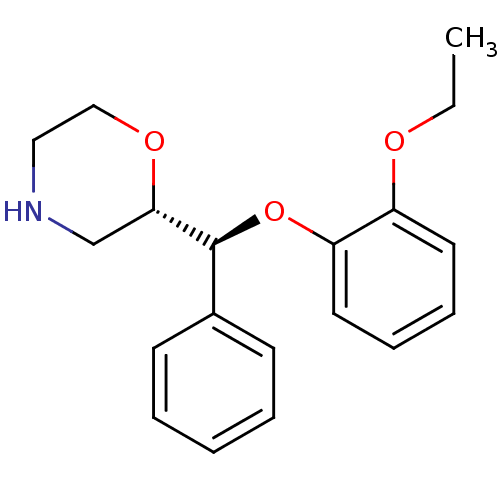
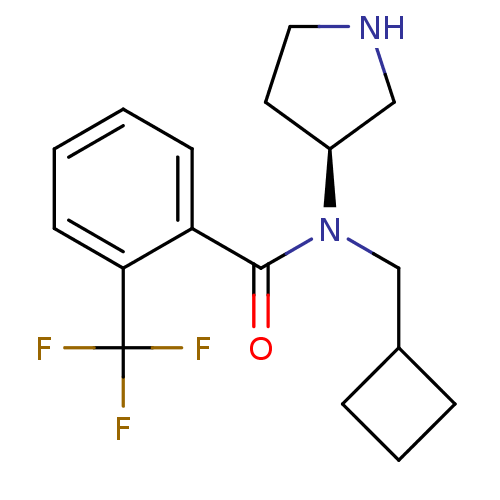
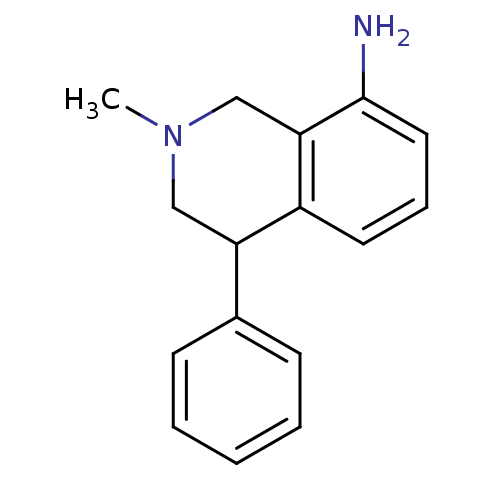
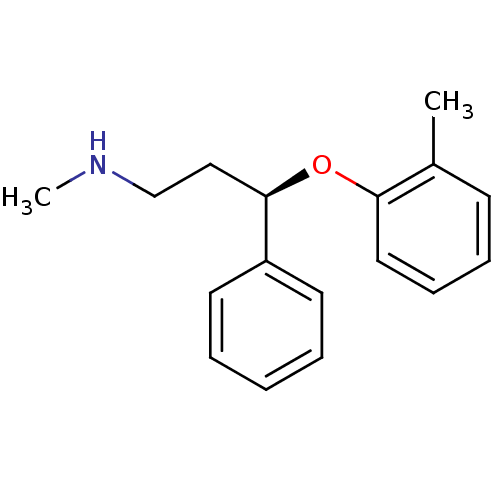
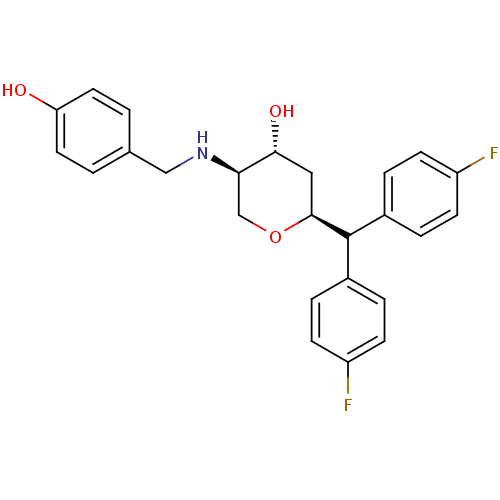
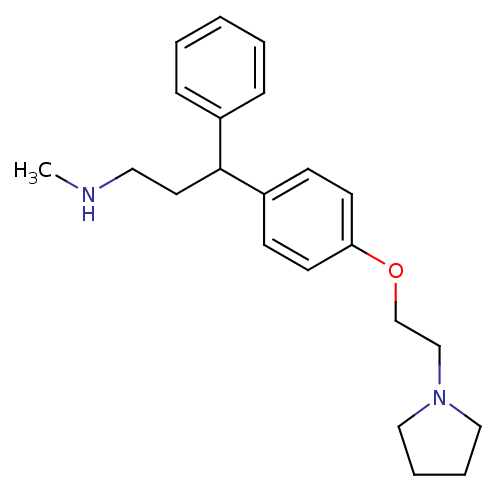
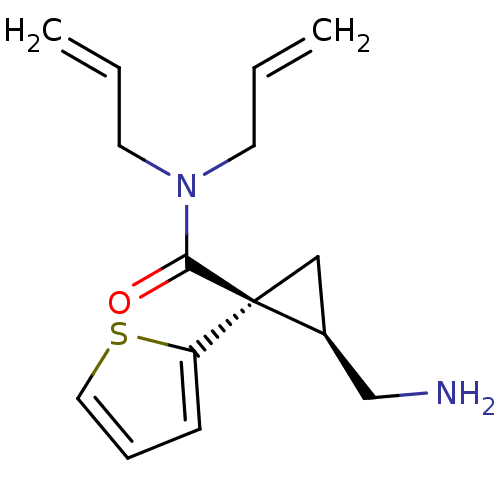
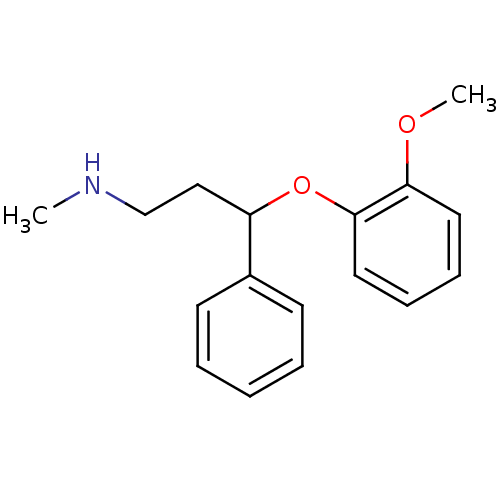
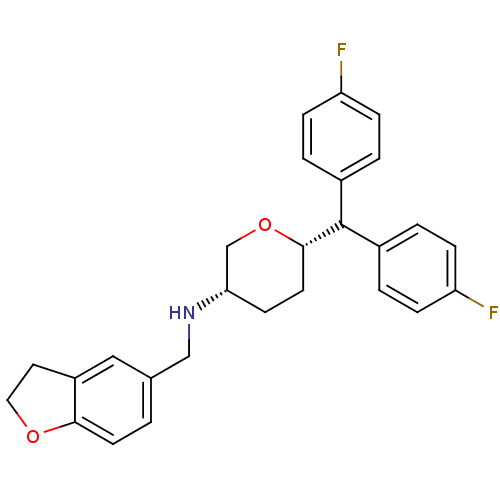
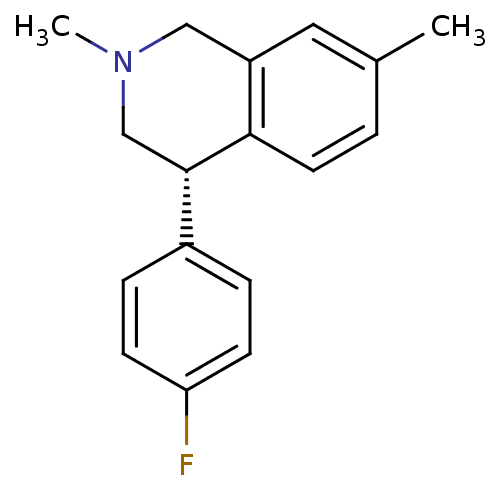
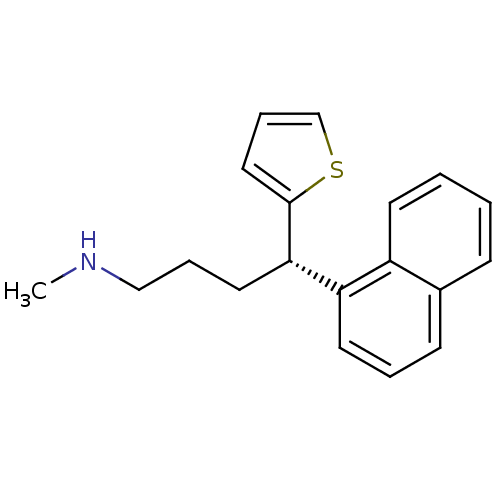
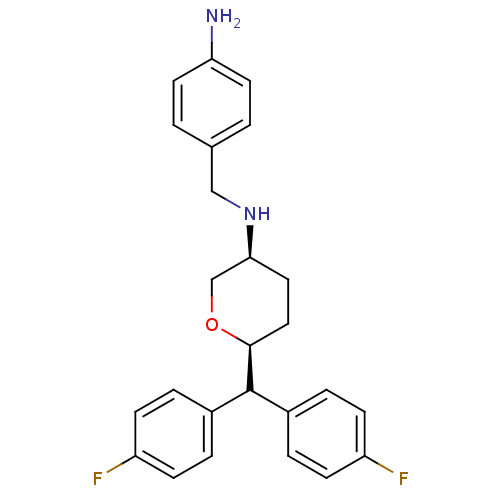
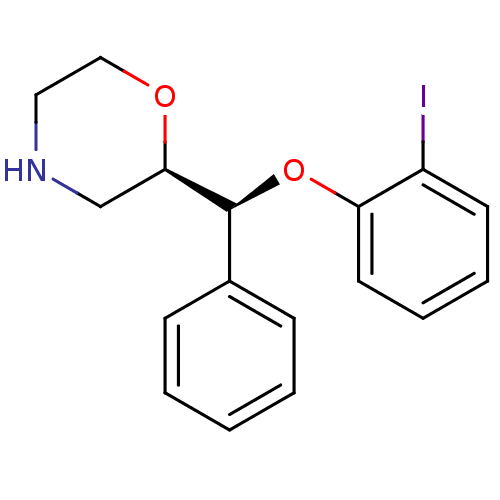
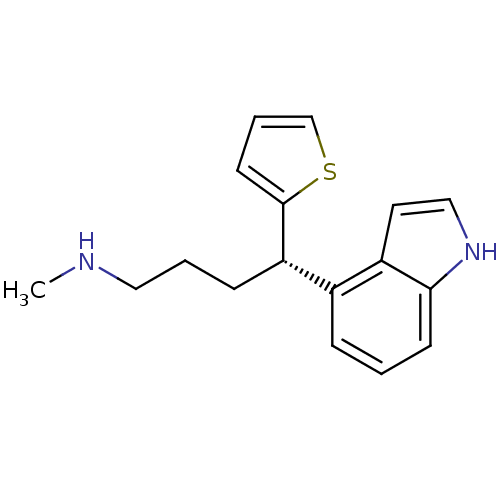
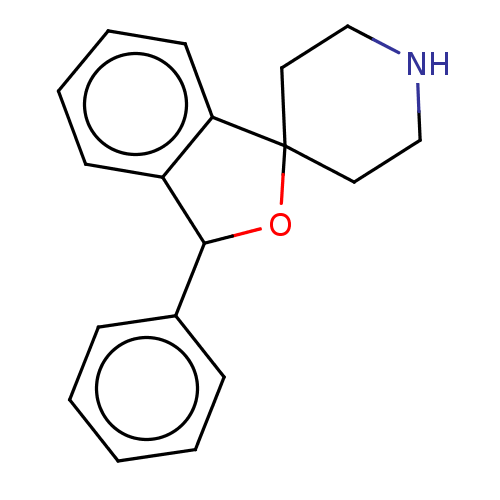
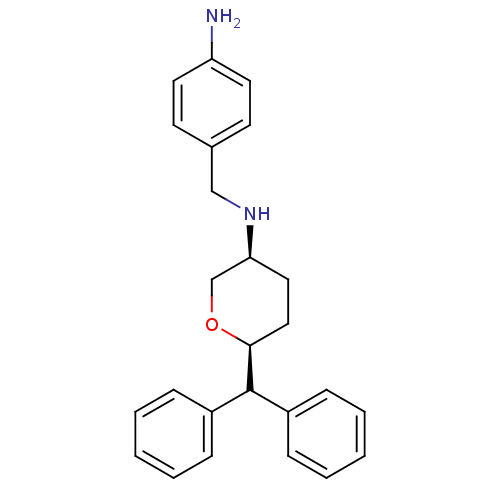
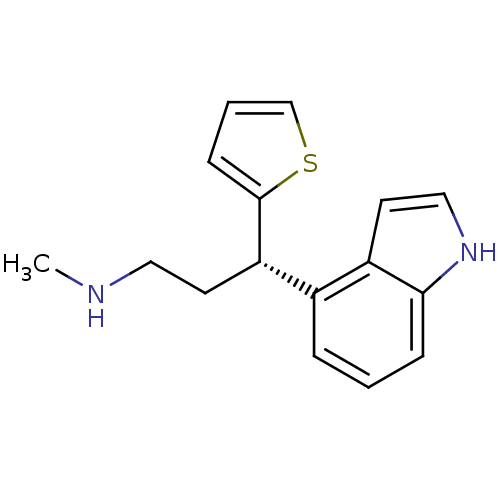
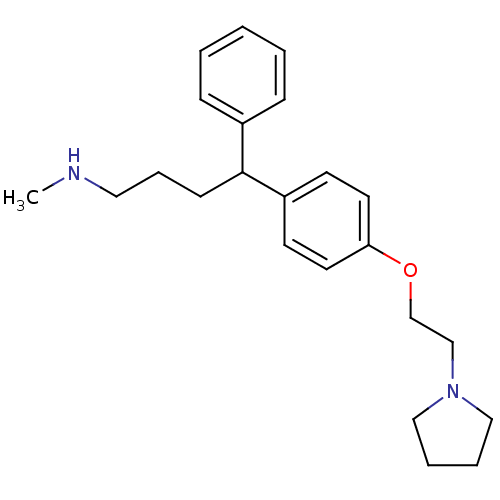
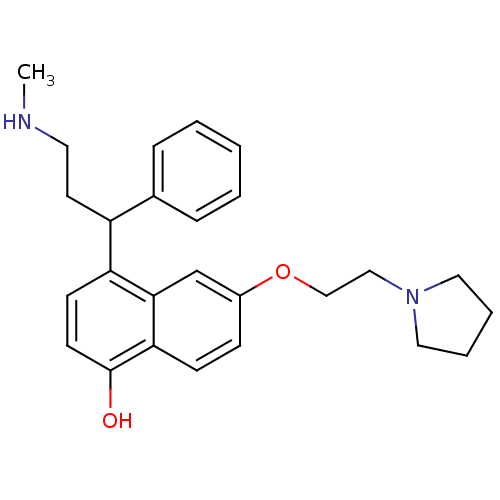
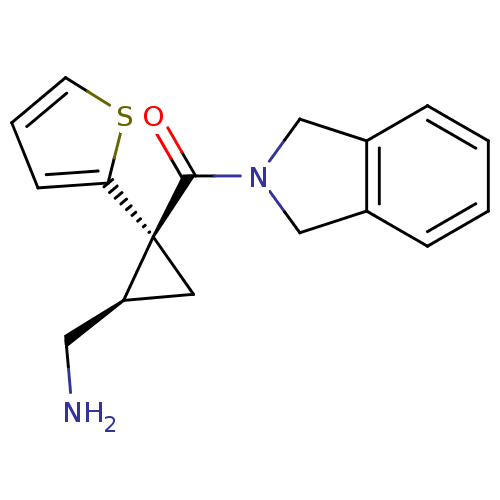
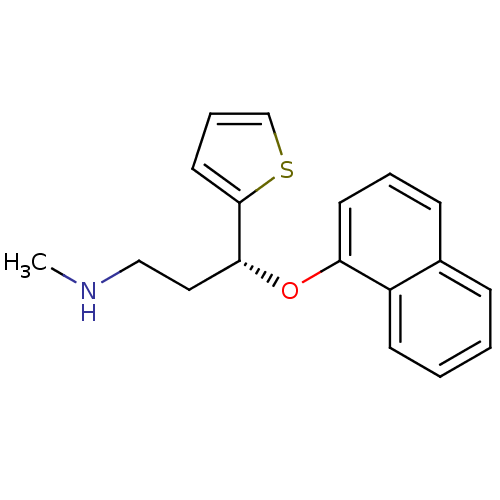
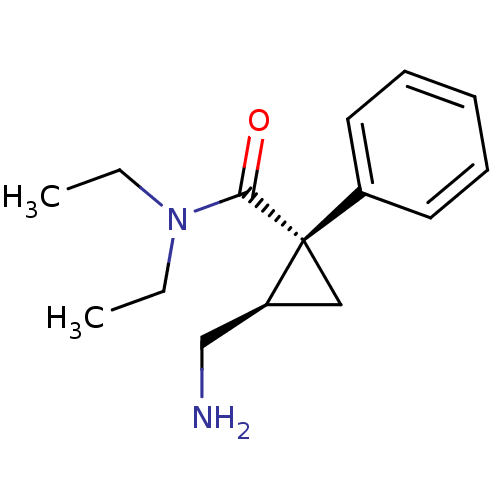
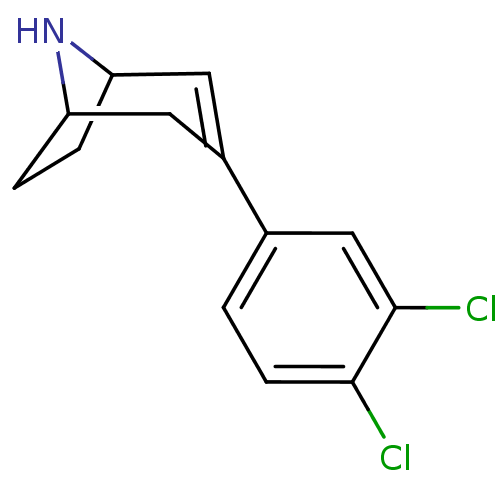
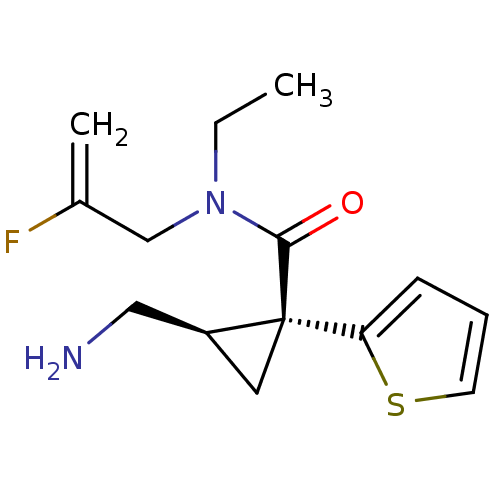
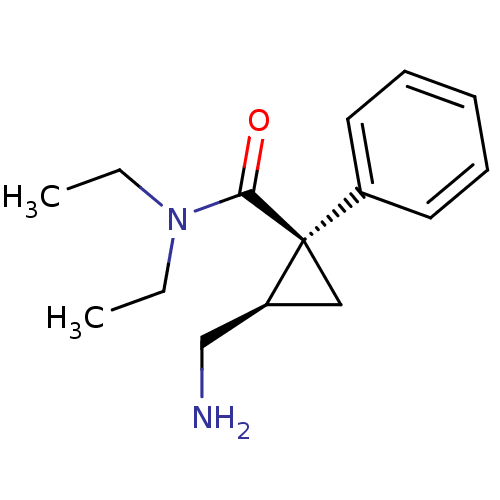
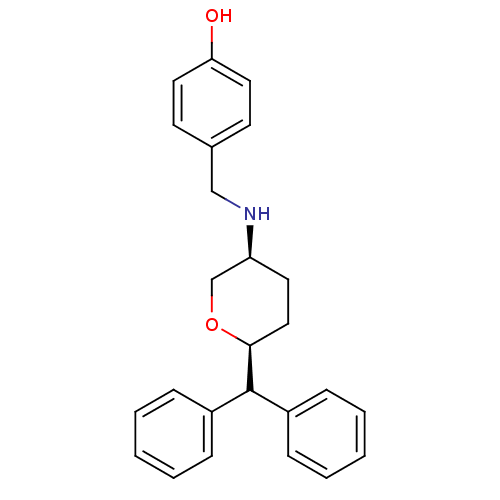
