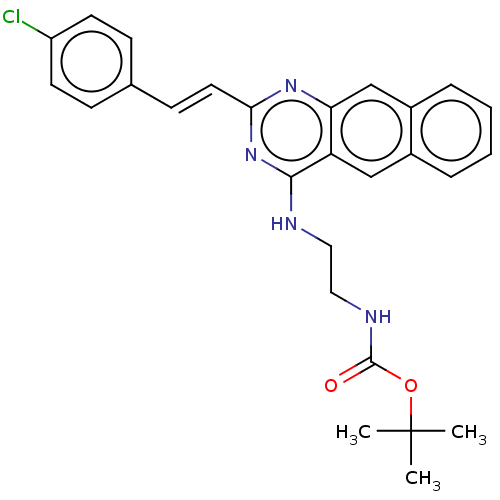
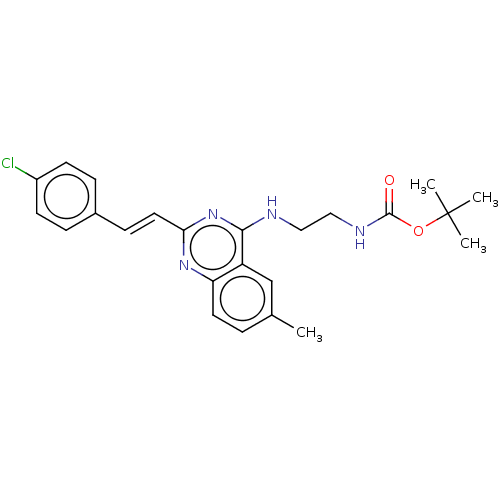
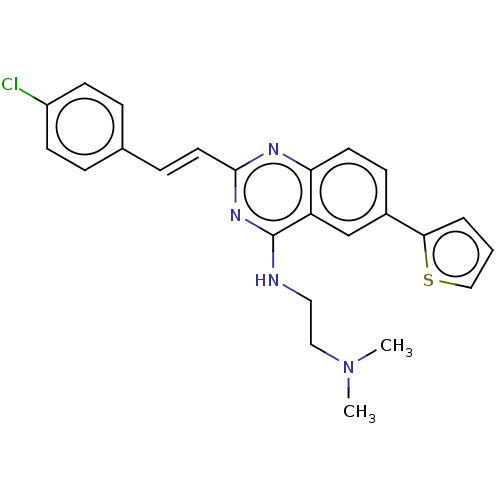
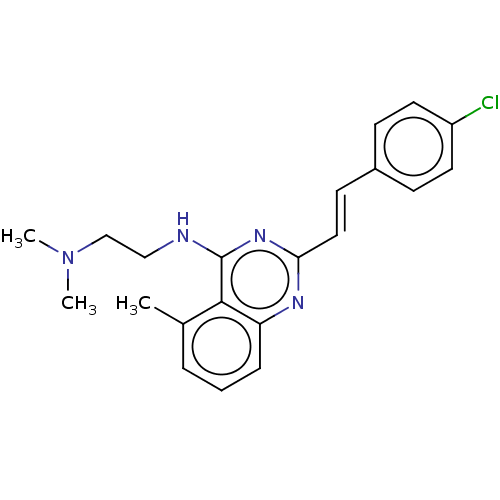
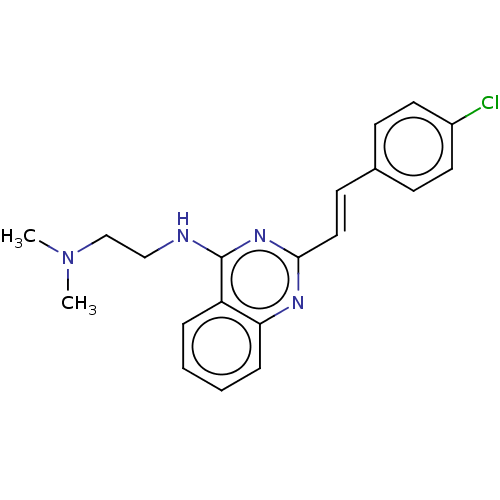
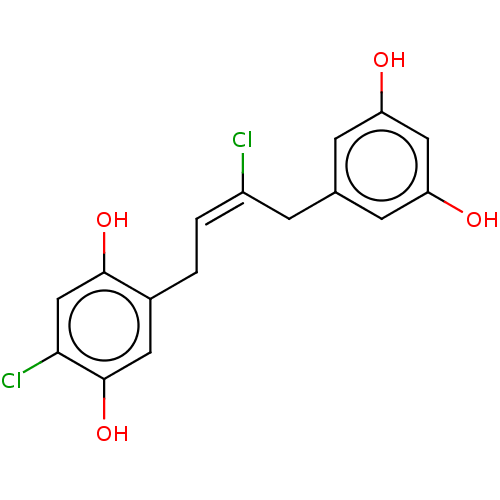
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
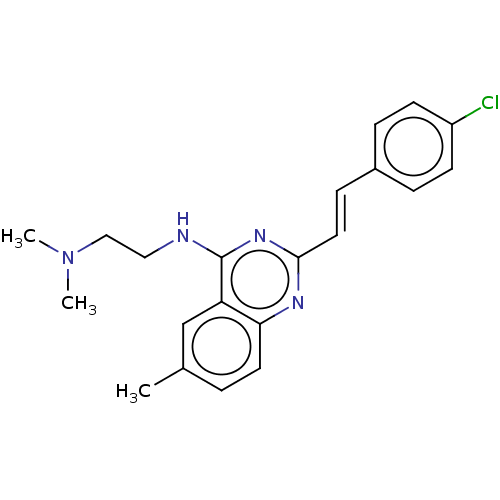
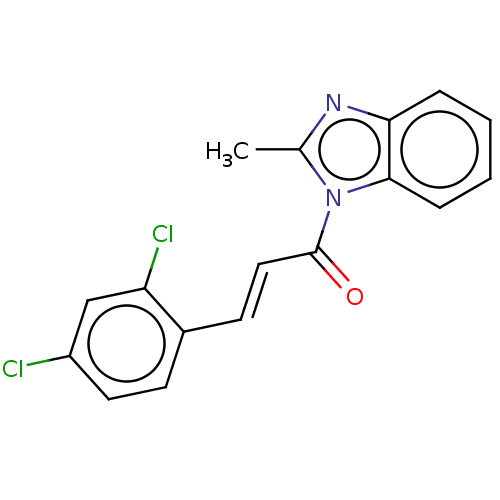
Affinity DataKd: 23nMAssay Description:Displacement of bis-ANS from Escherichia coli FtsZMore data for this Ligand-Target Pair
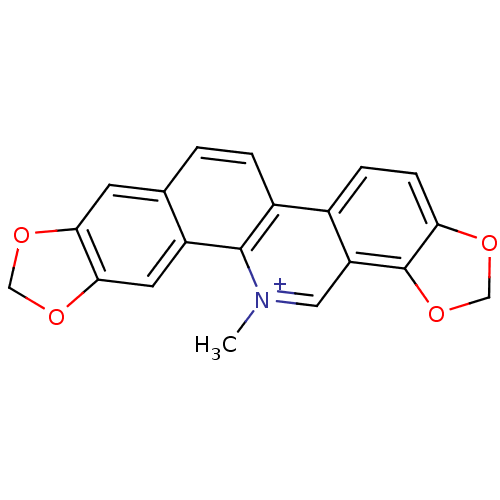
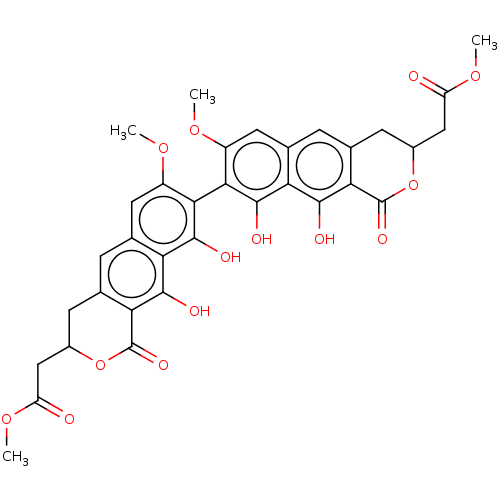
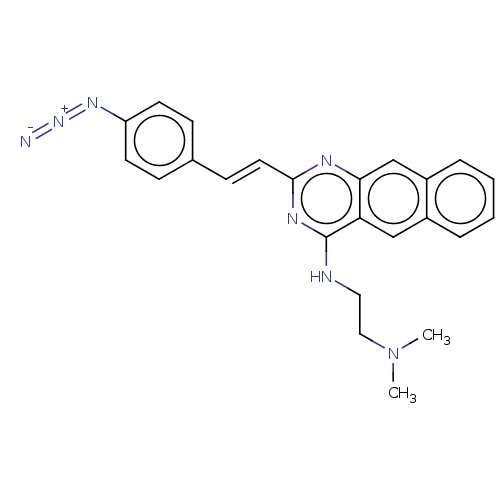
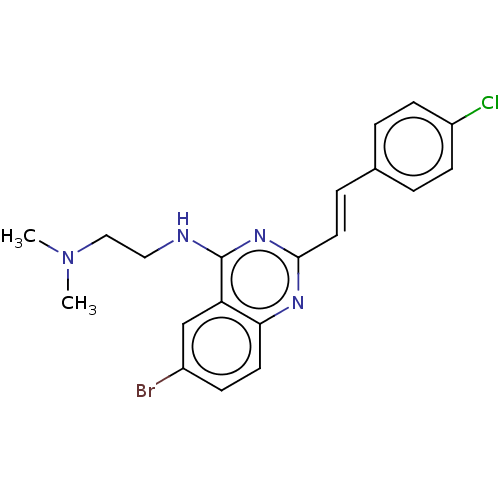
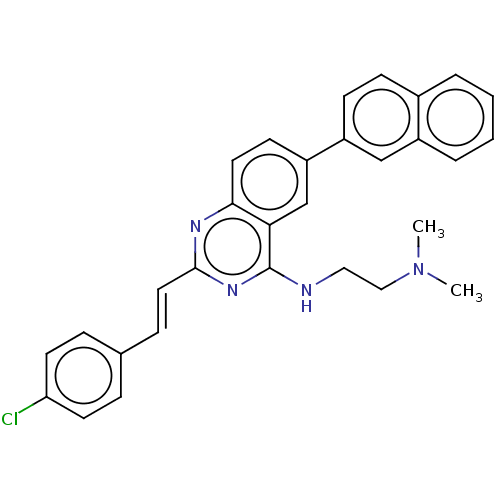
