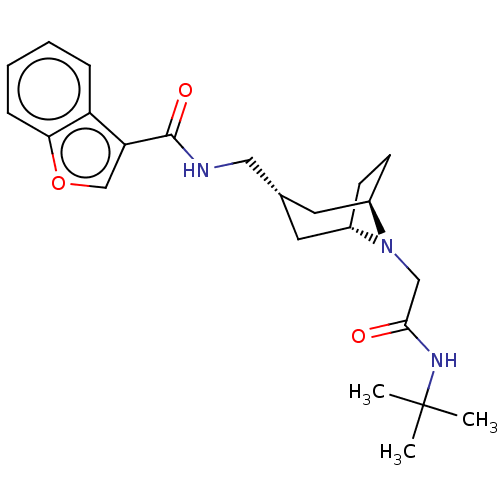
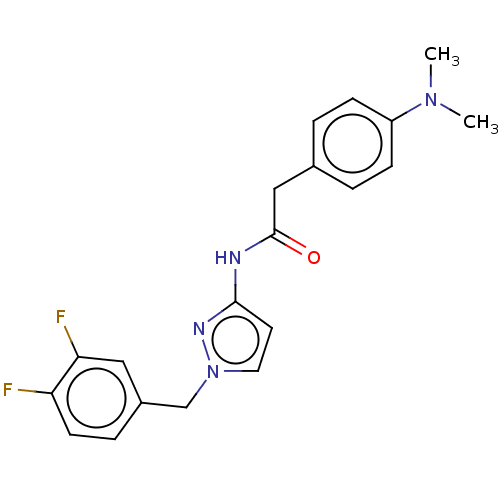
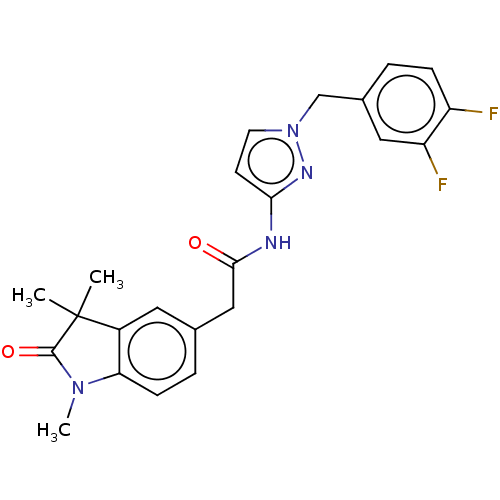
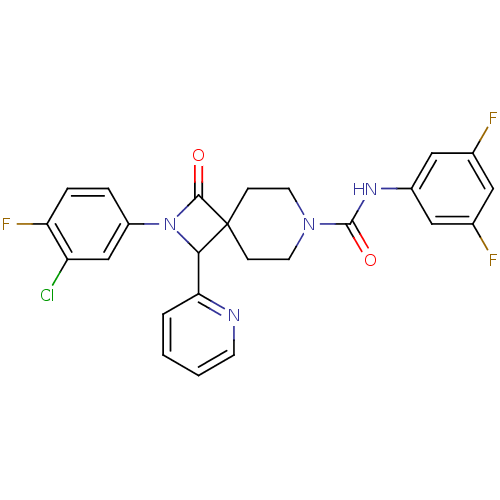
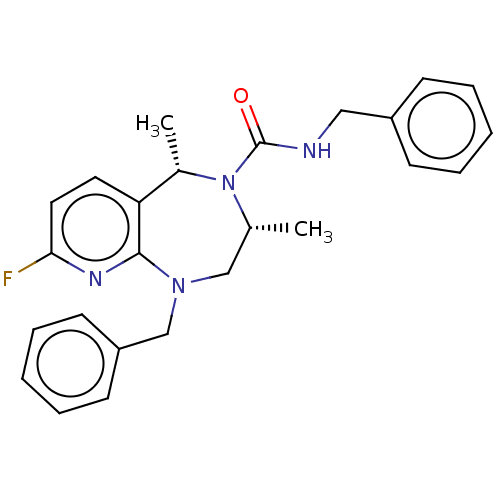
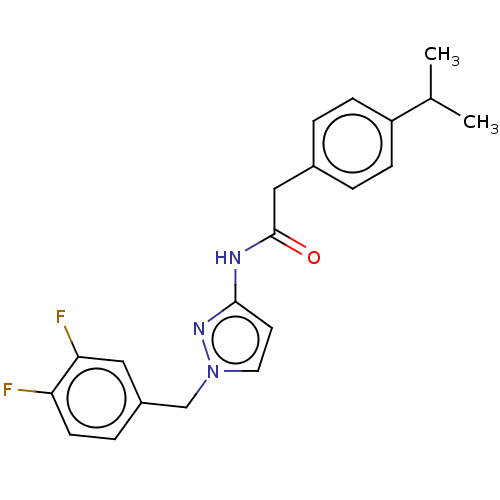
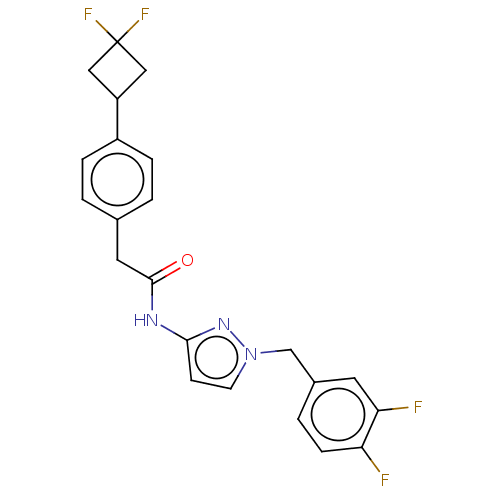
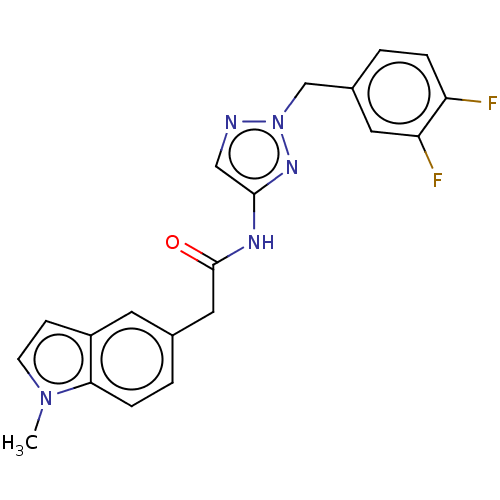
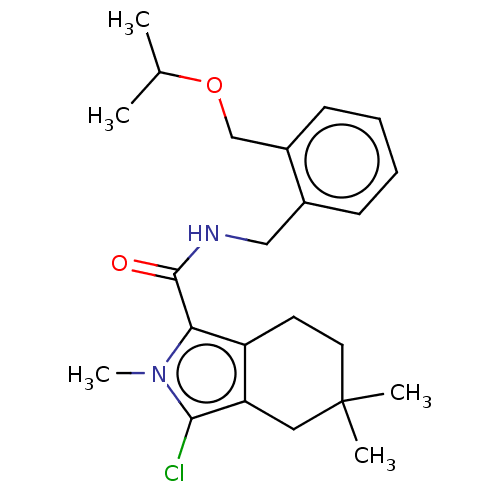
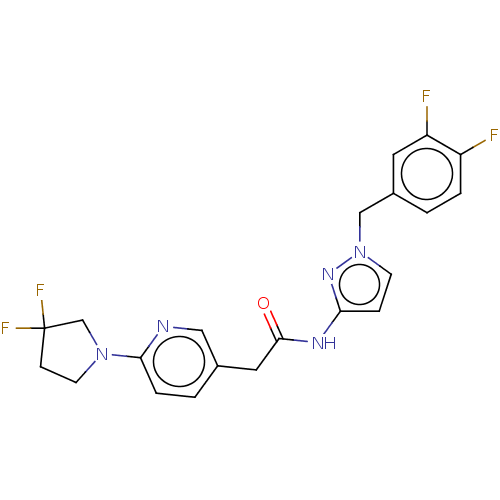
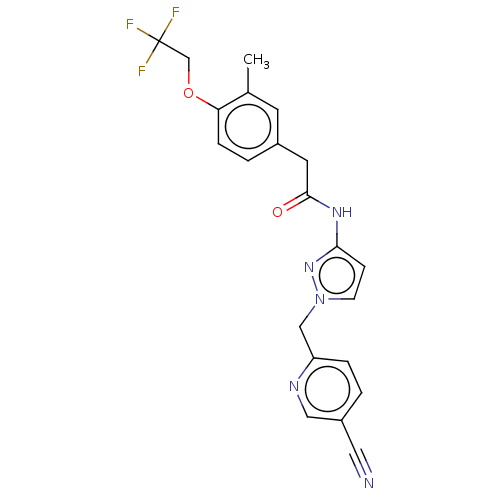
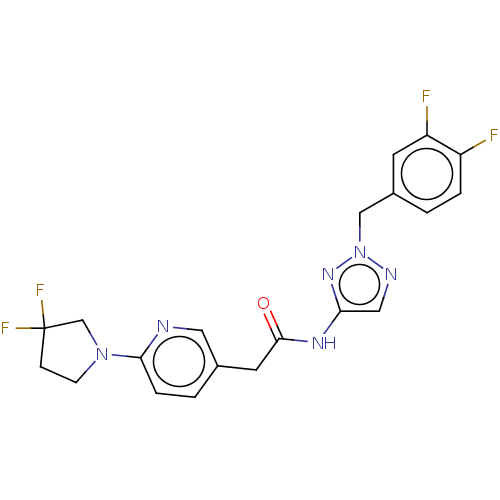
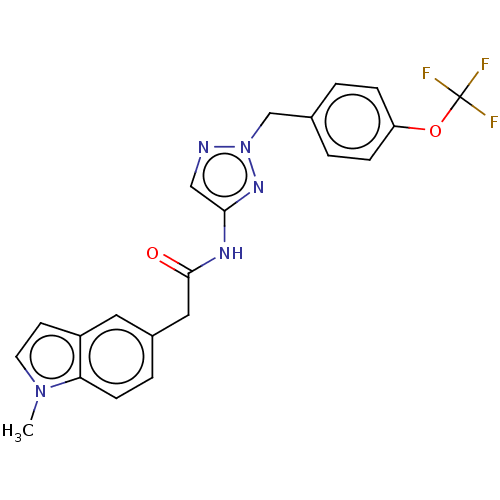
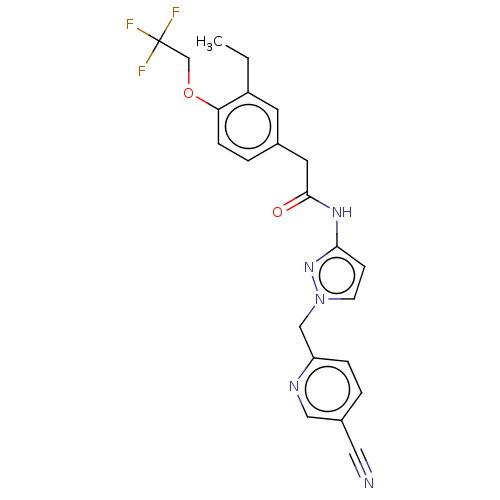
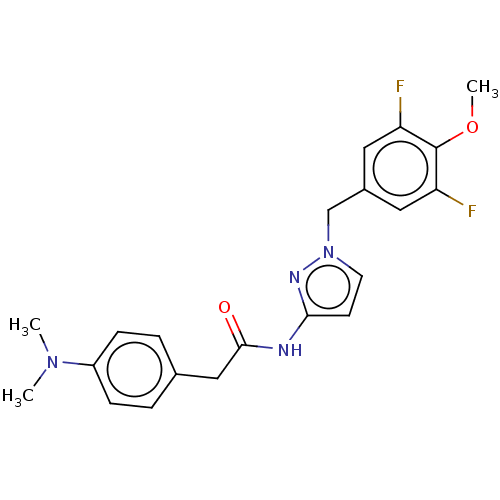
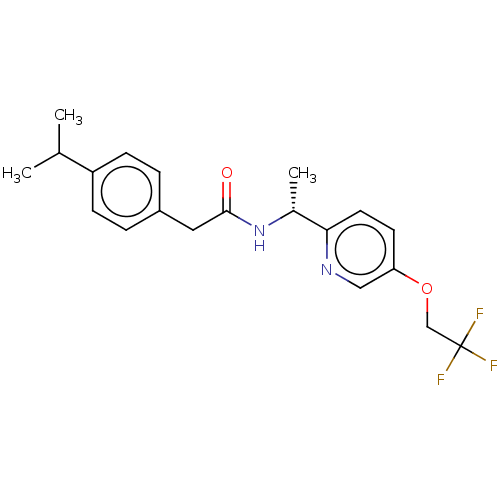
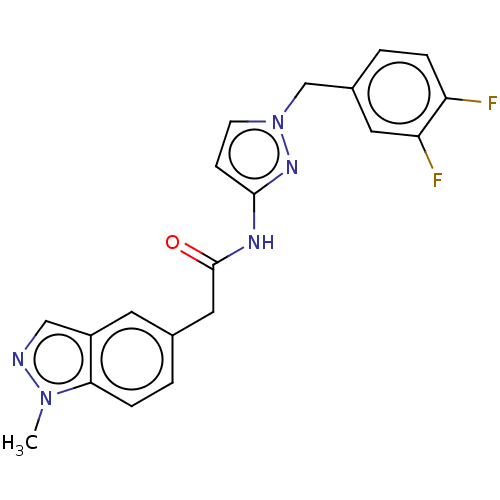
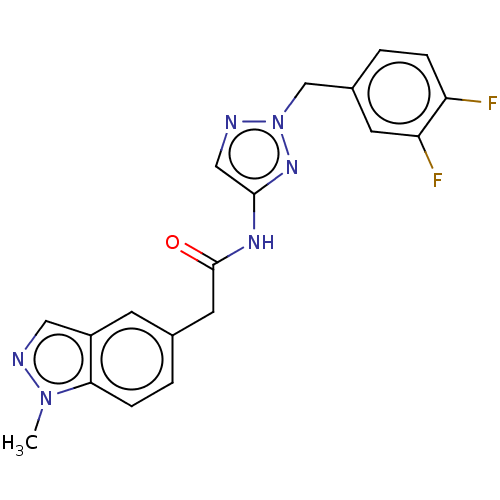