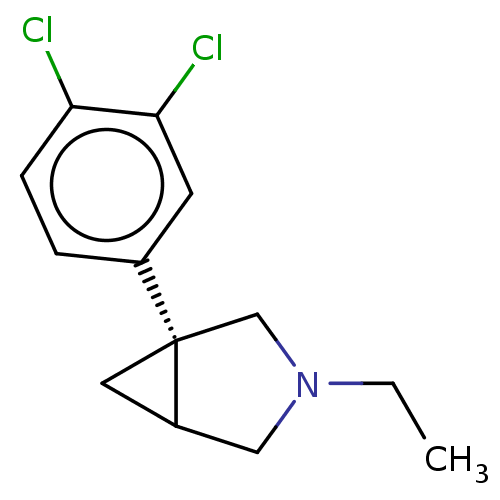
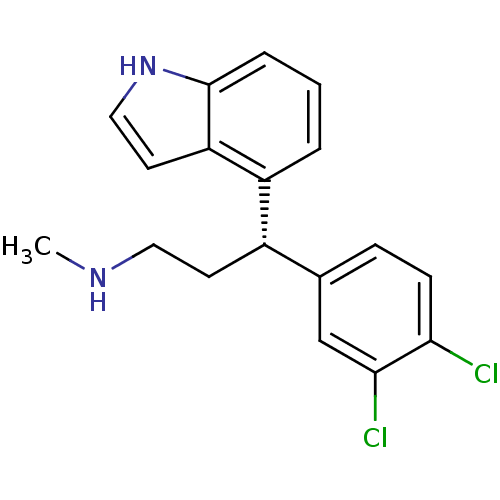
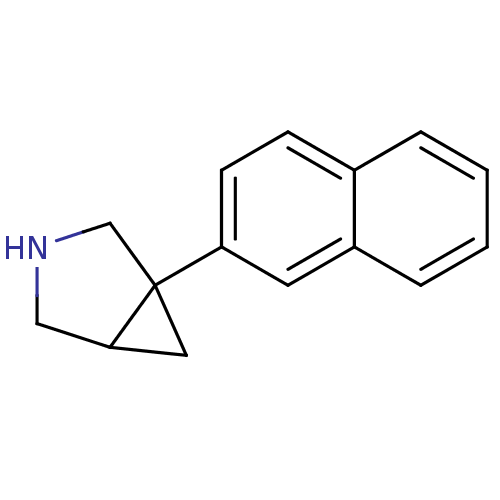
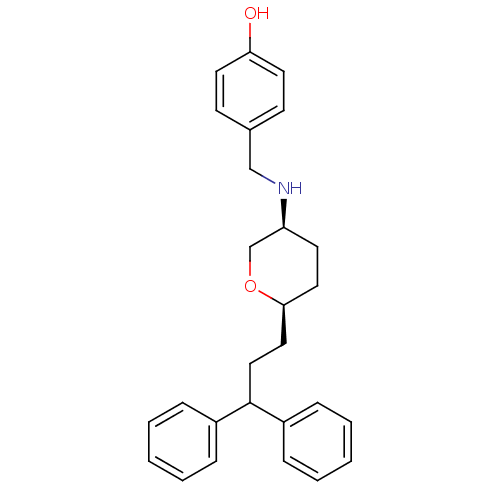
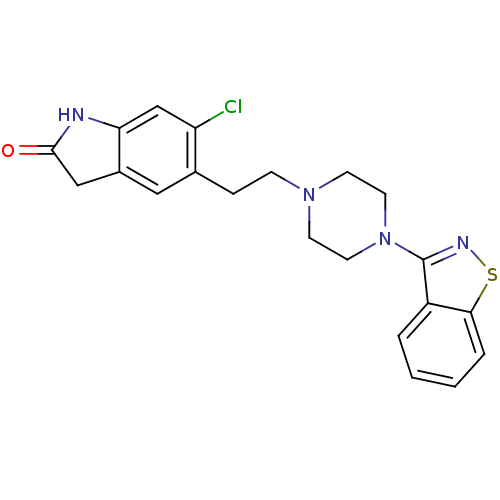
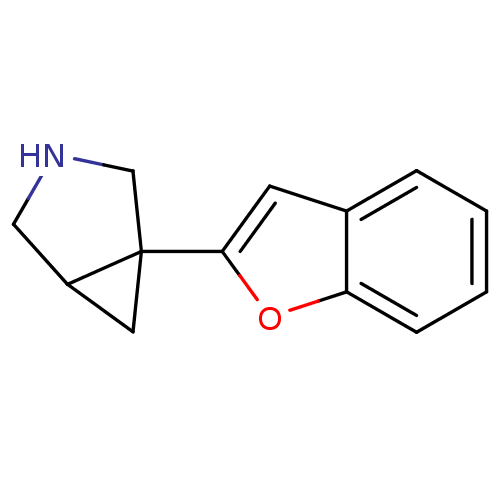
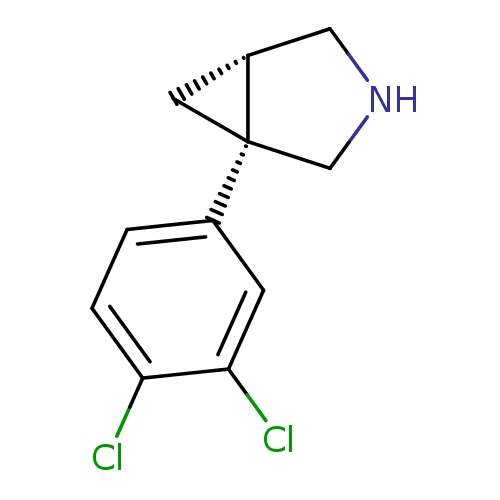
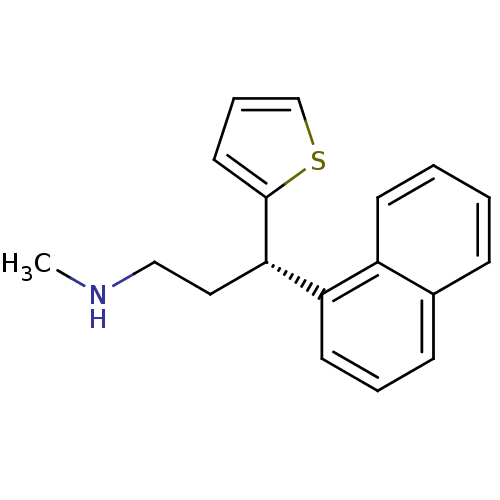
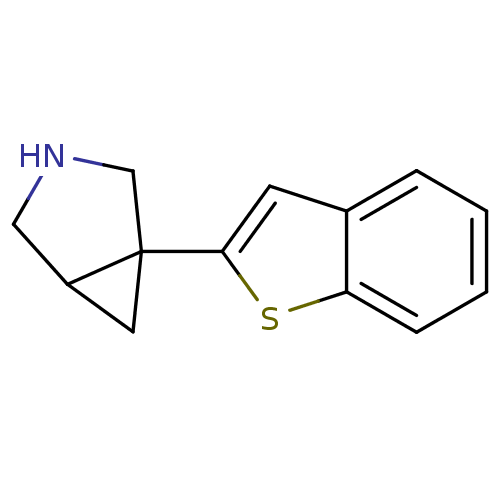
Affinity DataKi: 0.135nMAssay Description:Displacement of [N-methyl-3H]nisoxetine from rat hippocampus NET by filtration binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.690nMAssay Description:Displacement of [3H]-dopamine from rat cerebral cortex NET after 7 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.780nMAssay Description:Displacement of [3H]nisoxetine from rat NET in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 0.840nMAssay Description:Displacement of [3H]nisoxetine from NAT in Sprague-Dawley rat brain after 4 hrs by liquid scintillation analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:Inhibitory concentration against human Adenosine A3 receptor expressed in HEK293 cells using 0.1 nM [3H]AB-MECAMore data for this Ligand-Target Pair