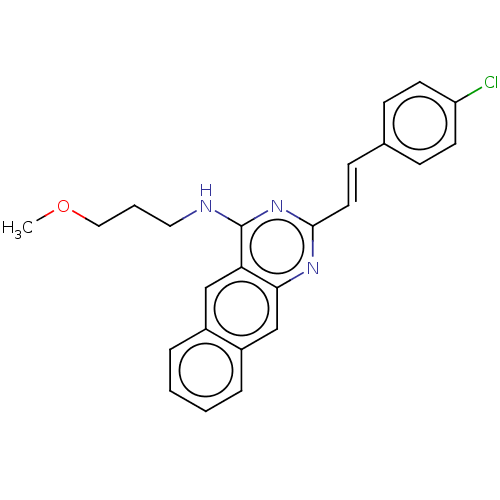
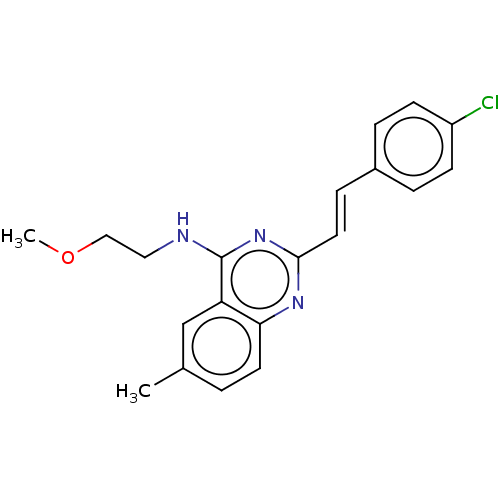
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
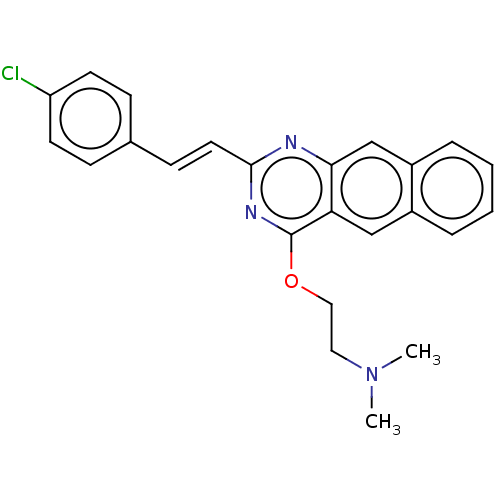
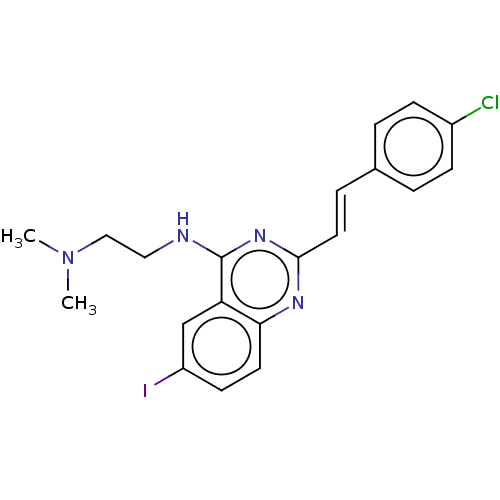
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of Escherichia coli recombinant FTsZ incubated for 10 mins in presence of GTP by spectrofluorimeter analysisMore data for this Ligand-Target Pair
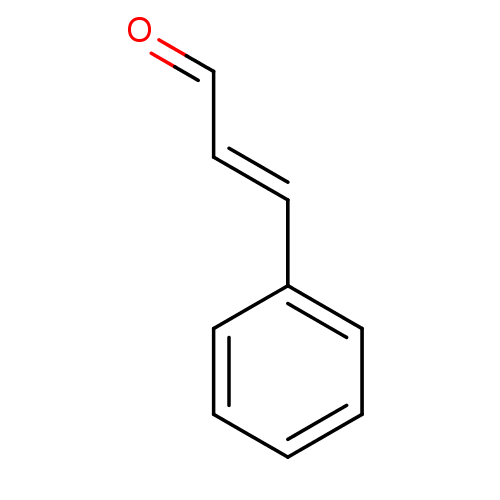
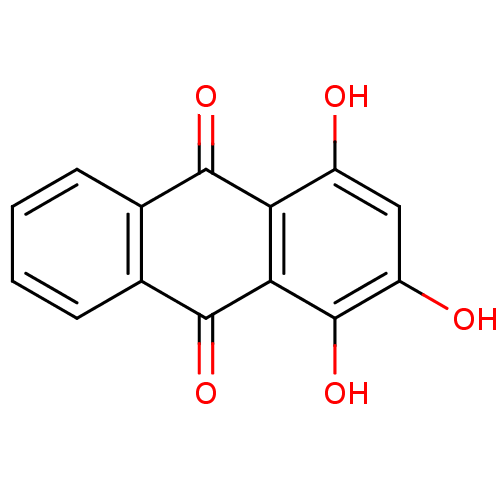
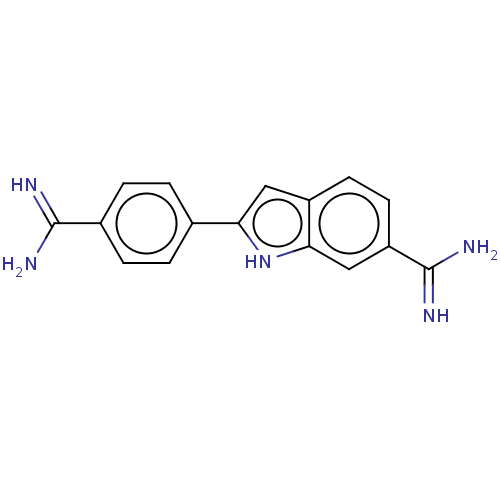
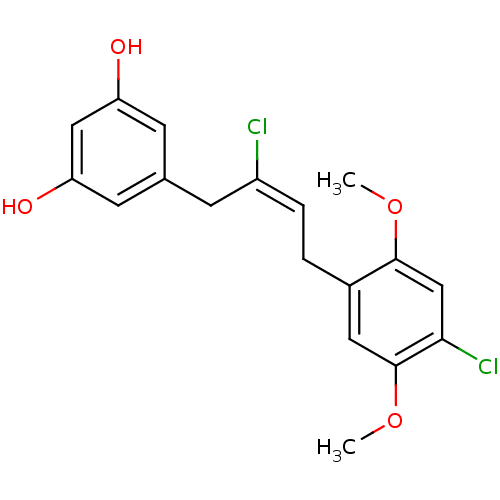
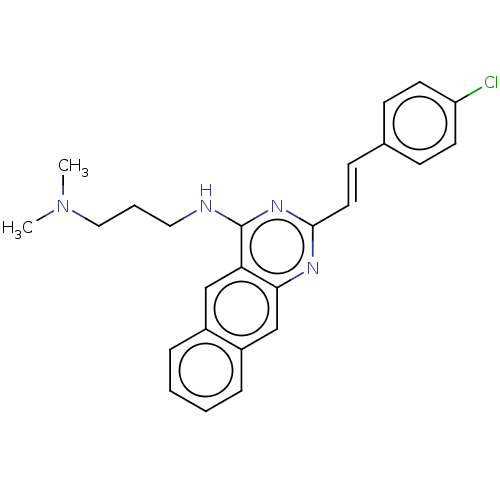
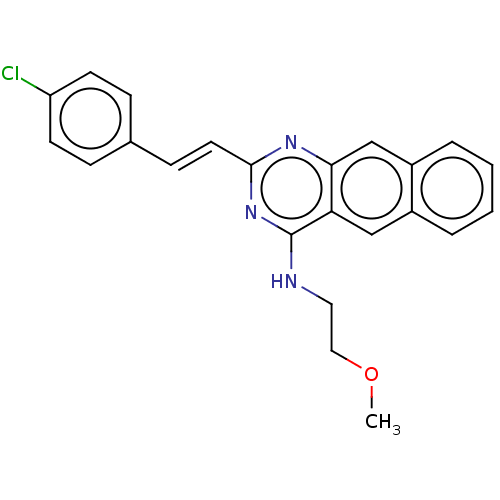
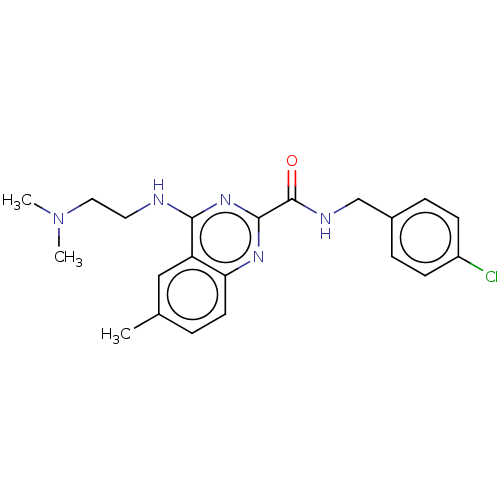
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