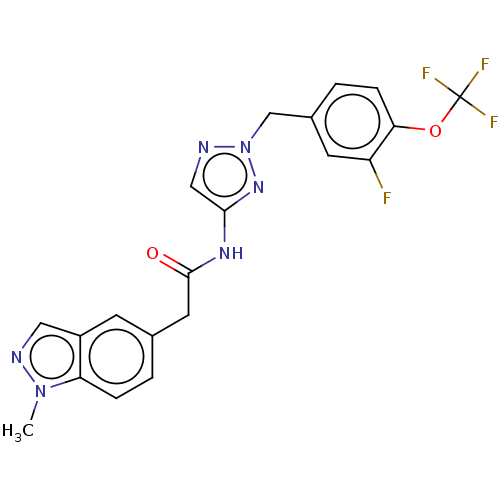
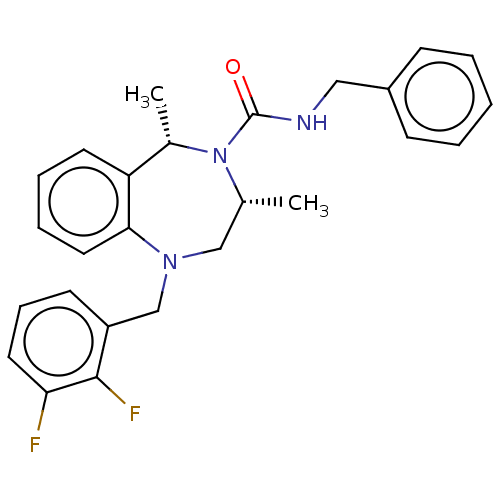
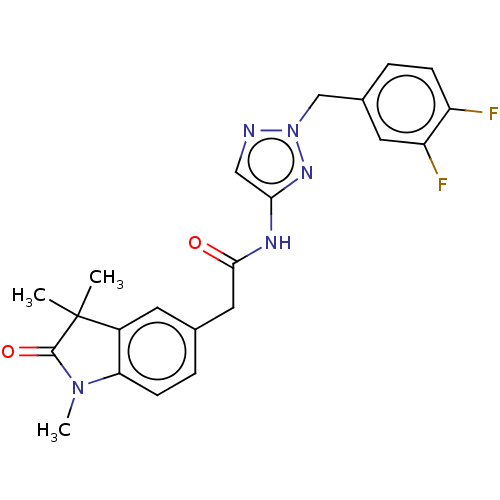
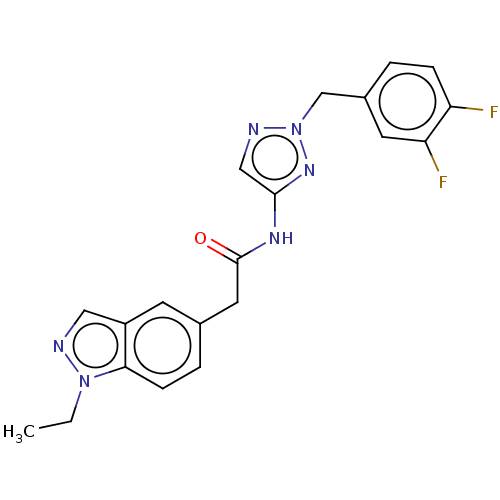
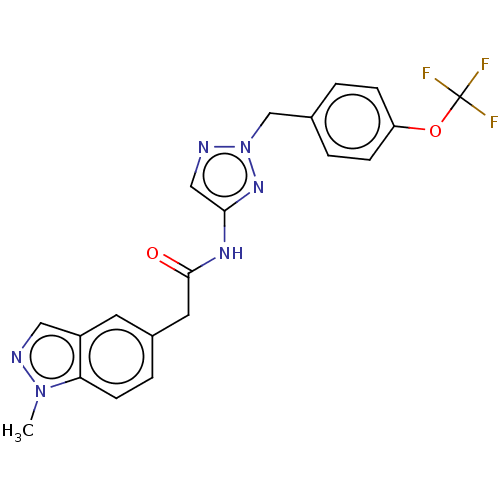
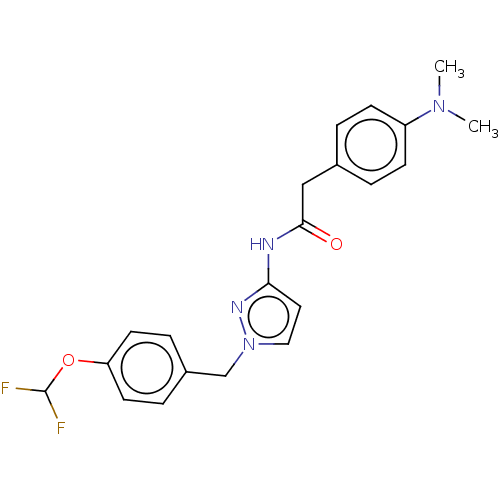
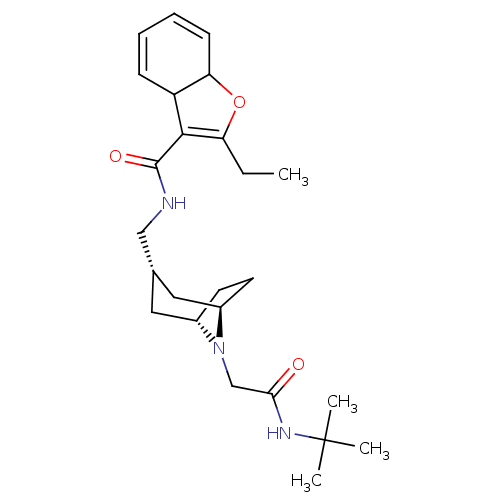
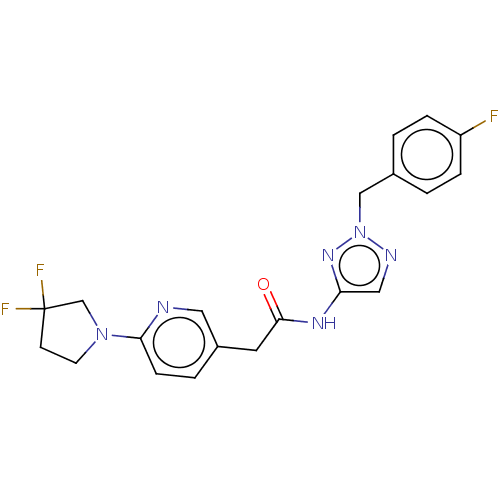
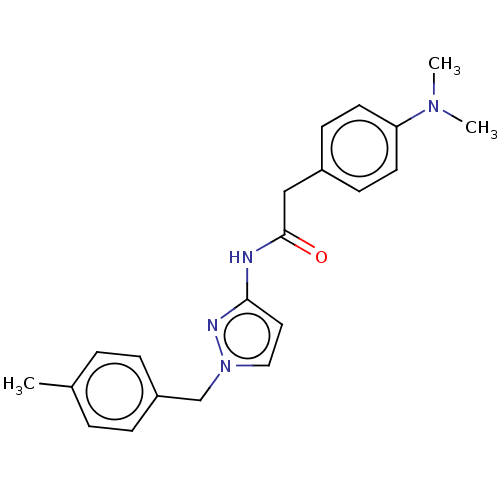
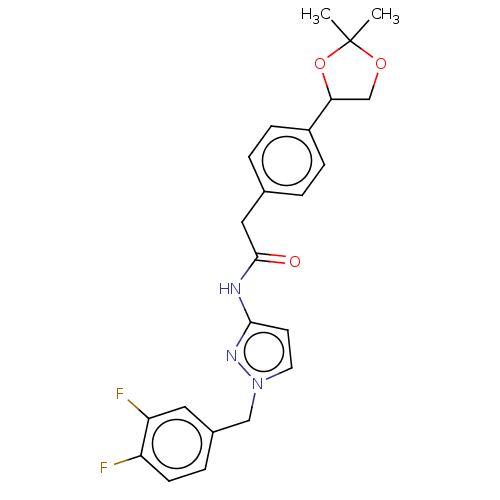
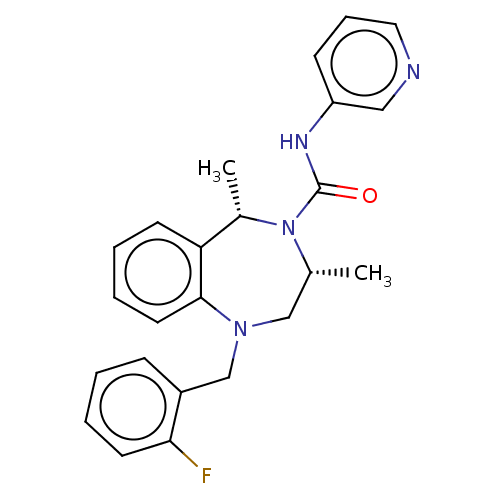
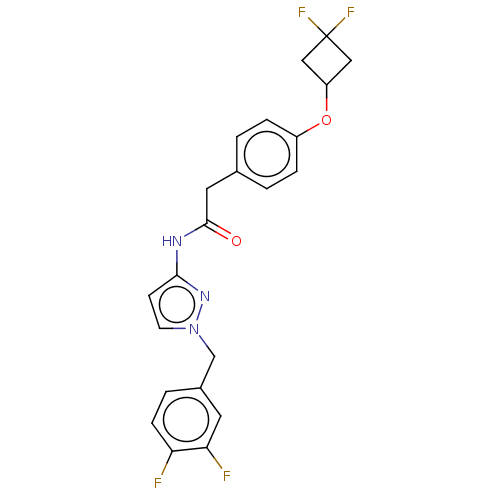
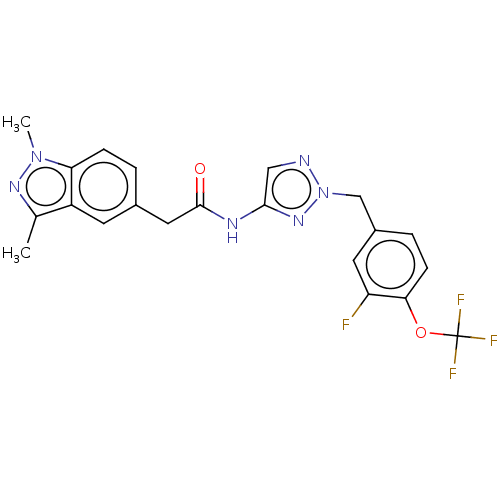
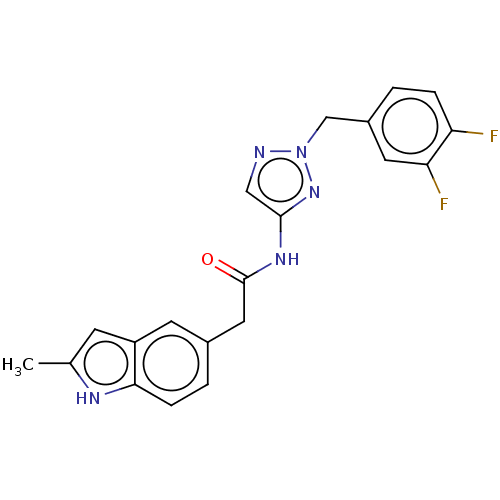
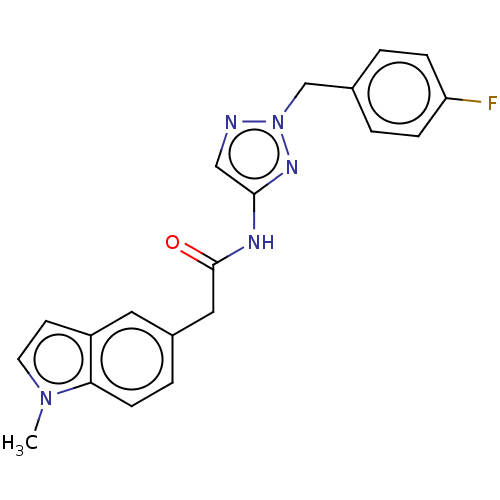
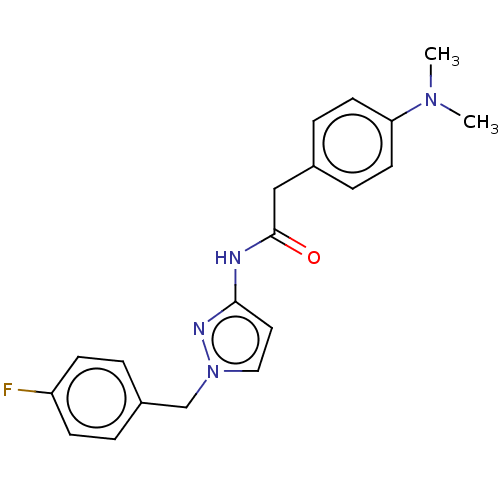
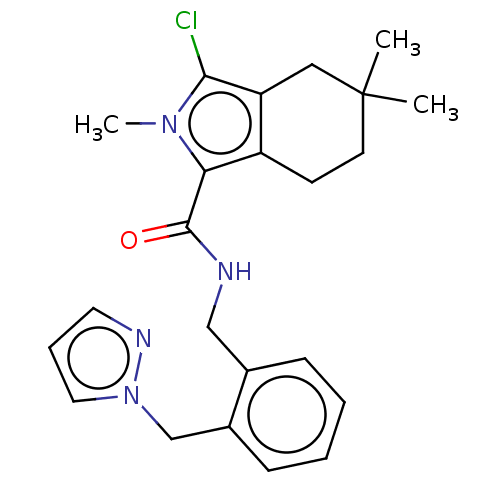
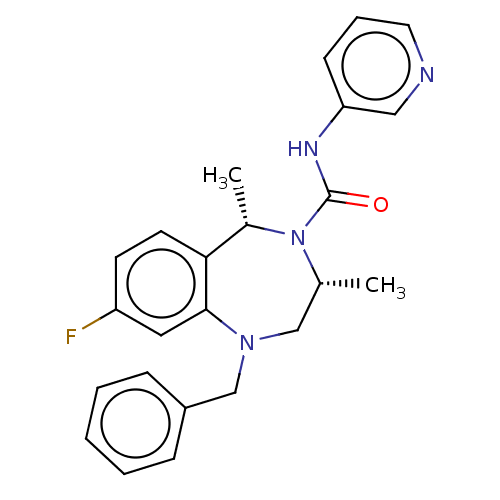
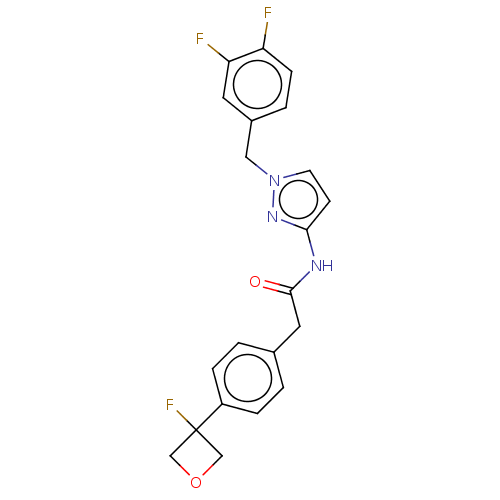