Affinity DataKi: 2.51nMAssay Description:Displacement of [3H]nisoxetine from NET in rat hippocampus after 2 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of [3H]nisoxetine from rat NET in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 3.10nMAssay Description:Inhibition of norepinephrine uptake at NET in rat brain synaptosomeMore data for this Ligand-Target Pair
Affinity DataKi: 3.90nMAssay Description:Binding affinity to rat NET expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Inhibition of NET in rat synaptosomal membraneMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]nisoxetine from Sprague-dawley rat NET by liquid scintillation spectrophotometryMore data for this Ligand-Target Pair
Affinity DataKi: 4.20nMAssay Description:Displacement of [3H]-dopamine from rat cerebral cortex NET after 7 minsMore data for this Ligand-Target Pair
Affinity DataKi: 4.30nMAssay Description:Displacement of [3H]nisoxetine from rat NET in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 4.60nMAssay Description:Displacement of Levo[ring-2,5,6-3H]norepinephrine from rat synaptosomal NET expressed in HEK293 cells by microplate scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Displacement of [3H]nisoxetine from Sprague-dawley rat NET by liquid scintillation spectrophotometryMore data for this Ligand-Target Pair
Affinity DataKi: 6.10nMAssay Description:Displacement of [3H]-dopamine from rat cerebral cortex NET after 7 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 6.40nMAssay Description:Displacement of Levo[ring-2,5,6-3H]norepinephrine from rat synaptosomal NET expressed in HEK293 cells by microplate scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 7.90nMAssay Description:Displacement of [3H]nisoxetine from Sprague-dawley rat NET by liquid scintillation spectrophotometryMore data for this Ligand-Target Pair
Affinity DataKi: 8.40nMAssay Description:Displacement of [3H]-dopamine from rat cerebral cortex NET after 7 minsMore data for this Ligand-Target Pair