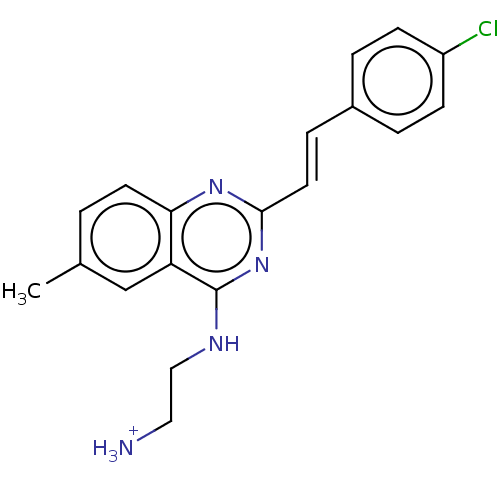
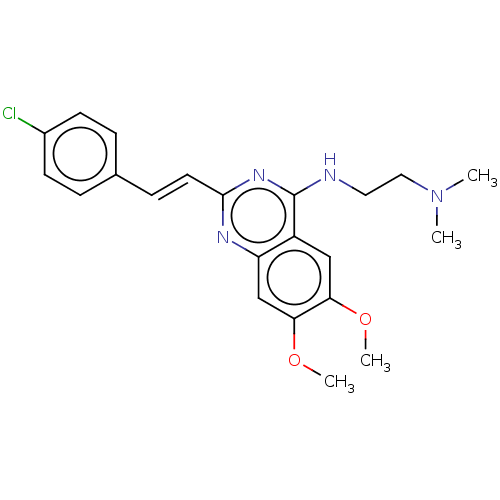
Affinity DataKd: 3.80E+3nMAssay Description:Binding affinity to Escherichia coli FtsZ at 25 degC by fluorescence anisotropy assayMore data for this Ligand-Target Pair
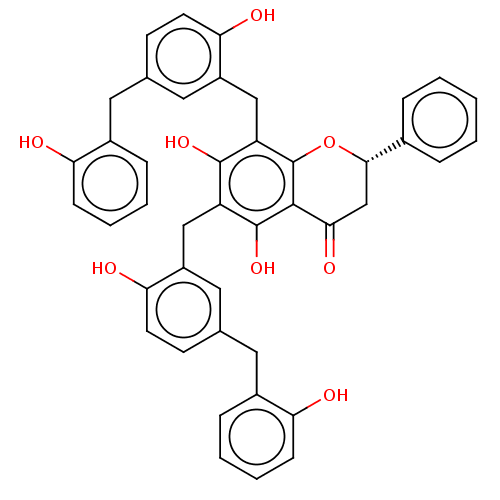
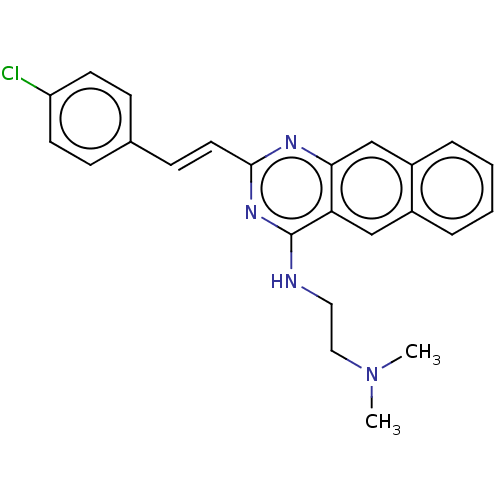
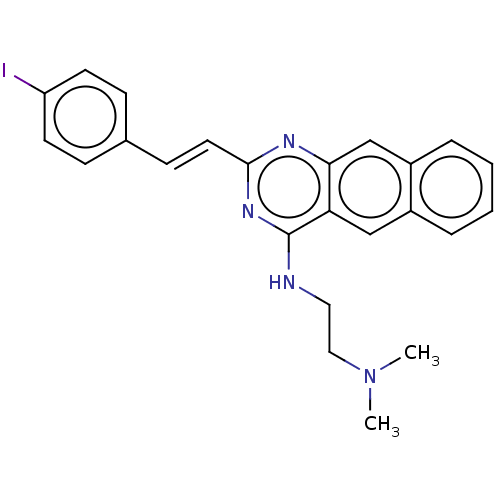
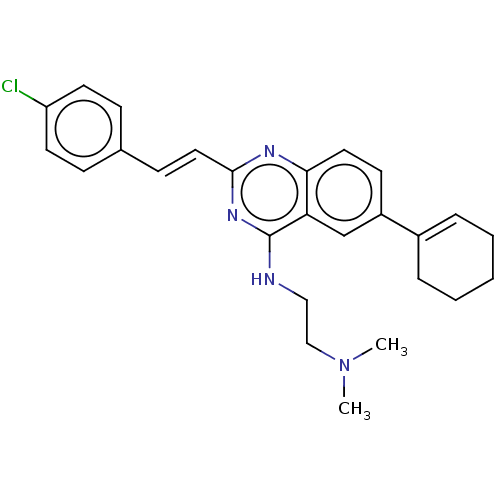
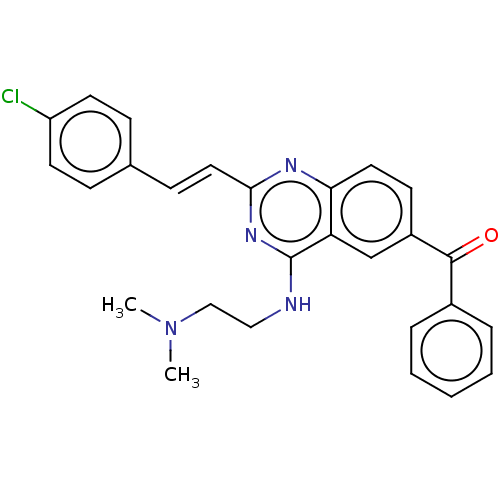
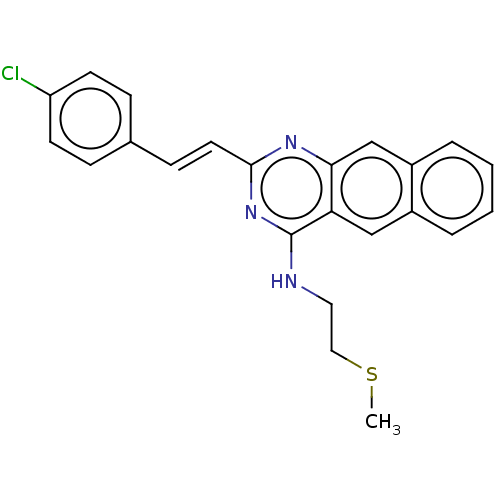
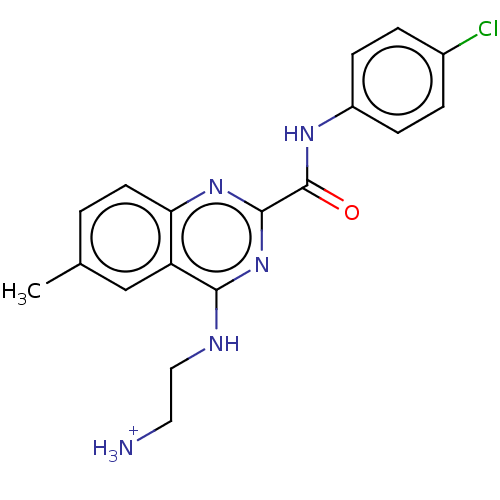
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
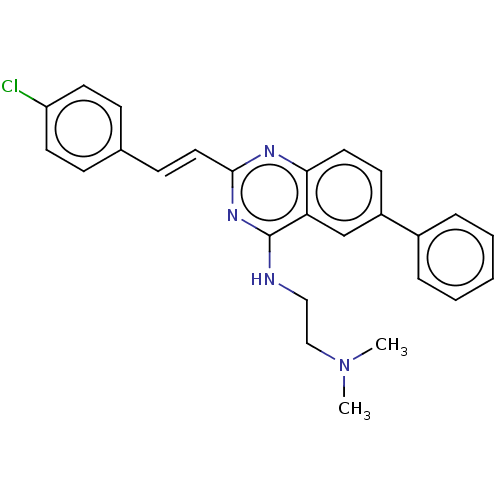
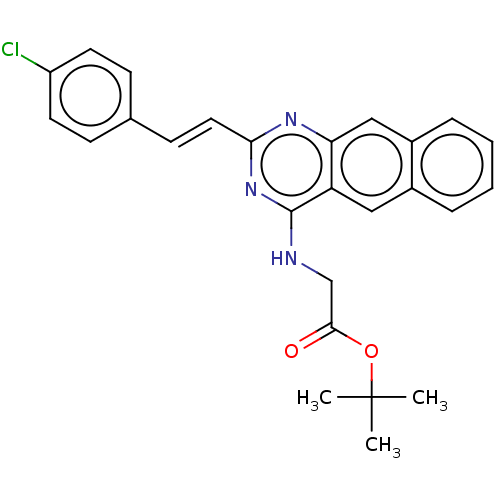