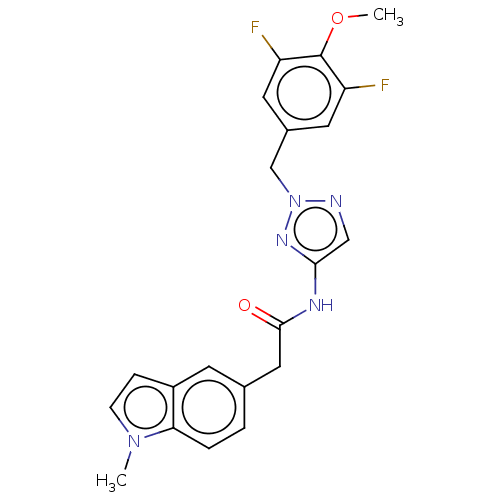
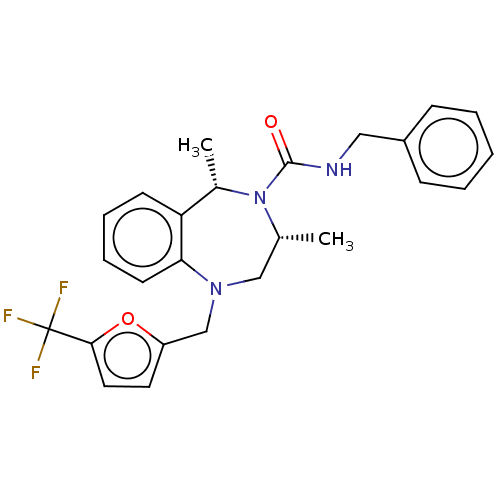
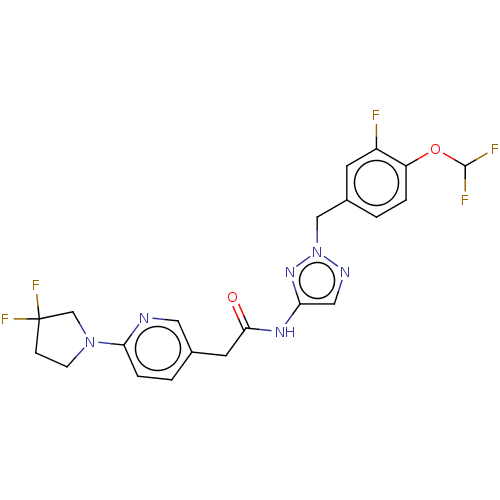
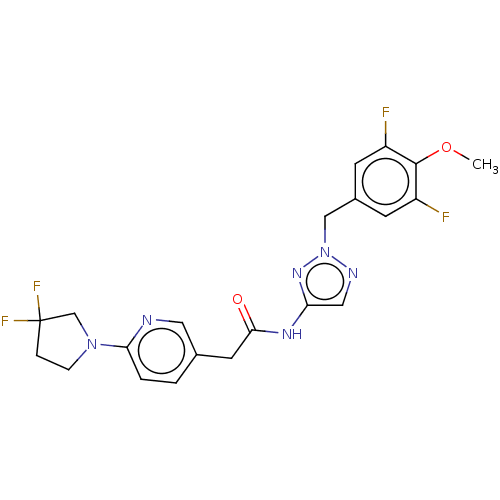
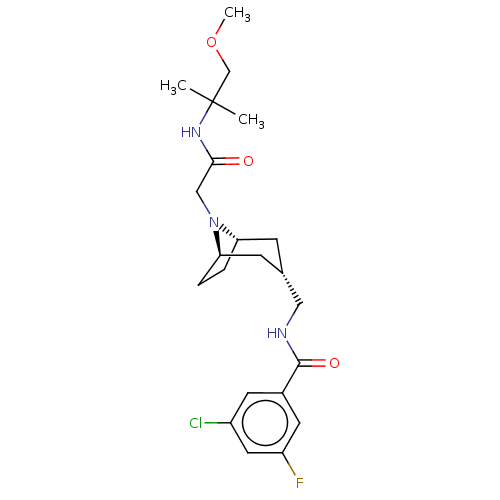
Affinity DataIC50: 3nMAssay Description:The HEK293 cells having human Cav3.2 stably expressed therein were cultured at 37° C. in Alpha-MEM, to which 10% (v/v) fetal bovine serum (FBS), peni...More data for this Ligand-Target Pair
Affinity DataIC50: 3.40nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.80nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of human CaV3.2 expressed in HEK293 cells at -100 mV membrane potential by Ionworks HT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of human CaV3.2 expressed in HEK293 