Affinity DataKi: 8.40nMAssay Description:Displacement of [3H]nisoxetine from NAT in Sprague-Dawley rat brain after 4 hrs by liquid scintillation analysisMore data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Displacement of [3H]nisoxetine from Sprague-dawley rat NET by liquid scintillation spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of NET-mediated norepinephrine uptake in rat brain synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [3H]-dopamine from rat cerebral cortex NET after 7 minsMore data for this Ligand-Target Pair
Affinity DataKi: 13.8nMAssay Description:Displacement of [3H]nisoxetine from Sprague-dawley rat NET by liquid scintillation spectrophotometryMore data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Displacement of [3H]nisoxetine from rat NET in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Displacement of [3H]nisoxetine from rat NET in rat cerebral cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Displacement of Levo[ring-2,5,6-3H]norepinephrine from rat synaptosomal NET expressed in HEK293 cells by microplate scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Displacement of [3H]nisoxetine from Sprague-dawley rat NET by liquid scintillation spectrophotometryMore data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Displacement of [3H]nisoxetine from Sprague-dawley rat NET by liquid scintillation spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Displacement of Levo[ring-2,5,6-3H]norepinephrine from rat synaptosomal NET expressed in HEK293 cells by microplate scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Displacement of [3H]nisoxetine from NET in rat brainMore data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:Displacement of Levo[ring-2,5,6-3H]norepinephrine from rat synaptosomal NET expressed in HEK293 cells by microplate scintillation counterMore data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Inhibition of [3H]norepinephrine reuptake at NET in rat brain synaptosome by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 