Affinity DataIC50: 5.81E+3nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activityMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 6.86E+3nMAssay Description:Inhibition of Escherichia coli FtsZ polymerizationMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 8.30E+3nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activityMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 9.00E+3nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 9.87E+3nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activityMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of recombinant Escherichia coli K12 FtsZ assembly by light-scattering assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.05E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activityMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKd: 1.10E+4nMAssay Description:Binding affinity to Escherichia coli BL21 FtsZ assessed as dissociation constant of hydrophobic probe ANS-FtsZ complex fluorescence quenching incubat...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of Escherichia coli FtsZMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.23E+4nMAssay Description:Inhibition of Escherichia coli FtsZ polymerizationMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.25E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activityMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.38E+4nMAssay Description:Inhibition of FtsZ in Escherichia coli assessed as disruption of Z ring formationMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.60E+4nMAssay Description:Inhibition of recombinant Escherichia coli K12 FtsZ GTPase activity by malachite green sodium molybdate assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKd: 1.66E+4nMAssay Description:Binding affinity to Escherichia coli FtsZ by fluorescence anisotrophyMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 2.40E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 2.40E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 2.60E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 2.70E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 2.80E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 3.60E+4nMpH: 6.5 T: 2°CAssay Description:GTPase assays were performed in 50 mM MES, pH 6.5, 50 mM KCl, 5 mM MgCl2 in the presence of 2 uM FtsZ in the presence of varying concentrations of co...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 3.70E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activityMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 4.90E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 5.20E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 5.80E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 6.50E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 8.50E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 9.05E+4nMAssay Description:Inhibition of Escherichia coli ATCC 25922 recombinant N-His6-tagged FtsZ polymerization expressed in Escherichia coli BL21(DE3) incubated for 15 mins...More data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 9.50E+4nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.28E+5nMAssay Description:Inhibition of Escherichia coli FtsZ GTPase activity after 5 mins by enzyme-coupled inhibition assayMore data for this Ligand-Target Pair
TargetCell division protein FtsZ(Escherichia coli (strain K12))
University of Wisconsin-Madison
Curated by ChEMBL
University of Wisconsin-Madison
Curated by ChEMBL
Affinity DataIC50: 1.60E+5nMpH: 6.5 T: 2°CAssay Description:GTPase assays were performed in 50 mM MES, pH 6.5, 50 mM KCl, 5 mM MgCl2 in the presence of 2 uM FtsZ in the presence of varying concentrations of co...More data for this Ligand-Target Pair