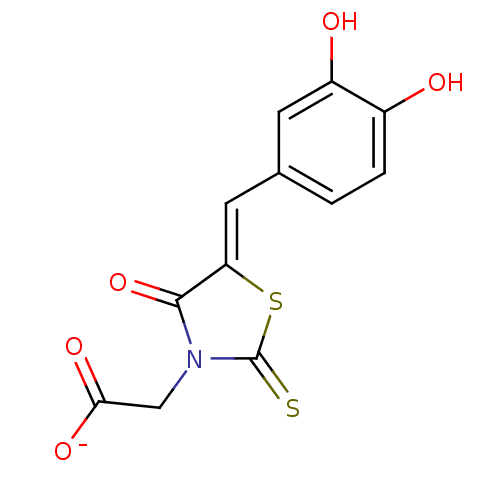
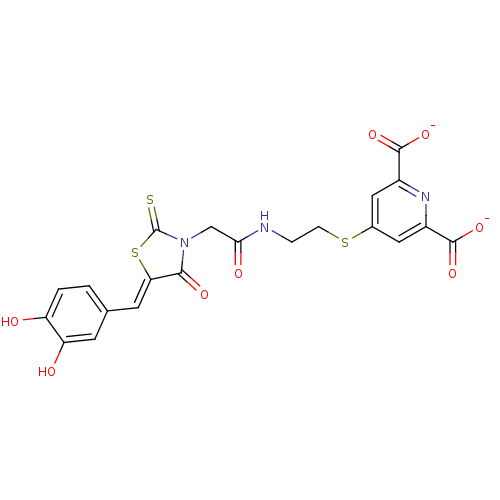
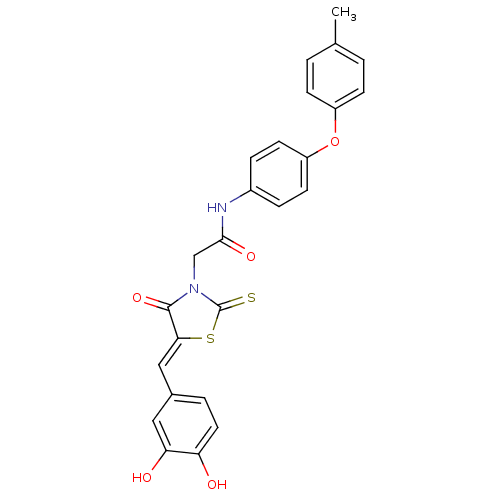
Affinity DataKi: 26nMpH: 7.4Assay Description:All reactions were monitored spectrophotometrically at 340 nm by using initial rates from the first 5% of reaction.More data for this Ligand-Target Pair
Affinity DataKi: 100nMpH: 7.4Assay Description:All reactions were monitored spectrophotometrically at 340 nm by using initial rates from the first 5% of reaction.More data for this Ligand-Target Pair
Affinity DataKi: >2.50E+4nMpH: 7.4Assay Description:All reactions were monitored spectrophotometrically at 340 nm by using initial rates from the first 5% of reaction.More data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMpH: 7.4Assay Description:All reactions were monitored spectrophotometrically at 340 nm by using initial rates from the first 5% of reaction.More data for this Ligand-Target Pair