26nMAssay Description:Displacement of [3H]-dopamine from rat cerebral cortex NET after 7 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Inhibition of norepinephrine uptake at NET in rat brain synaptosomeMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Displacement of Levo[ring-2,5,6-3H]norepinephrine from rat synaptosomal NET expressed in HEK293 cells by microplate scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 31nMAssay Description:Displacement of [3H]-dopamine from rat cerebral cortex NET after 7 minsMore data for this Ligand-Target Pair
Affinity DataKi: 32nMAssay Description:Binding assay using NET, DAT and SERT enzyme.More data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Inhibition of norepinephrine uptake at NET in rat brain synaptosomeMore data for this Ligand-Target Pair
Affinity DataKi: 35.4nMAssay Description:Displacement of [3H]nisoxetine from Sprague-dawley rat NET by liquid scintillation spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Inhibition of norepinephrine uptake at NET in rat brain synaptosomeMore data for this Ligand-Target Pair
Affinity DataKi: 38nMAssay Description:Displacement of [3H]-dopamine from rat cerebral cortex NET after 7 minsMore data for this Ligand-Target Pair
Affinity DataKi: 48nMAssay Description:Binding affinity to rat NETMore data for this Ligand-Target Pair
Affinity DataIC50: 48nMAssay Description:Inhibition of norepinephrine uptake at NET in rat brain synaptosomeMore data for this Ligand-Target Pair
Affinity DataIC50: 51nMAssay Description:Inhibition of norepinephrine uptake at NET in rat brain synaptosomeMore data for this Ligand-Target Pair
Affinity DataKi: 52.1nMAssay Description:Displacement of [3H]nisoxetine from Sprague-dawley rat NET by liquid scintillation spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 53nMAssay Description:Inhibition of norepinephrine uptake at NET in rat brain synaptosomeMore data for this Ligand-Target Pair
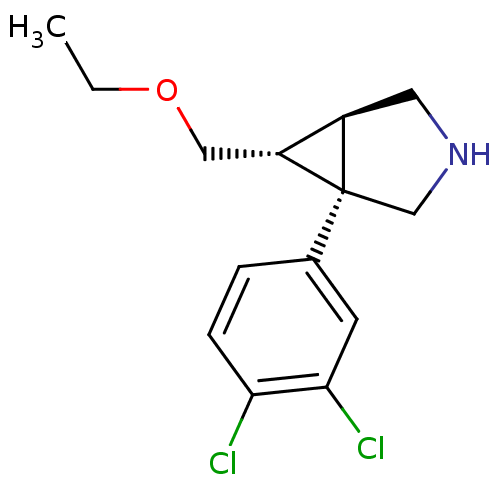
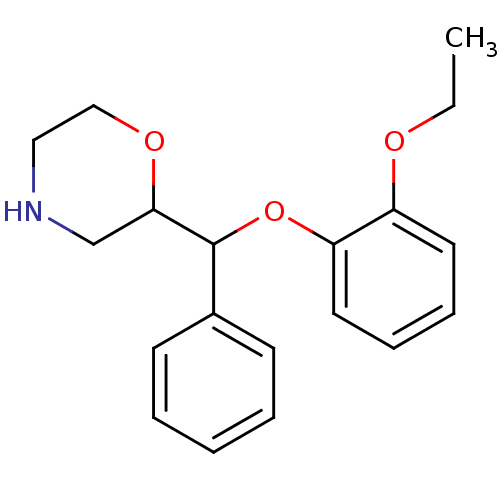

 3D Structure (crystal)
3D Structure (crystal)