cells at -100 mV membrane potential by Ionworks HT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.30nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.10nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.30nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Channel blocking activity at recombinant human CaV 3.2 channel expressed in HEK293 cells assessed as inhibition of Ca2+ flux preincubated for 3 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.10nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 6.40nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 6.60nMAssay Description:Inhibition of recombinant human Cav3.2 expressed in HEK293 cells assessed as reduction in CaCl2-induced intracellular calcium flux incubated for 20 m...More data for this Ligand-Target Pair
Affinity DataIC50: 6.70nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.30nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 7.40nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 7.5nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.80nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.80nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.20nMAssay Description:Inhibition of human T-type calcium channel Cav3.2 expressed in CHO cells assessed as inhibition of calcium tail current at -90 mV holding potential m...More data for this Ligand-Target Pair
Affinity DataIC50: 8.20nMAssay Description:Inhibition of CaV 3.2 channel (unknown origin) expressed in HEK293 cells by FLEPR Ca2+ flux assayMore data for this Ligand-Target Pair
Affinity DataIC50: 8.40nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.60nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.80nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:Channel blocking activity at recombinant human CaV 3.2 channel expressed in HEK293 cells assessed as inhibition of Ca2+ flux preincubated for 3 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.5nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 9.70nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 9.80nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 9.90nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:The HEK293 cells having human Cav3.2 stably expressed therein were cultured at 37° C. in Alpha-MEM, to which 10% (v/v) fetal bovine serum (FBS), peni...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Channel blocking activity at recombinant human CaV 3.2 channel expressed in HEK293 cells assessed as inhibition of Ca2+ flux preincubated for 3 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of recombinant human Cav3.2 expressed in HEK293 cells assessed as reduction in CaCl2-induced intracellular calcium flux incubated for 20 m...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Channel blocking activity at recombinant human CaV 3.2 channel expressed in HEK293 cells assessed as inhibition of Ca2+ flux preincubated for 3 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of recombinant Cav3.2 (unknown origin) expressed in HEK293 cells assessed as decrease in calcium flux preincubated for 3 mins followed by ...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Channel blocking activity at recombinant human CaV 3.2 channel expressed in HEK293 cells assessed as inhibition of Ca2+ flux preincubated for 3 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Stock solutions of test compounds are prepared to a concentration of 10 mM in DMSO. For the Cav3.2 assay, serial dilutions of the compounds are prepa...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:The HEK293 cells having human Cav3.2 stably expressed therein were cultured at 37° C. in Alpha-MEM, to which 10% (v/v) fetal bovine serum (FBS), peni...More data for this Ligand-Target Pair